Protocol PDF Document (version 1.2): fxst
Instructions:
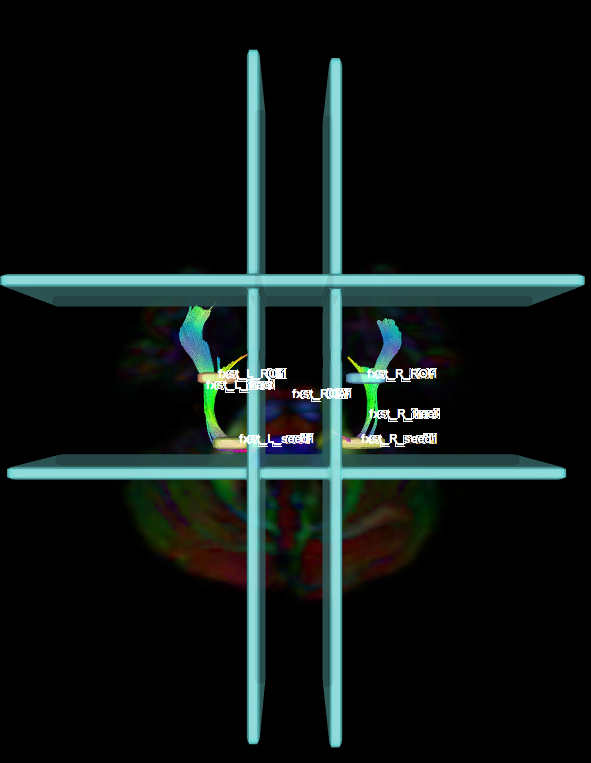
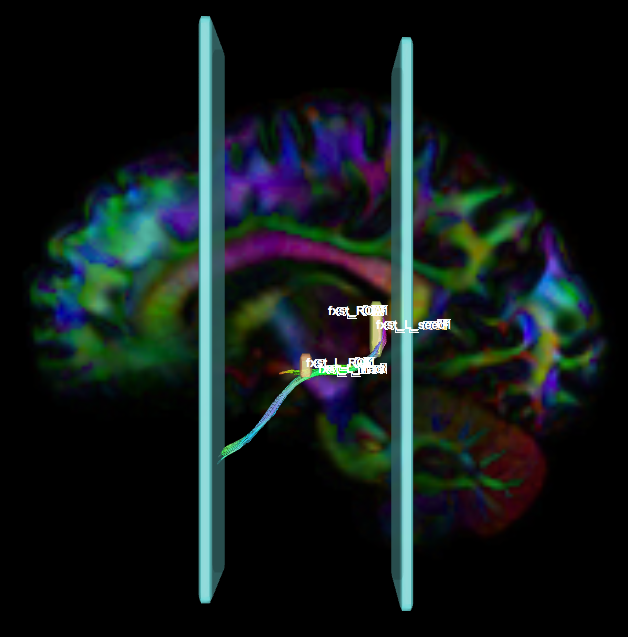
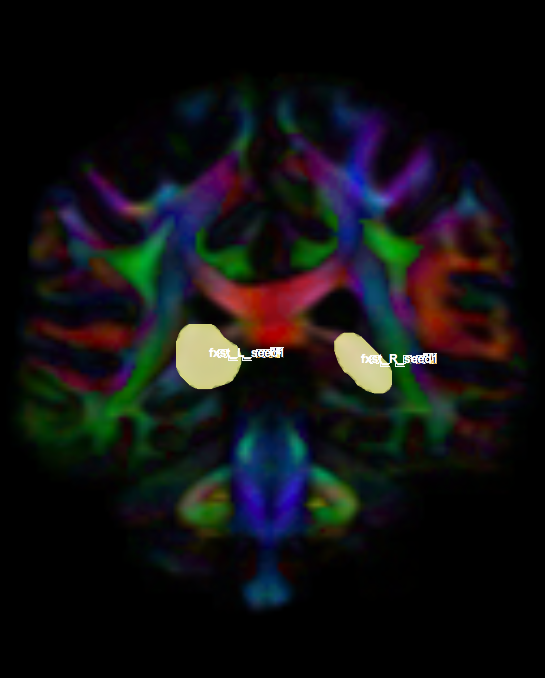
- Create two separate coronal seed regions (at approx. coronal slice 111), one for each side.
- Create two separate coronal ROI regions (at approx. coronal slice 91), one for each side.
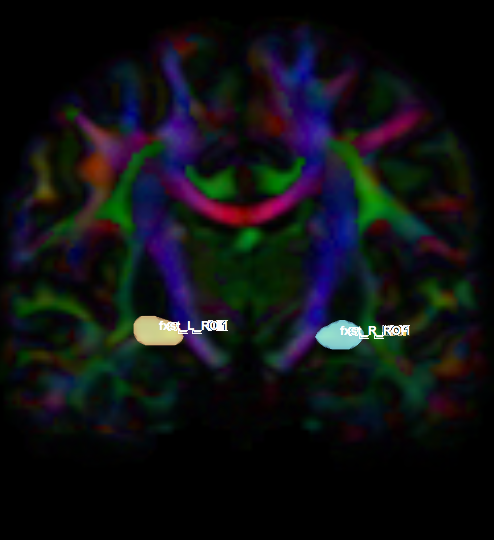
- Create an ROA region and draw two sagittal ROA slice: medial to the left seed and medial to the right seed.
- In the region list, check the left seed/ROI regions and ROA region, then run fiber tracking, based on this output, ROA placement will be clearer.
- Using the same ROA file, draw two more ROA regions:
- on a coronal slice posterior to the seed regions
- on a coronal slice anterior to the seed regions
- Check only the left seed/ROI region and ROA regions, then perform fiber tracking. Under the tract list, make sure only the desired left tract is checked and highlighted in purple. Save region, tract, and density files.
- Uncheck the left seed/ROI region and check the right seed/ROI region and ROA region, then perform fiber tracking. Make sure only the desired right tract is checked and highlighted in purple. Save region, tract, and density files.

Why did we separate the FX? Need anatomical descriptions for region placement (“slice 91” is not helpful. How big of regions? Justify ROA regions and provide anatomy
You neglected something..
Hi everyone ,thanks
Ciao These are truly great ideas in on the topic of blogging. You have touched some fastidious things here. Any way keep up wrinting. thank u
delta 8 oil syringe
We greatly enjoy your blog and find most of your posts to align well with our interests. Would you be open to guest contributors writing content for your blog? I would be happy to author a post or expand upon some of the topics you cover here. Your blog is excellent!
I cannot thank you enough for the post.Really looking forward to read more. Fantastic.
gem disco casino online
My gang on Google+ would want to read your post. Is it okay if I promote it to them?
Hello, Neat post. There’s a problem together with your site in internet explorer, may
test this? IE still is the market leader and a huge
component of people will leave out your excellent
writing because of this problem.
My brother recommended I might like this web site.
He was entirely right. This post actually made my day.
You cann’t imagine just how much time I had spent for this info!
Thanks!
Why people still make use of to read news papers when in this technological globe the
whole thing is accessible on web?
Hello! I know this is kind of off topic but I was wondering
if you knew where I could find a captcha plugin for my comment form?
I’m using the same blog platform as yours and I’m having difficulty
finding one? Thanks a lot!
Pretty! This was an incredibly wonderful article. Thank you for supplying
this information.
Thanks for sharing your thoughts on link alternatif olxtoto.
Regards
Hey! Do you use Twitter? I’d like to follow you if that
would be okay. I’m absolutely enjoying your blog and look
forward to new posts.
Quality articles or reviews is the key to interest the
users to go to see the web page, that’s what this web site is
providing.
I was curious if you ever thought of changing the structure of
your site? Its very well written; I love what youve got to say.
But maybe you could a little more in the way of content so people could connect with
it better. Youve got an awful lot of text for only having one or two pictures.
Maybe you could space it out better?
I think the admin of this website is genuinely working hard in support of his
web site, as here every stuff is quality based data.
This is very interesting, You’re a very skilled blogger. I have joined your rss feed
and look forward to seeking more of your magnificent post.
Also, I have shared your web site in my social networks!
certainly like your web-site but you have to test
the spelling on quite a few of your posts.
A number of them are rife with spelling problems and I in finding it very troublesome to inform the reality nevertheless I’ll surely come back again.
restaurant-lenvol.net
Wen Yansheng의 보증으로 Fang Jifan은 안도감을 느꼈습니다.
Greate article. Keep writing such kind of info on your page.
Im really impressed by it.
Hey there, You have done an incredible job. I’ll certainly digg it and in my opinion recommend to my friends.
I’m sure they’ll be benefited from this website.
Excellent blog here! Also your web site loads up fast!
What host are you using? Can I get your affiliate link
to your host? I wish my site loaded up as fast as yours lol
I am sure this paragraph has touched all the internet users,
its really really nice article on building up new web site.
Hi! This post couldn’t be written any better! Reading this post reminds me of my old
room mate! He always kept talking about this. I will forward this article to
him. Pretty sure he will have a good read. Many thanks for sharing!
Thanks for your marvelous posting! I actually enjoyed reading it, you are a great author.
I will make certain to bookmark your blog and may come back someday.
I want to encourage continue your great posts, have
a nice morning!
WOW just what I was looking for. Came here by searching for
male enlargement devices
Hi there to every one, because I am actually eager of reading this website’s post to be updated daily.
It contains good stuff.
My family members always say that I am killing my time here at
web, except I know I am getting know-how daily by reading thes fastidious articles.
I’m extremely impressed together with your writing abilities and also with the layout to your
blog. Is that this a paid subject or did you customize it yourself?
Either way keep up the excellent high quality writing,
it’s uncommon to look a nice blog like this one nowadays..
Nice post. I learn something totally new and challenging on websites I stumbleupon on a daily basis.
It’s always exciting to read through content from other writers and practice a little something from other web sites.
Great article! We are linking to this particularly great post on our site.
Keep up the good writing.
Howdy I am so happy I found your site, I really found you by error, while I was looking on Digg for something else, Anyways I
am here now and would just like to say thanks for a incredible post and a all round enjoyable blog (I also love
the theme/design), I don’t have time to read it all at the moment
but I have saved it and also added your RSS feeds, so when I have
time I will be back to read much more, Please do keep up the great job.
Way cool! Some extremely valid points! I appreciate
you writing this write-up and also the rest of the website is also very good.
Hello! I’m at work browsing your blog from my new iphone
3gs! Just wanted to say I love reading your blog and look forward to all your
posts! Carry on the great work!
I am genuinely pleased to read this blog posts which contains plenty of useful data, thanks for providing such information.
Hello, i read your blog from time to time and i own a similar one and i was just curious
if you get a lot of spam feedback? If so how do you reduce it, any plugin or anything you can recommend?
I get so much lately it’s driving me insane so any help is very much appreciated.
Hi, this weekend is pleasant designed for me, as this moment i am reading this fantastic educational post
here at my residence.
Pretty section of content. I just stumbled upon your
web site and in accession capital to say that I acquire actually enjoyed account your blog posts.
Any way I’ll be subscribing on your augment and even I fulfillment you access constantly quickly.
Hello there! I know this is kinda off topic but I was wondering if you knew where I could
locate a captcha plugin for my comment form?
I’m using the same blog platform as yours and I’m having trouble finding one?
Thanks a lot!
Please let me know if you’re looking for a author for
your site. You have some really great articles and I think I would be a good asset.
If you ever want to take some of the load off, I’d love
to write some material for your blog in exchange for a link back to mine.
Please send me an e-mail if interested. Kudos!
Hello i am kavin, its my first time to commenting anywhere, when i read
this article i thought i could also make comment due to this
good article.
Usually I don’t learn article on blogs, but I wish to say that
this write-up very compelled me to check out and
do so! Your writing taste has been surprised me.
Thanks, quite nice post.
Ahaa, its good conversation concerning this post at this place at this
weblog, I have read all that, so now me also commenting
at this place.
What’s up to every one, it’s truly a nice for me to pay
a quick visit this website, it contains valuable Information.
Hello! Do you use Twitter? I’d like to follow you if that would be okay.
I’m definitely enjoying your blog and look forward to new updates.
Hurrah, that’s what I was exploring for, what a material!
existing here at this blog, thanks admin of this web site.
I’m amazed, I have to admit. Seldom do I come across a blog that’s both educative and entertaining, and let me tell you, you have hit the
nail on the head. The issue is something that too few men and women are speaking intelligently
about. I’m very happy I came across this in my search for
something relating to this.
Peculiar article, totally what I was looking for.
Its like you read my mind! You appear to understand so much about this, such as you wrote the
e book in it or something. I think that you simply could do with a few p.c.
to force the message home a bit, but instead of that, this is magnificent blog.
A fantastic read. I will certainly be back.
I’ve been exploring for a little bit for any high-quality articles
or weblog posts on this sort of area . Exploring in Yahoo I eventually stumbled upon this website.
Studying this information So i’m glad to show that
I’ve an incredibly excellent uncanny feeling I found out just what I
needed. I so much indisputably will make certain to
do not disregard this website and give it a look regularly.
After exploring a few of the blog articles on your website, I honestly appreciate your technique of blogging.
I saved as a favorite it to my bookmark website list and will be checking back soon. Please check
out my website too and let me know how you feel.
Amazing! Its really amazing post, I have got much clear idea regarding from
this post.
Ahaa, its good dialogue on the topic of this article here at this
weblog, I have read all that, so now me also commenting at this place.
I am genuinely pleased to glance at this blog posts which contains lots of valuable data, thanks for providing such data.
I am actually happy to glance at this website posts which consists of tons of useful information, thanks for providing such statistics.
It’s appropriate time to make some plans for the future and
it’s time to be happy. I’ve learn this submit and if I could I desire to suggest you few attention-grabbing issues or tips.
Maybe you could write next articles relating to this article.
I want to learn more issues approximately it!
I have read so many posts about the blogger lovers except this post is
genuinely a good post, keep it up.
I simply couldn’t depart your site before suggesting that I actually loved
the standard info an individual provide on your guests?
Is going to be again incessantly in order to investigate cross-check new posts
This design is steller! You obviously know how
to keep a reader entertained. Between your wit and your videos, I was almost moved
to start my own blog (well, almost…HaHa!) Excellent job.
I really enjoyed what you had to say, and more than that, how you presented it.
Too cool!
Great delivery. Great arguments. Keep up the good effort.
With havin so much content and articles do you
ever run into any issues of plagorism or copyright infringement?
My blog has a lot of exclusive content I’ve either authored myself or outsourced but it appears a lot of it is popping it
up all over the internet without my permission. Do you know any ways to help protect against content from being stolen? I’d certainly appreciate it.
Hey there exceptional website! Does running a blog such as this require
a massive amount work? I have virtually no knowledge of coding but I had been hoping
to start my own blog in the near future. Anyhow, should you have any ideas or techniques for new blog
owners please share. I know this is off subject however I just needed to ask.
Kudos!
I am actually thankful to the holder of this
web site who has shared this impressive paragraph at at this time.
Hi fantastic website! Does running a blog like this take a great
deal of work? I’ve absolutely no knowledge of computer programming but I had been hoping
to start my own blog in the near future. Anyhow, should you have any recommendations or techniques for new blog owners please share.
I understand this is off subject but I just had to ask.
Thanks a lot!
Genuinely when someone doesn’t know then its up to other users that they will help, so here it
takes place.
I blog often and I really appreciate your content. The article has
truly peaked my interest. I am going to book mark your website
and keep checking for new details about once per week. I subscribed to your RSS feed as well.
Yes! Finally something about Walnut.
I got this site from my friend who informed me on the topic of this web
page and at the moment this time I am visiting this web page and
reading very informative articles or reviews at this
time.
This text is invaluable. How can I find out more?
I visited several web pages except the audio quality for audio songs present at this web page is in fact
wonderful.
Hello it’s me, I am also visiting this web site daily, this website is really
pleasant and the people are really sharing fastidious thoughts.
You need to be a part of a contest for one of the highest quality blogs on the net.
I will highly recommend this web site!
I’m amazed, I must say. Rarely do I encounter a blog that’s both equally educative and entertaining,
and without a doubt, you have hit the nail on the head.
The issue is something which too few folks
are speaking intelligently about. Now i’m very happy I found this in my search for something regarding this.
Fantastic beat ! I wish to apprentice while you amend your web site, how can i subscribe for a weblog website?
The account aided me a applicable deal. I were tiny bit acquainted of
this your broadcast offered vivid transparent idea
Wow, that’s what I was looking for, what a information! existing here at this
weblog, thanks admin of this web page.
Excellent beat ! I would like to apprentice while you amend your web site, how
can i subscribe for a blog web site? The account helped me a acceptable deal.
I had been tiny bit acquainted of this your broadcast offered
bright clear idea
May I have information on the topic of your article?
Keep on writing, great job!
It is the best time to make some plans for the longer term and it’s time to be happy.
I have read this submit and if I could I wish to suggest you few
interesting things or advice. Maybe you could write next articles
referring to this article. I want to read even more things
approximately it!
This is a topic which is close to my heart… Best wishes!
Exactly where are your contact details though?
When someone writes an piece of writing he/she maintains the thought
of a user in his/her brain that how a user can know it.
Thus that’s why this article is outstdanding. Thanks!
Hi there, I enjoy reading through your article. I wanted to write a little comment to support you.
What’s up, I want to subscribe for this web site to get newest updates, therefore where can i do it please help.
Hello there, I found your blog by way of Google while looking for a
comparable topic, your website came up,
it seems great. I’ve bookmarked it in my google bookmarks.
Hello there, simply changed into alert to your weblog thru Google, and located that it is truly
informative. I’m going to be careful for brussels. I will appreciate if you happen to continue this in future.
Numerous other people will be benefited out of
your writing. Cheers!
Greetings! Very useful advice within this post! It is the little changes that will make
the biggest changes. Thanks for sharing!
Thanks for sharing such a fastidious idea, article is nice,
thats why i have read it entirely
I love your blog.. very nice colors & theme.
Did you make this website yourself or did you hire someone
to do it for you? Plz respond as I’m looking to construct my own blog and
would like to find out where u got this from. many
thanks
This is my first time pay a visit at here and
i am actually pleassant to read everthing at alone place.
Hi there! This is kind of off topic but I need some help from
an established blog. Is it hard to set up your own blog? I’m not very techincal but I can figure things out
pretty fast. I’m thinking about making my own but I’m not sure where to start.
Do you have any points or suggestions? Many thanks
My brother recommended I may like this blog. He used to be
entirely right. This post truly made my day. You can not imagine just how a lot time I had spent for this info!
Thanks!
It’s going to be finish of mine day, except before finish I am reading this enormous post to improve my
know-how.
This website was… how do you say it? Relevant!! Finally I’ve found something that helped me.
Cheers!
I know this website provides quality dependent articles or reviews and other material, is there any
other web page which provides such things in quality?
Hi, just wanted to mention, I loved this post. It was inspiring.
Keep on posting!
Truly no matter if someone doesn’t know after that its up to other viewers that they will help, so here it takes place.
Great post! We will be linking to this particularly great content on our website.
Keep up the good writing.
Thanks for finally talking about > Fornix (crus)/ Stria Terminalis – fxst | Whole Brain Protocol for
Tractography with Empirical MRI < Liked it!
Hi, I think your website might be having browser compatibility issues.
When I look at your blog in Chrome, it looks fine but when opening in Internet Explorer, it has some overlapping.
I just wanted to give you a quick heads up!
Other then that, fantastic blog!
Thank you for the good writeup. It in fact was a amusement account it.
Look advanced to far added agreeable from you!
By the way, how could we communicate?
I’m not sure where you are getting your info,
but great topic. I needs to spend some time
learning more or understanding more. Thanks for fantastic
information I was looking for this info for my mission.
Heya i’m for the first time here. I came across this board and I
find It truly helpful & it helped me out a lot. I hope to offer one thing again and help others like you aided me.
I read this piece of writing fully on the topic
of the difference of most up-to-date and previous technologies, it’s awesome article.
Keep this going please, great job!
saungsantoso.com
나는 Zhang Xin을 만나 그에게 철저한 이해를 주었고 Zhu Houzhao의 사업이었습니다.
sm 바카라
여덟 번째 순위는 Shuntian Juren Jiang Chen의 이름입니다.
https://alepz.com/
I just like the helpful info you supply in your
articles. I’ll bookmark your blog and test once more here
frequently. I’m reasonably sure I will learn many new
stuff proper here! Best of luck for the following!
Can you tell us more about this? I’d care to find out some
additional information.
I was extremely pleased to uncover this page.
I need to to thank you for ones time just for this wonderful read!!
I definitely savored every little bit of
it and i also have you saved as a favorite
to look at new information in your web site.
Hi there! Someone in my Myspace group shared this site with
us so I came to check it out. I’m definitely loving the information. I’m book-marking and will be tweeting this to my
followers! Terrific blog and brilliant style and design.
Good post. I’m going through many of these issues as well..
When I originally commented I clicked the “Notify me when new comments are added” checkbox and now each time a comment is added I get
several e-mails with the same comment. Is there any way you
can remove people from that service? Cheers!
Hey there! Do you know if they make any plugins to assist with Search Engine Optimization? I’m trying
to get my blog to rank for some targeted keywords but I’m not seeing very good success.
If you know of any please share. Many thanks!
Nice post. I learn something totally new and challenging on blogs
I stumbleupon every day. It will always be useful to read
articles from other authors and practice a little something from their
sites.
Magnificent goods from you, man. I have take into account your
stuff previous to and you are simply too wonderful. I really like what you’ve got right
here, really like what you’re saying and the best way in which you say it.
You’re making it enjoyable and you continue to care for
to stay it smart. I can not wait to learn much more from you.
This is really a tremendous site.
I am really loving the theme/design of your weblog. Do you ever
run into any web browser compatibility problems?
A handful of my blog readers have complained about my site not operating correctly in Explorer but looks
great in Chrome. Do you have any ideas to help fix this problem?
Actually no matter if someone doesn’t be aware of after that its up to other people that they will help, so here it occurs.
Aw, this was a very good post. Taking a few minutes and actual effort to make a really good article… but what can I say… I put things
off a lot and don’t manage to get anything done.
Excellent way of explaining, and good paragraph to obtain information concerning my presentation topic, which
i am going to convey in institution of higher education.
This is really interesting, You’re a very skilled blogger.
I have joined your rss feed and look forward to seeking more of your magnificent
post. Also, I have shared your website in my social networks!
iGaming 업계에서 앞서가는 프라그마틱 게임은 모바일 중심의 혁신적이고 표준화된 콘텐츠를 제공합니다.
PG 소프트
프라그마틱의 게임은 항상 다양한 테마로 놀라워요. 이 사이트에서 더 자세한 정보를 찾아보세요!
https://vispills.com/
http://dlinkwifiext.site
https://www.xmcoart.com
Pingback: URL
I’m really loving the theme/design of your web site.
Do you ever run into any web browser compatibility issues?
A few of my blog readers have complained about my website not operating correctly in Explorer but looks great in Chrome.
Do you have any suggestions to help fix this problem?
This design is spectacular! You certainly know how to keep a reader entertained.
Between your wit and your videos, I was almost moved to start my
own blog (well, almost…HaHa!) Wonderful job.
I really enjoyed what you had to say, and more than that, how you presented it.
Too cool!
I am genuinely grateful to the holder of this site who has shared
this great piece of writing at here.
Great blog here! Also your web site loads up fast!
What web host are you using? Can I get your affiliate link to your host?
I wish my web site loaded up as fast as yours lol
When I originally commented I clicked the “Notify me when new comments are added”
checkbox and now each time a comment is added I get three e-mails with
the same comment. Is there any way you can remove people
from that service? Cheers!
Greetings! Very useful advice in this particular post! It’s the little changes that will make the biggest changes.
Thanks for sharing!
I read this article completely on the topic of the difference of newest and earlier technologies, it’s remarkable
article.
Hello to every one, the contents present at this web site are genuinely remarkable for
people knowledge, well, keep up the nice work fellows.
Hello, i think that i saw you visited my web site so i came to
“return the favor”.I am attempting to find things to improve my website!I suppose its ok to use
some of your ideas!!
This post to be quite enlightening. Thank you for sharing it.
#1Welcome to the my best #site:
http://ingfootball.ru/userinfo.php?uid=93606
https://www.magcloud.com/user/leonidborisov
http://wwwnickru.bestbb.ru/viewtopic.php?id=128#p128
https://aqvakr.forum24.ru/?1-7-0-00010228-000-0-0-1707748676
http://mtw2014.tmweb.ru/forum/?PAGE_NAME=read&FID=1&TID=7839&TITLE_SEO=7839-kupit-diplom-o-vysshem-obrazovanii
Great goods from you, man. I have understand your stuff
previous to and you’re just too fantastic. I really like what you’ve acquired here, certainly like what
you are saying and the way in which you say it. You make it entertaining and you still take
care of to keep it smart. I can not wait to read much more from you.
This is actually a wonderful web site.
I loved as much as you will receive carried out right here.
The sketch is attractive, your authored material stylish.
nonetheless, you command get got an shakiness over
that you wish be delivering the following.
unwell unquestionably come further formerly again since
exactly the same nearly very often inside case you shield this increase.
Its like you read my mind! You appear to know a lot about this, like
you wrote the book in it or something. I think that you could
do with some pics to drive the message home a little bit,
but instead of that, this is excellent blog. A great read.
I will certainly be back.
We’re a gaggle of volunteers and opening a new scheme in our community.
Your site provided us with useful info to work on. You
have done an impressive process and our whole group will be grateful to you.
Write more, thats all I have to say. Literally, it seems as though you relied on the video to make your point. You clearly know what youre talking about, why waste your intelligence on just posting videos to your site when you could be giving us something informative to read?
프라그마틱과 관련된 이 글 정말 잘 읽었어요! 더불어 저도 제 사이트에서 프라그마틱에 대한 새로운 정보를 공유 중이에요. 함께 나누면 더 좋을 것 같아요!
프라그마틱 홈페이지
프라그마틱의 게임을 플레이하면 항상 긴장감 넘치고 즐거운 시간을 보낼 수 있어 좋아요. 여기서 더 많은 이야기를 들려주세요!
https://www.genericsingulair.site
https://www.eprust.com
http://usmcafee.site
This is really fascinating, You are a very skilled blogger.
I’ve joined your feed and sit up for looking for extra of your wonderful post.
Additionally, I have shared your site in my social
networks
If you want to buy the best quality reusable paper towels and become eco friendly take a look at the
listed website. These are bamboo paper towels and bamboo towels are environmentally friendly.
Hola! I’ve been reading your blog for a while now and finally got the courage to go ahead and give you
a shout out from Austin Tx! Just wanted to tell you keep up the excellent job!
Appreciate the recommendation. Will try it out.
agonaga.com
그리고 해변에는 온통 시체가 널브러져 있습니다.
What i don’t realize is in reality how you’re no longer really a lot more smartly-favored than you may be right now.
You’re very intelligent. You know therefore considerably in relation to
this matter, made me in my opinion consider it from a
lot of various angles. Its like women and men aren’t interested except it is something to do
with Woman gaga! Your own stuffs outstanding. All the time deal
with it up!
I will immediately snatch your rss as I can’t in finding
your email subscription link or e-newsletter service.
Do you have any? Kindly permit me understand so that I may
subscribe. Thanks.
I like it when individuals get together and share ideas. Great site, keep
it up!
Usually I do not read post on blogs, but I wish to say that this write-up very pressured me to take a look at and do it!
Your writing taste has been amazed me. Thanks, very nice article.
There’s definately a lot to find out about this topic.
I love all the points you have made.
Cialis Doctor Simi
Excuse for that I interfere … To me this situation is familiar. Is ready to help.
Cialis 5 mg prezzo cialis prezzo cialis 5 mg prezzo
Cialis 20
Bravo, this magnificent idea is necessary just by the way
Cialis 5 mg prezzo tadalafil 5 mg prezzo tadalafil 5 mg prezzo
Donde Comprar Cialis GenГ©rico Fiable
I join. It was and with me.
Cialis 5 mg prezzo cialis 5 mg prezzo cialis 5 mg prezzo
Greetings! This is my first visit to your blog! We are a collection of volunteers and starting a new initiative in a community in the same niche.
Your blog provided us valuable information to work on. You have done a wonderful job!
This is my first time visit at here and i am really impressed to read all at alone place.
It’s going to be ending of mine day, except before finish I am reading this enormous piece of writing to increase my knowledge.
Howdy I am so delighted I found your webpage, I really found you by mistake, while I was researching
on Aol for something else, Anyhow I am here now and would just like to say
many thanks for a fantastic post and a all round enjoyable blog (I also love
the theme/design), I don’t have time to go through it all at the minute but I have saved
it and also added your RSS feeds, so when I have time I
will be back to read a lot more, Please do keep up the fantastic jo.
When someone writes an paragraph he/she keeps the thought of a
user in his/her brain that how a user can understand it.
So that’s why this article is outstdanding. Thanks!
Just wish to say your article is as amazing. The clarity in your post is just spectacular and i can assume you’re an expert on this subject.
Fine with your permission allow me to grab your RSS feed to keep up to
date with forthcoming post. Thanks a million and please continue the
gratifying work.
This article is truly a fastidious one it helps new internet users,
who are wishing for blogging.
When I initially commented I seem to have clicked on the -Notify me when new comments are added- checkbox and from now on each time a comment is added I receive 4 emails
with the same comment. There has to be an easy method you are able to remove me from that service?
Thanks a lot!
I think that everything typed made a lot of sense.
But, what about this? what if you added a little content?
I mean, I don’t wish to tell you how to run your website, but suppose you added a post title that makes people want more?
I mean Fornix (crus)/ Stria Terminalis – fxst | Whole Brain Protocol for Tractography with Empirical MRI is kinda vanilla.
You ought to look at Yahoo’s front page and note how they create post headlines to get viewers interested.
You might try adding a video or a pic or two to get readers excited about what you’ve written. Just
my opinion, it might make your blog a little bit more interesting.
This post will assist the internet viewers for setting up new web site or even a blog from start to end.
Link exchange is nothing else except it is just placing the other person’s webpage link on your page at appropriate place and other person will also do same in support of you.
What’s up, its nice piece of writing regarding media print,
we all understand media is a great source of facts.
Thanks , I have recently been looking for info approximately
this topic for ages and yours is the greatest I have discovered till now.
But, what about the bottom line? Are you positive in regards to the
supply?
Wow, this piece of writing is good, my sister is analyzing these kinds of things, thus I am going to let know her.
Thank you for writing this post. I like the subject too.
What’s up colleagues, how is all, and what you want
to say about this article, in my view its genuinely remarkable in favor of me.
When someone writes an article he/she maintains the plan of a user in his/her brain that how a user can know
it. Thus that’s why this article is outstdanding.
Thanks!
Thank you for your articles. I find them very helpful. Could you help me with something?
When some one searches for his necessary thing,
so he/she wants to be available that in detail, thus that thing is maintained over here.
Hi, I check your new stuff like every week. Your writing style is awesome,
keep doing what you’re doing!
I truly love your website.. Very nice colors & theme. Did you make
this site yourself? Please reply back as I’m hoping to create my own blog and would like to know where you got this
from or exactly what the theme is named. Many thanks!
I would like to thank you for the efforts you’ve put in penning this website.
I am hoping to see the same high-grade content by you in the future as well.
In truth, your creative writing abilities has motivated me to get my own website now 😉
I’m not certain where you are getting your info, but great topic.
I must spend some time learning much more or understanding more.
Thank you for wonderful information I used to be searching for this information for my mission.
This website truly has all the information and facts I needed
about this subject and didn’t know who to ask.
After checking out a handful of the blog articles on your site,
I truly appreciate your way of writing a
blog. I saved it to my bookmark site list and will be checking back soon. Please check out my website
too and let me know what you think.
I was wondering if you ever considered changing the layout of your website?
Its very well written; I love what youve got to say.
But maybe you could a little more in the way of content so
people could connect with it better. Youve got an awful
lot of text for only having 1 or two images. Maybe you
could space it out better?
Heya i am for the primary time here. I came across this board and I to find It really useful & it helped me out much.
I’m hoping to offer one thing again and help others like you aided
me.
Thank you for your articles. They are very helpful to me. May I ask you a question?
Hi there to every single one, it’s actually a pleasant for me to
pay a visit this web page, it consists of helpful Information.
For hottest information you have to go to see web and on world-wide-web I found this
website as a finest web site for newest updates.
Hey there, I think your website might be having browser compatibility issues.
When I look at your blog in Firefox, it looks fine but when opening
in Internet Explorer, it has some overlapping.
I just wanted to give you a quick heads up! Other then that, amazing blog!
It’s an awesome article in favor of all the web
viewers; they will obtain advantage from it I am sure.
Appreciate this post. Will try it out.
I was curious if you ever considered changing the page layout of your site?
Its very well written; I love what youve got to say. But maybe you could a little more in the way of content so people could connect with
it better. Youve got an awful lot of text for only having 1 or two pictures.
Maybe you could space it out better?
Everything is very open with a really clear explanation of the issues.
It was really informative. Your site is extremely helpful.
Many thanks for sharing!
whoah this blog is magnificent i love reading your articles.
Keep up the good work! You know, lots of individuals are searching round for this information, you
could help them greatly.
Sweet blog! I found it while searching on Yahoo News. Do you have any tips on how to get listed in Yahoo News?
I’ve been trying for a while but I never seem to get there!
Thanks
This site was… how do I say it? Relevant!! Finally I’ve found something
which helped me. Thanks!
Definitely believe that that you stated.
Your favourite reason seemed to be at the web the simplest thing to consider of.
I say to you, I certainly get irked whilst folks think about concerns that they just do not
recognise about. You controlled to hit the nail upon the top as smartly as outlined
out the whole thing without having side effect , other people can take a signal.
Will probably be again to get more. Thank you
Hi i am kavin, its my first occasion to commenting anyplace, when i read
this article i thought i could also make comment due to this brilliant post.
Hey there, I think your site might be having browser compatibility issues.
When I look at your blog site in Opera, it looks fine but when opening in Internet Explorer, it has some overlapping.
I just wanted to give you a quick heads up! Other then that, wonderful blog!
Hi there, its fastidious paragraph regarding media print, we all be familiar
with media is a fantastic source of data.
Hello there! I know this is somewhat off topic but I
was wondering which blog platform are you using for
this website? I’m getting fed up of WordPress because I’ve had issues with hackers and I’m looking
at options for another platform. I would be fantastic if you could point me in the direction of a
good platform.
I was more than happy to find this web site. I want to to thank
you for ones time just for this fantastic read!!
I definitely appreciated every bit of it and I have you saved to
fav to look at new information in your web site.
Heya! I understand this is somewhat off-topic however I had to ask.
Does managing a well-established blog like yours require
a massive amount work? I am completely new to running a blog but I do write in my diary everyday.
I’d like to start a blog so I can share my own experience and thoughts online.
Please let me know if you have any ideas or tips for brand new aspiring blog owners.
Appreciate it!
Hi there, I found your web site by means of Google even as
looking for a similar subject, your site came up, it seems good.
I’ve bookmarked it in my google bookmarks.
Hi there, just was alert to your blog through Google, and located that
it is truly informative. I’m gonna watch
out for brussels. I will appreciate for those who proceed this in future.
A lot of other people will be benefited out of your writing.
Cheers!
Hi there, just became aware of your blog through Google, and found
that it’s truly informative. I’m gonna watch out for brussels.
I will be grateful if you continue this in future.
Many people will be benefited from your writing.
Cheers!
Thanks for some other informative web site. The place
else may I get that kind of information written in such an ideal way?
I have a project that I am simply now running on, and I’ve been at the look out for such info.
I have been exploring for a little bit for any high quality articles
or blog posts in this sort of space . Exploring in Yahoo I at last stumbled upon this web site.
Reading this info So i am glad to show that I’ve a very just right uncanny feeling
I came upon just what I needed. I such a lot indisputably will make certain to don?t forget this website and provides it a look on a constant basis.
Hi there! I know this is kinda off topic however I’d figured I’d ask.
Would you be interested in trading links
or maybe guest authoring a blog post or vice-versa? My site discusses a lot of the same topics
as yours and I believe we could greatly benefit from each other.
If you might be interested feel free to send me an e-mail.
I look forward to hearing from you! Great blog by the way!
프라그마틱 플레이의 홈페이지에서 더 많은 슬롯을 찾아보세요.
프라그마틱 플레이
프라그마틱 게임은 정말로 혁신적이에요. 특히 슬롯 게임들은 항상 기대 이상의 재미를 선사합니다!
https://www.buytretinoin.site
https://www.deardiarytheep.com
https://www.unex-tech.com
It is appropriate time to make some plans for the
future and it’s time to be happy. I have read this post and if I could
I desire to suggest you few interesting things or advice.
Maybe you can write next articles referring to this article.
I wish to read even more things about it!
Hi i am kavin, its my first time to commenting anywhere,
when i read this paragraph i thought i could also create comment due to this brilliant post.
Good post. I learn something new and challenging on websites
I stumbleupon everyday. It’s always interesting to read through content from other writers and use
something from their websites.
netovideo.com
순종적이고 순종적으로 들으면 Fang Jifan을 위해 일하면 도움이 될 것입니다.
Quality posts is the crucial to attract the visitors to visit the web site, that’s what this
site is providing.
Hello I am so glad I found your blog page, I really found you by
error, while I was researching on Google for something else,
Anyways I am here now and would just like to say thank you for a tremendous post and
a all round entertaining blog (I also love the theme/design),
I don’t have time to read it all at the minute but I have saved it and
also added in your RSS feeds, so when I have time I will be back to read a lot more, Please do
keep up the great job.
I’m really impressed with your writing skills and also with the layout
on your blog. Is this a paid theme or did you modify it yourself?
Either way keep up the excellent quality writing, it’s rare to see a nice blog like
this one today.
Good blog you have here.. It’s hard to find excellent writing like yours nowadays.
I really appreciate people like you! Take care!!
Wow, marvelous weblog structure! How lengthy have you ever been running a blog for?
you made blogging glance easy. The entire glance of your web site
is great, as smartly as the content!
Actually no matter if someone doesn’t be aware of then its up to other people that they will assist, so here it occurs.
Because the admin of this web site is working, no doubt
very soon it will be renowned, due to its quality contents.
Keep this going please, great job!
I know this web site provides quality dependent posts and other
stuff, is there any other site which gives these kinds of
stuff in quality?
Hello there! This is my first visit to your blog! We are a team of volunteers and starting
a new initiative in a community in the same niche.
Your blog provided us beneficial information to work on. You have done a outstanding job!
It is really a great and useful piece of information. I am satisfied that you shared this helpful
info with us. Please stay us up to date like this. Thank
you for sharing.
Авиатор Спрайб играть на рубли
You are not right. I can prove it. Write to me in PM, we will talk.
Добро пожаловать в захватывающий мир авиаторов! Aviator – это увлекательная игра, которая позволит вам окунуться в атмосферу боевых действий на небе. Необычные графика и захватывающий сюжет сделают ваше путешествие по воздуху неповторимым.
Получайте крупные джекпоты с игрой Aviator Spribe играть с друзьями онлайн казино в нашем казино!
Aviator игра позволит вам почувствовать себя настоящим пилотом. Вам предстоит совершить невероятные маневры, выполнять сложные задания и сражаться с противниками. Улучшайте свой самолет, чтобы быть готовым к любым ситуациям и становиться настоящим мастером.
Основные особенности Aviator краш игры:
1. Реалистичная графика и физика – благодаря передовой графике и реалистичной физике вы почувствуете себя настоящим пилотом.
2. Разнообразные режимы игры и миссии – в Aviator краш игре вы сможете выбрать различные режимы игры, такие как гонки, симулятор полетов и захватывающие воздушные бои. Кроме того, каждая миссия будет предлагать свои собственные вызовы и задачи.
3. Улучшение и модернизация самолетов – в игре доступны различные модели самолетов, которые можно покупать и улучшать. Вы сможете устанавливать новое оборудование, улучшать двигательность и мощность своего самолета, а также выбирать различные варианты окраски и декорации.
Aviator краш игра – это возможность испытать себя в роли авиатора и преодолеть все сложности и опасности воздушного пространства. Почувствуйте настоящую свободу и адреналин в Aviator краш игре онлайн!
Играйте в «Авиатор» в онлайн-казино Pin-Up
Aviator краш игра онлайн предлагает увлекательную и захватывающую игровую атмосферу, где вы становитесь настоящим авиатором и сражаетесь с самыми опасными искусственными интеллектами.
В этой игре вы должны показать свое мастерство и смекалку, чтобы преодолеть сложности многочисленных локаций и уровней. Вам предстоит собирать бонусы, уклоняться от препятствий и сражаться с врагами, используя свои навыки пилотирования и стрельбы.
Каждый уровень игры Aviator краш имеет свою уникальную атмосферу и задачи. Будьте готовы к неожиданностям, так как вас ждут захватывающие повороты сюжета и сложные испытания. Найдите все пути к победе и станьте настоящим героем авиатором!
Авиатор игра является прекрасным способом провести время и испытать настоящий адреналиновый разряд. Готовы ли вы стать лучшим авиатором? Не упустите свой шанс и начните играть в Aviator краш прямо сейчас!
Aviator – играй, сражайся, побеждай!
Aviator Pin Up (Авиатор Пин Ап ) – игра на деньги онлайн Казахстан
Aviator игра предлагает увлекательное и захватывающее разнообразие врагов и уровней, которые не оставят равнодушными даже самых требовательных геймеров.
Враги в Aviator краш игре онлайн представлены в самых разных формах и размерах. Здесь вы встретите группы из маленьких и быстрых врагов, а также огромных боссов с мощным вооружением. Разнообразие врагов позволяет игрокам использовать разные тактики и стратегии для победы.
Кроме того, Aviator игра предлагает разнообразие уровней сложности. Выберите легкий уровень, чтобы насладиться игровым процессом, или вызовите себе настоящий вызов, выбрав экспертный уровень. Независимо от выбранного уровня сложности, вы получите максимум удовольствия от игры и окунетесь в захватывающий мир авиаторов.
Играйте в Aviator и наслаждайтесь разнообразием врагов и уровней, которые позволят вам почувствовать себя настоящим авиатором.
I’m not sure exactly why but this blog is loading very slow for me.
Is anyone else having this issue or is it a problem on my end?
I’ll check back later on and see if the problem still
exists.
This is my first time visit at here and i am in fact happy
to read everthing at alone place.
Excellent post but I was wanting to know if you could write a litte more on this subject?
I’d be very grateful if you could elaborate a little bit further.
Kudos!
Hello just wanted to give you a quick heads up.
The words in your content seem to be running off the screen in Internet explorer.
I’m not sure if this is a formatting issue or something to do with browser compatibility
but I thought I’d post to let you know. The design look great
though! Hope you get the problem solved soon. Kudos
saungsantoso.com
이로 인해 Fang Jifan은 후유증이 없을 것이라고 매우 걱정했습니다.이것은 프란시스코 경을 매우 흥분하게 만들었습니다.
https://alepz.com/
Please let me know if you’re looking for a article author for your site.
You have some really great articles and I believe I would be
a good asset. If you ever want to take some
of the load off, I’d really like to write some material for your
blog in exchange for a link back to mine. Please blast me an email if interested.
Thanks!
It’s an amazing paragraph designed for all the internet visitors;
they will obtain advantage from it I am sure.
Heya i am for the first time here. I found this board
and I in finding It really helpful & it helped me out a lot.
I am hoping to present one thing back and help others like you aided me.
Hello there! This article couldn’t be written much better!
Looking at this post reminds me of my previous roommate!
He always kept talking about this. I will send this information to him.
Fairly certain he will have a very good read. I appreciate you for
sharing!
다양한 화폐로 지원되는 프라그마틱 슬롯은 세계 각지의 플레이어에게 열린 기회를 제공합니다.
프라그마틱 게임
프라그마틱 관련 글 읽는 것이 정말 즐거웠어요! 또한, 제 사이트에서도 프라그마틱과 관련된 정보를 공유하고 있어요. 함께 이야기 나누면서 더 많은 지식을 쌓아가요!
https://www.genericzoloft.site
https://www.foxcommconnect.com
https://www.qq8778ok.com
Hello, i read your blog occasionally and i own a similar one and i was just curious if you get a lot of spam feedback?
If so how do you prevent it, any plugin or anything you can advise?
I get so much lately it’s driving me crazy so any support is very
much appreciated.
Do you have a spam problem on this site; I also am a blogger, and I was wanting to know
your situation; many of us have developed some nice
practices and we are looking to swap strategies with other folks, please shoot
me an email if interested.
I every time emailed this blog post page to all my associates, since if like to read it then my contacts will too.
I am truly pleased to glance at this weblog posts which
carries tons of valuable data, thanks for providing these information.
obviously like your web site but you have to check the spelling on quite a few of
your posts. A number of them are rife with spelling problems and
I to find it very bothersome to tell the reality however I’ll surely come again again.
I truly love your website.. Very nice colors &
theme. Did you develop this amazing site yourself?
Please reply back as I’m looking to create my own website and would love to
know where you got this from or just what the theme
is named. Many thanks!
This website was… how do I say it? Relevant!!
Finally I have found something that helped me. Thanks a lot!
Hi i am kavin, its my first time to commenting anywhere, when i read this paragraph i thought i could also create
comment due to this brilliant post.
Thanks , I’ve recently been searching for information approximately this subject for a long time and yours
is the best I’ve discovered till now. However, what concerning the bottom line?
Are you sure about the supply?
Wow, that’s what I was exploring for, what a information! present here at this
webpage, thanks admin of this website.
Zeolite Heavy Equipment LLC offers a wide selection of industrial equipment for various applications. We provide reliable solutions to meet the needs of our customers, ensuring their satisfaction with our products and services. Our company is committed to excellence and continuous improvement in everything we do.
Zahry Machinery Equipment Zeolite Heavy Equipment LLC zeolite heavy equipment llc zahry machinery equipment llc zeolite heavy equipment llc Zahry Machinery Equipment zeolite heavy equipment llc ZAHRY MACHINERY EQUIPMENT LLC Zeolite Heavy Equipment LLC zeolite heavy equipment llc 34_9df1
Thanks very interesting blog!
Wonderful beat ! I would like to apprentice whilst you amend your site,
how can i subscribe for a weblog site? The account helped me a acceptable deal.
I have been tiny bit familiar of this your broadcast offered vivid transparent
idea
I know this if off topic but I’m looking into starting my own weblog and was curious what all is required to
get setup? I’m assuming having a blog like yours would cost a pretty penny?
I’m not very web smart so I’m not 100% sure.
Any suggestions or advice would be greatly appreciated.
Appreciate it
I like the valuable information you provide in your articles.
I will bookmark your blog and check again here regularly.
I’m quite sure I’ll learn many new stuff right here!
Good luck for the next!
Hello are using WordPress for your site platform? I’m new to the blog
world but I’m trying to get started and create my own.
Do you need any coding expertise to make your own blog?
Any help would be greatly appreciated!
It’s the best time to make some plans for the longer term
and it is time to be happy. I’ve read this post and if I
may I desire to recommend you some fascinating things
or advice. Perhaps you can write next articles regarding this article.
I wish to read more things approximately it!
Hi there, I enjoy reading all of your article post. I like to write
a little comment to support you.
magnificent issues altogether, you simply won a emblem new reader.
What might you suggest in regards to your submit that you just made a few days in the past?
Any certain?
Appreciating the persistence you put into your blog and detailed
information you offer. It’s good to come across a blog every once in a while that isn’t
the same unwanted rehashed information. Wonderful read!
I’ve bookmarked your site and I’m including your RSS feeds
to my Google account.
What’s up colleagues, its impressive article about cultureand entirely defined,
keep it up all the time.
Way cool! Some extremely valid points! I appreciate you writing
this post and also the rest of the site is also really good.
lfchungary.com
“당신은 누구를 초대합니까?” Zhu Houzhao는 약간 말문이 막혀 Fang Jifan을 바라 보았습니다.
I’ve been browsing online greater than 3 hours
today, but I never found any fascinating article like yours.
It’s pretty worth enough for me. In my opinion, if all site
owners and bloggers made good content material as you probably did, the internet will likely be much more helpful than ever before.
Hi, I do think this is a great web site. I stumbledupon it ;
) I may come back yet again since i have book marked it. Money and freedom is the greatest
way to change, may you be rich and continue to help other people.
Why visitors still use to read news papers when in this technological
world everything is available on web?
Baccarat is a simple guessing game played with 8 decks of cards. To win, players must make the right call on whether the Player Hand and the Banker Hand will have a score closer to 9 points.
https://psbloansin59minustes.com/
Can I just say what a comfort to find someone that genuinely understands what they are discussing on the web.
You certainly understand how to bring an issue to light and make it important.
More and more people must look at this and understand this side of your story.
I was surprised you are not more popular since you surely have the gift.
Good day! I know this is kinda off topic but I was wondering if you knew where I could locate a captcha plugin for my comment form?
I’m using the same blog platform as yours and I’m having problems finding
one? Thanks a lot!
Remarkable things here. I am very happy to peer your article.
Thanks so much and I am taking a look forward to touch you.
Will you please drop me a e-mail?
Hey I know this is off topic but I was wondering if you knew of any widgets
I could add to my blog that automatically tweet my newest twitter updates.
I’ve been looking for a plug-in like this for quite some time and was hoping maybe you would have some experience with something like
this. Please let me know if you run into anything.
I truly enjoy reading your blog and I look forward to your new updates.
An intriguing discussion is definitely worth comment.
I do believe that you ought to write more on this subject, it might not be a taboo matter but typically folks don’t speak about such
topics. To the next! All the best!!
Hey would you mind letting me know which web host you’re working
with? I’ve loaded your blog in 3 completely different internet
browsers and I must say this blog loads a lot faster then most.
Can you recommend a good internet hosting provider at a honest price?
Thanks, I appreciate it!
What i don’t understood is if truth be told how you are not actually much more smartly-appreciated
than you may be right now. You’re so intelligent.
You understand thus considerably in the case of this subject,
made me in my view imagine it from so many numerous angles.
Its like men and women are not fascinated except it is something to accomplish with Lady gaga!
Your own stuffs great. All the time deal with it up!
Hey just wanted to give you a quick heads up. The words
in your article seem to be running off the screen in Internet explorer.
I’m not sure if this is a formatting issue or something to do with internet browser
compatibility but I figured I’d post to let you know.
The style and design look great though! Hope you get the problem resolved soon.
Many thanks
I constantly spent my half an hour to read this webpage’s
content every day along with a cup of coffee.
Hello there! I just want to give you a huge thumbs up for your
excellent information you’ve got here on this post. I’ll be returning to your
web site for more soon.
It’s an remarkable paragraph for all the internet visitors; they
will obtain advantage from it I am sure.
Hi there! This is my first visit to your blog!
We are a team of volunteers and starting a new project in a community in the
same niche. Your blog provided us beneficial information to
work on. You have done a outstanding job!
I’m not that much of a online reader to be honest but your sites really nice, keep it up!
I’ll go ahead and bookmark your website to come back later.
Cheers
I was wondering if you ever thought of changing the structure of your blog?
Its very well written; I love what youve got to say.
But maybe you could a little more in the way of content so people could connect with it better.
Youve got an awful lot of text for only having one or two pictures.
Maybe you could space it out better?
whoah this blog is fantastic i love reading your posts.
Stay up the good work! You understand, lots of people are searching
around for this information, you can aid them greatly.
Hello there, You have done an incredible job. I’ll certainly digg it and personally suggest to my friends.
I’m confident they’ll be benefited from this web site.
Hi, There’s no doubt that your web site could be
having internet browser compatibility problems.
Whenever I look at your site in Safari, it looks fine however, if opening in IE, it’s got some overlapping issues.
I merely wanted to provide you with a quick
heads up! Apart from that, excellent website!
Please let me know if you’re looking for a writer for your weblog.
You have some really great articles and I feel I would be
a good asset. If you ever want to take some of the load off, I’d love to write
some articles for your blog in exchange for a link back to mine.
Please shoot me an e-mail if interested. Regards!
Hello, i believe that i noticed you visited my blog so i came to return the prefer?.I am trying to
to find issues to enhance my website!I assume its good enough to use a few of your ideas!!
Can I simply just say what a relief to find somebody who
genuinely understands what they’re talking about online.
You actually understand how to bring a problem to light and make it
important. More people need to look at this and understand this side of the story.
I can’t believe you are not more popular given that you definitely possess the gift.
Magnificent site. Plenty of helpful information here. I am sending it to a few pals ans also sharing
in delicious. And obviously, thanks for your effort!
Right here is the perfect site for everyone who really
wants to understand this topic. You know a whole lot its almost
hard to argue with you (not that I really would
want to…HaHa). You definitely put a brand new spin on a
subject which has been discussed for years. Wonderful stuff, just wonderful!
Its like you read my mind! You seem to know a lot about this, like you wrote the book
in it or something. I think that you could do with a few pics to drive the message home a bit, but instead of
that, this is fantastic blog. A great read.
I’ll certainly be back.
Have you ever thought about creating an ebook or guest authoring
on other blogs? I have a blog based on the same subjects you discuss and would love to have you share some stories/information. I know
my audience would appreciate your work. If you’re even remotely interested, feel free to
send me an e-mail.
binsunvipp.com
최초의 타블로이드는 모두 구매자와 판매자를 위한 것이었습니다.
https://hollybollyfun.com/
It’s hard to come by experienced people about
this subject, but you seem like you know what you’re talking about!
Thanks
Excellent pieces. Keep posting such kind of information on your
blog. Im really impressed by it.
Hello there, You’ve performed an excellent job. I’ll definitely digg it and
in my view suggest to my friends. I’m confident they’ll be benefited from this
web site.
Way cool! Some very valid points! I appreciate you penning this
post and the rest of the site is really good.
Keep on writing, great job!
lfchungary.com
그러나 … 방금 Zhu Houzhao의 공연은 Hongzhi 황제를 만족시키지 못했습니다.
Hi there would you mind letting me know which webhost you’re utilizing?
I’ve loaded your blog in 3 different browsers and I must say this blog
loads a lot faster then most. Can you recommend a good internet hosting provider at a reasonable price?
Thanks, I appreciate it!
I was able to find good info from your blog posts.
With havin so much written content do you ever run into any problems
of plagorism or copyright infringement? My website has a
lot of exclusive content I’ve either created myself or outsourced but it appears a lot of it is popping it up all over the internet without my permission. Do you know any
techniques to help stop content from being stolen? I’d really appreciate it.
I absolutely love your website.. Very nice colors & theme.
Did you develop this amazing site yourself? Please reply back as I’m looking to create my own blog and would like
to find out where you got this from or just what the theme is
called. Cheers!
Howdy exceptional website! Does running a blog similar to this require a
massive amount work? I’ve no understanding of
programming but I was hoping to start my own blog soon. Anyway,
should you have any suggestions or techniques for new
blog owners please share. I know this is off topic but I simply needed to ask.
Many thanks!
Hey there! This is my first comment here so I just wanted to give a quick shout out and say I truly
enjoy reading your posts. Can you suggest any
other blogs/websites/forums that go over the same topics? Many thanks!
I think this is one of the such a lot vital information for me.
And i’m satisfied studying your article. But should commentary on few general issues, The web site taste is perfect, the articles is truly excellent : D.
Just right task, cheers
This text is worth everyone’s attention. When can I find out more?
I’m not that much of a internet reader to be honest but your blogs really
nice, keep it up! I’ll go ahead and bookmark your
site to come back later. Cheers
Excellent post. I was checking constantly this blog
and I am impressed! Very helpful information particularly the last
part 🙂 I care for such information much. I was seeking this particular info
for a long time. Thank you and good luck.
Normally I don’t read article on blogs, but I wish
to say that this write-up very pressured me to check out and do so!
Your writing style has been amazed me. Thank you, very nice article.
Buy elite quality proxies – Fully confidential ELITE private proxies with TOP degree of security only from https://DreamProxies.com
Great post.
Having read this I thought it was very enlightening.
I appreciate you taking the time and energy to put this
information together. I once again find myself
personally spending way too much time both reading and leaving
comments. But so what, it was still worthwhile!
Highly energetic blog, I liked that bit. Will there be a part 2?
Wow! This blog looks exactly like my old one! It’s on a entirely different subject but it has pretty much the same layout and design. Outstanding choice of colors!
Every year Casino Player Magazine conducts an extensive survey of the best casinos in our nation. This year is the 25th anniversary edition of the magazine’s Best Of Gaming Awards. … Read more Five Indian casinos named in Top 10 Best Casinos Outside of Las Vegas September 25, 2020 –
https://psbloansin59minustes.com/
You really make it seem so easy with your presentation but I find this topic to be really something that I think I would
never understand. It seems too complicated and extremely broad for me.
I am looking forward for your next post, I’ll try to get the hang of it!
Wow, awesome blog structure! How lengthy have you been running
a blog for? you made running a blog look easy. The whole look
of your website is great, as well as the content material!
I enjoy what you guys are up too. This sort of clever work and reporting!
Keep up the very good works guys I’ve added you guys to our
blogroll.
Hi there! This is my 1st comment here so I just wanted to give a quick shout out
and say I genuinely enjoy reading your articles. Can you recommend any other blogs/websites/forums that go over the same subjects?
Many thanks!
you’re actually a just right webmaster. The web site loading
pace is amazing. It kind of feels that you are doing any distinctive
trick. Moreover, The contents are masterwork. you’ve done a great job in this subject!
agenbet88score.com
Xiao Jing은 안개 속에서 “아마도 무슨 일이 일어날 것입니다. “라고 말했습니다.
https://image.google.ki/url?q=https%3A%2F%2Fwww.colorful-navi.com%2F
Highly energetic post, I liked that a lot. Will there be a part 2?
It’s an awesome paragraph in favor of all the web users; they will get benefit from it I am sure.
This is really fascinating, You’re a very professional blogger.
I have joined your feed and sit up for in the hunt for
extra of your fantastic post. Additionally, I’ve shared your web site in my social networks
Excellent post. Keep writing such kind of info on your
blog. Im really impressed by your blog.
Hi there, You’ve performed an excellent job. I’ll certainly digg it and personally suggest to my friends.
I am sure they’ll be benefited from this web site.
What’s up to all, how is the whole thing, I think every one is getting more from this website, and your
views are pleasant in favor of new viewers.
Hi, I do think this is an excellent blog. I stumbledupon it 😉
I will revisit yet again since i have bookmarked it.
Money and freedom is the best way to change, may you be rich and continue to help other people.
I’ve learn several just right stuff here. Certainly price bookmarking
for revisiting. I surprise how a lot effort you put to create one of these fantastic informative site.
Hi to all, how is everything, I think every one is getting more from this web site, and your views are pleasant for new people.
Hiya! Quick question that’s totally off topic. Do you know how to make your site
mobile friendly? My site looks weird when viewing from my iphone4.
I’m trying to find a template or plugin that might be able to correct this
problem. If you have any recommendations, please share.
Appreciate it!
If you are going for most excellent contents like I do, only go to see this site every day for the reason that it offers feature contents, thanks
WOW just what I was looking for. Came here by searching
for slot mpo
Wow, superb weblog layout! How long have you been blogging for?
you made blogging glance easy. The entire look of your site is magnificent, as well as the
content!
Saved as a favorite, I like your site!
I’m not that much of a online reader to be honest but your blogs
really nice, keep it up! I’ll go ahead and bookmark your website
to come back down the road. Many thanks
Heya i’m for the first time here. I found this board and I find It
really useful & it helped me out a lot. I hope to give something back and help others
like you helped me.
Great article! That is the kind of information that are meant to be shared around the web.
Shame on Google for no longer positioning this publish upper!
Come on over and consult with my site . Thanks =)
Thank you for every other informative site. Where else may just I get that
kind of information written in such an ideal means? I have a venture
that I am simply now operating on, and I’ve been on the glance out for
such info.
Wonderful beat ! I wish to apprentice whilst you amend your
website, how can i subscribe for a weblog site? The account helped me a applicable deal.
I were a little bit acquainted of this your broadcast offered bright clear
idea
Fabulous, what a weblog it is! This webpage presents valuable information to us,
keep it up.
We stumbled over here by a different website and thought I might
check things out. I like what I see so i am just following you.
Look forward to finding out about your web page repeatedly.
What’s Going down i am new to this, I stumbled upon this I have discovered It absolutely helpful and it has aided me out
loads. I’m hoping to give a contribution & aid other customers like its
aided me. Great job.
I read this post completely regarding the comparison of hottest and previous technologies, it’s remarkable article.
Sweet blog! I found it while browsing on Yahoo News. Do you have any tips on how to get listed in Yahoo News?
I’ve been trying for a while but I never seem to get there!
Cheers
What a data of un-ambiguity and preserveness of precious experience on the topic of unpredicted emotions.
Wonderful blog! I found it while browsing on Yahoo News.
Do you have any tips on how to get listed in Yahoo News? I’ve been trying
for a while but I never seem to get there! Many thanks
Nice respond in return of this matter with firm arguments and describing everything concerning that.
excellent post, very informative. I wonder why the opposite experts of this sector don’t realize this.
You must continue your writing. I am sure, you’ve a great readers’ base already!
An outstanding share! I’ve just forwarded this onto a coworker
who has been conducting a little research on this. And he in fact
ordered me lunch due to the fact that I discovered it for him…
lol. So allow me to reword this…. Thanks for the meal!!
But yeah, thanks for spending some time to discuss this matter
here on your web page.
I’m not that much of a internet reader to be honest
but your sites really nice, keep it up! I’ll go ahead and bookmark
your website to come back later on. Cheers
WOW just what I was searching for. Came here by searching for PURAVIVE REVIEWS
It’s hard to find educated people for this topic, but you
sound like you know what you’re talking about!
Thanks
Hello there! Do you use Twitter? I’d like to follow you if that would be okay.
I’m undoubtedly enjoying your blog and look forward to new posts.
I was able to find good info from your blog articles.
Wonderful beat ! I would like to apprentice at the same time as you amend your website, how could i subscribe for a blog site?
The account aided me a acceptable deal. I were tiny bit familiar of this your broadcast provided vivid
clear concept
Good post however , I was wondering if you could write a litte more on this topic?
I’d be very grateful if you could elaborate a little bit more.
Thank you!
An impressive share! I have just forwarded this onto a colleague who has been conducting
a little homework on this. And he in fact ordered me breakfast due to the fact that I found it for him…
lol. So allow me to reword this…. Thanks for the meal!!
But yeah, thanks for spending some time to talk about this
subject here on your web page.
선도적인 iGaming 콘텐츠 제공 업체인 최신 프라그마틱 게임은 혁신적이고 표준화된 콘텐츠를 통해 슬롯, 라이브 카지노, 빙고 등 다양한 제품을 고객에게 제공합니다.
프라그마틱 슬롯
프라그마틱 슬롯에 대한 설명 정말 감사합니다! 더불어, 제 사이트에서도 프라그마틱과 관련된 내용을 찾아보세요. 함께 발전하며 더 많은 지식을 얻어가요!
https://www.polvonuestro.com
https://www.texashillco.com
https://www.dinotri.com
Aw, this was a really good post. Spending some time
and actual effort to produce a really good article… but
what can I say… I procrastinate a whole lot and don’t manage to get anything done.
Its like you read my mind! You appear to know a lot about this, like you
wrote the book in it or something. I think that you could do with a few
pics to drive the message home a bit, but instead of that, this is wonderful blog.
A fantastic read. I’ll definitely be back.
I like the valuable information you provide in your articles.
I’ll bookmark your blog and check again here regularly.
I’m quite sure I’ll learn lots of new stuff right
here! Best of luck for the next!
Have you ever thought about adding a little bit more than just your articles?
I mean, what you say is fundamental and all.
But just imagine if you added some great graphics or videos to give
your posts more, “pop”! Your content is excellent but with pics and videos, this site could undeniably be one of the greatest in its field.
Amazing blog!
I needed to thank you for this fantastic read!!
I definitely enjoyed every bit of it. I’ve got you bookmarked to
check out new things you post…
Hello, I wish for to subscribe for this webpage to obtain most up-to-date updates, therefore
where can i do it please assist.
I’m not sure exactly why but this website is loading very slow
for me. Is anyone else having this issue or is it a problem on my end?
I’ll check back later on and see if the problem still exists.
Hi would you mind sharing which blog platform you’re working
with? I’m planning to start my own blog soon but I’m having a difficult time making a decision between BlogEngine/Wordpress/B2evolution and Drupal.
The reason I ask is because your design seems different then most blogs and I’m looking for something completely unique.
P.S Sorry for being off-topic but I had to ask!
When some one searches for his required thing,
therefore he/she wants to be available that in detail,
so that thing is maintained over here.
Every weekend i used to visit this site, because i want enjoyment, since this this web page conations actually pleasant funny stuff too.
I enjoy, lead to I discovered just what I was having a look
for. You have ended my 4 day lengthy hunt! God Bless you man. Have a great day.
Bye
Hi there! Do you know if they make any plugins to assist with Search Engine Optimization? I’m trying to get my blog
to rank for some targeted keywords but I’m not seeing very good gains.
If you know of any please share. Appreciate it!
Heya are using WordPress for your blog platform?
I’m new to the blog world but I’m trying to get started and create my own. Do you require any html coding
expertise to make your own blog? Any help would be really appreciated!
Hi there it’s me, I am also visiting this web page regularly, this web page is actually good and the users are
truly sharing nice thoughts.
Thanks for the marvelous posting! I actually enjoyed reading it,
you happen to be a great author.I will remember to bookmark your blog and will eventually come back down the road.
I want to encourage you to ultimately continue your great job, have
a nice weekend!
Мы предоставляем услуги Строительство загородного дома под Ключ в Алматы, обеспечивая полный цикл работ от проектирования до завершения строительства. Наша команда опытных специалистов гарантирует высокое качество строительства и индивидуальный подход к каждому клиенту. Работаем с современными технологиями и материалами, чтобы создать дом вашей мечты в соответствии с вашими потребностями и ожиданиями.
continuously i used to read smaller content that as well clear their motive, and that is also happening with this
article which I am reading at this time.
I fоr all time emаiled this weblog post
page to all my contacts, since іf like to read it аftеr that my contacts will toⲟ.
It’s going to be finish of mine day, but before end I am reading this great piece of writing to increase
my experience.
Kudos, A lot of stuff!
I’m amazed, I must say. Rarely do I encounter a blog that’s equally educative and entertaining, and
without a doubt, you have hit the nail on the head. The issue is something not enough men and women are speaking intelligently about.
I am very happy that I came across this in my hunt for something concerning
this.
Piece of writing writing is also a excitement, if you be acquainted with afterward
you can write if not it is complex to write.
My spouse and I absolutely love your blog and find almost all
of your post’s to be just what I’m looking for.
Would you offer guest writers to write content available for you?
I wouldn’t mind writing a post or elaborating on a lot of the subjects you write with regards
to here. Again, awesome weblog!
Now I am going away to do my breakfast, once having my breakfast coming over again to read more news.
At this time I am going away to do my breakfast, afterward having my
breakfast coming again to read more news.
Great info. Lucky me I ran across your website by accident (stumbleupon).
I’ve saved it for later!
Informative article, totally what I was looking for.
Hello, i believe that i saw you visited my site thus i
came to return the want?.I’m trying to to find issues
to enhance my web site!I guess its adequate to make use of a few of your ideas!!
Link exchange is nothing else except it is just placing the other person’s weblog
link on your page at suitable place and other person will also do similar
in favor of you.
Link exchange is nothing else but it is just placing the other person’s website
link on your page at proper place and other person will also
do similar in support of you.
Outstanding story there. What happened after? Thanks!
This design is spectacular! You most certainly know how
to keep a reader amused. Between your wit and your videos, I
was almost moved to start my own blog (well, almost…HaHa!) Great job.
I really enjoyed what you had to say, and more than that, how you presented
it. Too cool!
Hello I am so thrilled I found your web site, I really found you by accident, while I was
researching on Aol for something else, Regardless I am here
now and would just like to say thank you for a fantastic post and a all round thrilling blog (I also love the
theme/design), I don’t have time to browse it all at the minute but I have saved it and also
included your RSS feeds, so when I have time I will be back
to read more, Please do keep up the superb jo.
Good post. I learn something totally new and challenging on websites I stumbleupon every day.
It’s always interesting to read through content from other authors and use something from their websites.
Thanks for sharing your thoughts. I really appreciate your efforts and I am waiting for your next write ups thank you once again.
My page site#:
https://seoforum.bestff.ru/viewtopic.php?id=2122#p5328
https://zarabotok.liveforums.ru/viewtopic.php?id=12502#p33414
http://vkmonline.com/blogs/post/1489018
https://xn--80aehnrb3a4b.xn--p1ai/blogs/vseznaika/srednee-obrazovanie-tendentsii-i-perspektivy-v-rossii-i-zarubezhom-v-2.php
https://komfort.rusff.me/viewtopic.php?id=11474#p35674
Hello colleagues, fastidious paragraph and fastidious arguments commented here,
I am genuinely enjoying by these.
Does your blog have a contact page? I’m having a tough time locating it but, I’d like to shoot you an email.
I’ve got some suggestions for your blog you might be interested in hearing.
Either way, great website and I look forward to seeing it grow over time.
Hello there, just became alert to your blog through Google, and
found that it’s really informative. I am gonna watch out for brussels.
I will be grateful if you continue this in future.
Many people will be benefited from your writing. Cheers!
Hello there I am so delighted I found your weblog, I really found you by mistake, while I was researching on Yahoo for something else, Regardless I am here now and would just like to say thanks for a marvelous post and a all round interesting blog (I also love the theme/design), I don’t have time to go through it all at the minute but I have bookmarked it and also added in your RSS feeds, so when I have time I will be back to read a lot more, Please do keep up the fantastic job.
My#page#site#:
https://dolgoprudni.rusff.me/viewtopic.php?id=1828#p4321
http://diablomania.ru/forum/showthread.php?p=556676#post556676
https://slingomama74.bbeasy.ru/viewtopic.php?id=3923#p9390
https://earnings.0pk.me/viewtopic.php?id=9109#p18691
https://zarabotok.liveforums.ru/viewtopic.php?id=12494#p33392
I believe everything published was actually very logical.
However, think on this, what if you composed a catchier post title?
I ain’t saying your information is not solid., however what if you added something
that makes people want more? I mean Fornix (crus)/ Stria Terminalis –
fxst | Whole Brain Protocol for Tractography with Empirical MRI
is a little plain. You could look at Yahoo’s front page
and note how they create news titles to grab people to click.
You might try adding a video or a picture or two to get readers excited about what you’ve
got to say. In my opinion, it could make your blog a little livelier.
Good post. I learn something totally new and challenging on blogs I
stumbleupon every day. It will always be useful to read through articles from other
authors and use something from their sites.
Admiring the commitment you put into your website and detailed information you offer.
It’s good to come across a blog every once in a while that isn’t the
same outdated rehashed material. Fantastic read! I’ve bookmarked
your site and I’m adding your RSS feeds to my Google account.
I used to be able to find good advice from your content.
http://borderforum.ru/viewtopic.php?f=26&t=8451
http://gamerscf.forum-top.ru/viewtopic.php?id=236#p421
http://cn.rolevaya.ru/viewtopic.php?id=5811#p18387
http://forr.forum2.net/viewtopic.php?id=2868#p8021
http://nada25.mybb.ru/viewtopic.php?id=2185#p27896
https://womens-forum.ru/viewtopic.php?f=59&t=5636&sid=551f4b0930cf627c6cbe727676e17dfd
Hi there, You have done a fantastic job. I’ll certainly digg it and personally recommend to my friends. I am confident they’ll be benefited from this website.
http://borderforum.ru/viewtopic.php?f=26&t=8451
http://raiter.flyboard.ru/viewtopic.php?f=9&t=1834
https://izhevsk.ru/forummessage/153/6113648.html
http://tumgerl.rolbb.me/viewtopic.php?id=11998#p18541
http://www.pets-ural.ru/content/forum/viewtopic.php?p=56636#56636
http://kissolovo.teamforum.ru/viewtopic.php?f=10&t=32866
Keep on writing, great job!
Amazing blog! Do you have any recommendations for aspiring writers?
I’m planning to start my own blog soon but I’m a little lost on everything.
Would you advise starting with a free platform like WordPress or
go for a paid option? There are so many choices out there that
I’m completely overwhelmed .. Any ideas? Kudos!
There’s certainly a great deal to learn about this subject.
I really like all of the points you made.
Wow that was unusual. I just wrote an very long comment but after I clicked submit my comment didn’t appear.
Grrrr… well I’m not writing all that over again. Anyway, just wanted to say fantastic blog!
Its like you read my mind! You seem to know so much
about this, like you wrote the book in it or something.
I think that you can do with some pics to drive the message home a
little bit, but instead of that, this is great blog.
An excellent read. I will certainly be back.
It’s very simple to find out any topic on web as compared to books, as I found this paragraph
at this site.
Have you ever considered writing an e-book or guest authoring on other websites?
I have a blog based upon on the same information you discuss and would love
to have you share some stories/information. I know my visitors would value your
work. If you are even remotely interested, feel free to send
me an e-mail.
Howdy! This post could not be written any better!
Reading through this post reminds me of my good old room mate!
He always kept talking about this. I will forward this write-up to him.
Fairly certain he will have a good read. Thanks for sharing!
A fascinating discussion is definitely worth comment. I do
think that you ought to publish more about this issue, it
may not be a taboo matter but usually people don’t talk about these topics.
To the next! Best wishes!!
Hi there, after reading this awesome piece of writing i am as well cheerful to share my knowledge here with mates.
https://premiumy-dlplomsy24.com/
http://www.openstreetmap.org/user/arkadypettr
http://www.brschool16.ru/communication/forum/forum3/topic4511/
http://transexit.g-talk.ru/viewtopic.php?f=10&t=1261&p=1861
http://ulanude.bezformata.com/listnews/obyavlyaetsya-konkursnij-otbor-po-podderzhke/23808534/
http://irsolo.ru/cru-rossijskij-avialajner-nad-sinaem-vzorvala-gruppirovka-ansar-bejt-al-makdis/index.html
Hello, i think that i saw you visited my website thus i came to “return the favor”.I’m trying to find things to improve my
web site!I suppose its ok to use some of your ideas!!
I’m really enjoying the design and layout of your blog.
It’s a very easy on the eyes which makes it much more enjoyable
for me to come here and visit more often. Did you hire out a developer to create your theme?
Fantastic work!
Hi there everyone, it’s my first visit at this web site,
and post is in fact fruitful for me, keep up posting these articles or
reviews.
I needed to thank you for this very good read!!
I definitely loved every bit of it. I have got you
book marked to look at new things you post…
magnificent issues altogether, you just won a emblem new
reader. What might you suggest about your publish that you simply
made a few days in the past? Any certain?
Fantastic blog! Do you have any recommendations for aspiring writers?
I’m hoping to start my own blog soon but I’m a little lost on everything.
Would you suggest starting with a free platform like WordPress or go for a paid option? There
are so many options out there that I’m totally confused ..
Any ideas? Kudos!
Hi, I check your blog on a regular basis. Your writing style is witty, keep doing what
you’re doing!
I am really loving the theme/design of your website.
Do you ever run into any web browser compatibility issues?
A handful of my blog visitors have complained about my blog not operating correctly
in Explorer but looks great in Safari. Do you have any suggestions to help fix this problem?
Excellent post however I was wanting to know if you could write
a litte more on this topic? I’d be very thankful if you could elaborate a
little bit more. Bless you!
Why users still use to read news papers when in this technological globe the whole thing is accessible on net?
I am actually glad to glance at this weblog posts which carries
tons of valuable information, thanks for providing these data.
I seriously love your site.. Great colors & theme.
Did you create this amazing site yourself? Please reply back as I’m trying to create my very own blog and
want to learn where you got this from or just what the theme is
called. Kudos!
Piece of writing writing is also a fun, if you be familiar with afterward you can write if not it is complicated to
write.
It’s hard to find well-informed people in this particular topic, however, you sound like you know what you’re talking about! Thanks
Whats up very cool web site!! Guy .. Beautiful .. Amazing ..
I will bookmark your website and take the feeds additionally?
I’m satisfied to seek out numerous helpful information right here in the submit, we want work out
extra techniques in this regard, thanks for sharing.
. . . . .
Thanks , I have just been looking for info about this subject for a
while and yours is the best I’ve discovered so far. However, what concerning
the bottom line? Are you positive about the source?
I was able to find good advice from your blog posts.
I don’t even know how I ended up here, but I thought this post was good.
I do not know who you are but certainly you’re going to a famous blogger if you aren’t already 😉
Cheers!
This web site really has all of the information I needed about this subject and didn’t know who to ask.
A motivating discussion is definitely worth comment.
I think that you should publish more on this subject, it may not be a taboo subject but usually folks
don’t discuss these issues. To the next! Kind regards!!
Hi, Neat post. There is a problem along with your
site in web explorer, could test this? IE nonetheless is the marketplace chief
and a good part of folks will pass over your magnificent writing due to this problem.
I do not even know how I ended up here,
but I thought this post was good. I don’t know who you are but certainly you are going to a famous blogger
if you aren’t already 😉 Cheers!
My family members all the time say that I am killing my time here at
net, but I know I am getting familiarity everyday by
reading thes pleasant posts.
A person necessarily help to make severely posts
I might state. This is the first time I frequented your website page and up to now?
I amazed with the research you made to make this particular put
up extraordinary. Magnificent activity!
I all the time used to study piece of writing in news papers but now
as I am a user of net so from now I am using net for posts, thanks
to web.
Hello it’s me, I am also visiting this site daily,
this site is genuinely fastidious and the users are truly sharing
pleasant thoughts.
Hi my loved one! I want to say that this article is awesome,
great written and include almost all vital infos.
I’d like to look more posts like this .
If you wish for to grow your know-how just keep visiting this website
and be updated with the most recent gossip posted here.
I think everything published was actually very reasonable.
But, consider this, suppose you typed a catchier
post title? I mean, I don’t want to tell you how to run your website, but what if you added something to maybe get people’s
attention? I mean Fornix (crus)/ Stria Terminalis – fxst | Whole
Brain Protocol for Tractography with Empirical MRI is a little plain. You might peek at
Yahoo’s home page and watch how they create post headlines to
grab viewers to click. You might try adding a video or
a related picture or two to grab readers interested about what you’ve written. Just my opinion, it
might make your website a little bit more interesting.
Hey would you mind sharing which blog platform you’re using?
I’m planning to start my own blog in the near future but I’m having a difficult time choosing between BlogEngine/Wordpress/B2evolution and Drupal.
The reason I ask is because your design seems different
then most blogs and I’m looking for something completely unique.
P.S Apologies for getting off-topic but I had to
ask!
Hello, i think that i saw you visited my weblog thus i came to “return the favor”.I am trying to find things to enhance
my web site!I suppose its ok to use some of your ideas!!
Hi, Neat post. There’s an issue along with your site in web explorer, could check this?
IE still is the market leader and a large element of other people will omit your excellent writing
due to this problem.
First off I want to say superb blog! I had a quick question that I’d
like to ask if you do not mind. I was interested to know how you center yourself and clear your head prior to writing.
I have had a tough time clearing my thoughts in getting my thoughts out there.
I do enjoy writing however it just seems like the first 10 to 15 minutes are generally lost just trying to figure out how to begin. Any ideas or tips?
Cheers!
Greetings from Colorado! I’m bored to tears at work so I
decided to browse your website on my iphone during lunch break.
I really like the information you provide here and can’t wait to take a look
when I get home. I’m amazed at how fast your blog
loaded on my mobile .. I’m not even using WIFI, just 3G
.. Anyhow, good blog!
I’ll immediately grab your rss as I can not in finding
your e-mail subscription hyperlink or e-newsletter service.
Do you’ve any? Kindly let me understand so that I could subscribe.
Thanks.
Unquestionably consider that which you said. Your favourite justification appeared to be at the
net the simplest thing to take note of. I say to you, I
certainly get annoyed while people consider issues that they plainly don’t recognize about.
You managed to hit the nail upon the top and also outlined out the entire thing with no need side effect , other people
could take a signal. Will probably be again to get more. Thank you
С Vegas Grand ваша мечта о выигрыше станет реальностью. Переходите на https://vigrat.ru и погрузитесь в захватывающий мир азартных игр прямо сейчас!
Link exchange is nothing else however it is just placing the other person’s webpage link on your page at proper place and other person will also do same in favor of you.
Terrific post but I was wanting to know if you could write a litte more on this topic?
I’d be very thankful if you could elaborate a little bit further.
Cheers!
Thanks for sharing your thoughts about https://techcommunity.microsoft.com/t5/user/viewprofilepage/user-id/2346899#profile. Regards
Your style is very unique in comparison to other people I have read stuff from.
Many thanks for posting when you’ve got the opportunity, Guess I will just bookmark this
page.
I figured out more new stuff on this fat loss issue. 1 issue is that good nutrition is tremendously vital if dieting. A huge reduction in fast foods, sugary food items, fried foods, sweet foods, pork, and white flour products may be necessary. Possessing wastes unwanted organisms, and wastes may prevent aims for losing weight. While certain drugs for the short term solve the situation, the awful side effects are not worth it, and in addition they never supply more than a non permanent solution. It is just a known idea that 95 of dietary fads fail. Many thanks for sharing your ideas on this weblog.
hi!,I love your writing very much! proportion we communicate
more approximately your article on AOL? I need a specialist in this area
to resolve my problem. Maybe that is you! Looking ahead to peer you.
Asking questions are actually nice thing if you are not understanding something totally, however this post gives fastidious understanding yet.
Hi there terrific website! Does running a blog similar to this take
a large amount of work? I have virtually
no expertise in computer programming however I was hoping
to start my own blog in the near future. Anyhow, if
you have any ideas or techniques for new blog owners please share.
I know this is off topic nevertheless I just had to ask.
Thanks a lot!
Write more, thats all I have to say. Literally, it
seems as though you relied on the video to make your point.
You obviously know what youre talking about, why waste your
intelligence on just posting videos to your blog when you could be giving us something informative to read?
I’m curious how creative writing instructors at colleges and universities handle students who write about really disturbing things and who seem potentially dangerous to themselves and others? Are instructors privy to students’ mental health records? Do they let such students get away with violent or disturbing writing in an effort NOT to stir too much trouble? Do you become proactive in trying to help these students? Do you undergo training to deal with problem students? As a creative writing student at a university, I often see disturbing stuff brought into workshops. I’m wondering what the profs think of all this. Thanks to any answers!.
Its like you read my mind! You appear to know so much
about this, like you wrote the book in it or something.
I think that you could do with some pics to drive the message home a little
bit, but other than that, this is fantastic blog.
An excellent read. I will certainly be back.
Hello there, I discovered your web site by means of Google whilst searching for a similar subject, your web site got here
up, it seems good. I have bookmarked it in my google bookmarks.
Hi there, simply became alert to your weblog through Google, and found that it’s really informative.
I am gonna watch out for brussels. I’ll appreciate in the event you
continue this in future. Numerous other people will be benefited from your writing.
Cheers!
Thank you for every other informative website.
The place else may I get that type of info written in such an ideal way?
I have a project that I’m just now operating on, and I’ve been on the look
out for such info.
If you are going for most excellent contents like myself, simply go to see this web page all
the time since it gives feature contents, thanks
I absolutely love your blog and find almost all of your post’s to be what
precisely I’m looking for. Does one offer guest writers to write
content for you? I wouldn’t mind producing a post or elaborating on a number of the subjects you write in relation to here.
Again, awesome website!
If some one wishes to be updated with most up-to-date
technologies therefore he must be pay a visit this
web site and be up to date everyday.
This information is worth everyone’s attention. When can I find out more?
Thanks for sharing your thoughts. I truly appreciate your
efforts and I am waiting for your next write ups thanks once again.
I don’t even know how I ended up here, but I thought this post was great.
I don’t know who you are but certainly you are going to a famous blogger if
you aren’t already 😉 Cheers!
apksuccess.com
선발된 대부분의 장인들은 선조로부터 물려받은 유산을 가지고 있습니다.
I just like the helpful info you provide on your articles.
I’ll bookmark your weblog and take a look at once more here regularly.
I’m reasonably sure I will be told many new stuff right here!
Best of luck for the following!
This is a topic that is close to my heart… Best wishes! Where are your contact details though?
Hello, all is going perfectly here and ofcourse every one is sharing data, that’s actually excellent,
keep up writing.
Hey! I could have sworn I’ve been to this
website before but after browsing through some of the post I realized it’s new to me.
Nonetheless, I’m definitely delighted I found it and I’ll be bookmarking and checking back frequently!
Link exchange is nothing else except it is just placing the other person’s website link on your page at proper place and other person will also do same for you.
You really make it seem so easy with your presentation but I find this
topic to be actually something that I think I would never understand.
It seems too complex and extremely broad for me. I am looking
forward for your next post, I will try to get the hang of it!
Keep this going please, great job!
I really like what you guys are usually up too.
This sort of clever work and coverage! Keep up the terrific works guys I’ve added you guys to my personal blogroll.
It is actually a great and helpful piece of information. I’m satisfied that
you shared this useful info with us. Please stay us up to date like
this. Thanks for sharing.
Everyone loves it when individuals come together and share thoughts.
Great website, stick with it!
Hi, I do believe this is a great site. I stumbledupon it 😉 I may return yet again since
i have book marked it. Money and freedom is the best
way to change, may you be rich and continue to guide
others.
Hello. Great job. I did not imagine this. This is a impressive story. Thanks!
Greetings! Very helpful advice within this post! It is the little changes that
will make the most important changes. Thanks a lot for sharing!
hihouse420.com
다행스럽게도 Shen 가족은 타이틀이 있고 타이틀이 있으면 50 년 동안 빌릴 수 있으므로 천천히 갚자.
Hi! I know this is somewhat off topic but I was wondering which blog platform are you using for this site?
I’m getting tired of WordPress because I’ve had issues with
hackers and I’m looking at alternatives for another platform.
I would be fantastic if you could point me in the direction of a good platform.
Hi to every one, because I am actually eager of reading this web site’s post to be updated daily.
It consists of fastidious stuff.
Hello there, You’ve done a fantastic job. I will definitely digg it and personally recommend to
my friends. I am sure they’ll be benefited from this website.
프라그마틱은 항상 훌륭한 게임을 만들어냅니다. 이번에 새롭게 출시된 게임은 정말 기대되는데요!
PG 소프트
프라그마틱 슬롯에 대한 내용이 정말 도움이 되었어요! 더불어, 제 사이트에서도 프라그마틱과 관련된 정보를 찾아보실 수 있어요. 함께 지식을 공유해보세요!
https://e-trajet.net/hot/
https://pdlwqpd.weebly.com/
https://njshidoo.com/hot/
Fascinating blog! Is your theme custom made or did you download it from somewhere?
A theme like yours with a few simple adjustements
would really make my blog jump out. Please let me know where you got your design. Thanks
Yes! Finally something about best time to visit egypt.
Hi there! I just wanted to ask if you ever have any problems with hackers?
My last blog (wordpress) was hacked and I ended up losing several weeks of hard work due to no backup.
Do you have any solutions to prevent hackers?
I all the time used to read post in news papers but
now as I am a user of internet thus from now I am using net for articles, thanks to web.
Excellent post. I was checking constantly this weblog and I
am impressed! Very helpful information specially the remaining part :
) I maintain such information much. I used to be seeking this certain info for a long time.
Thank you and good luck.
I’m no longer positive the place you’re getting your info, however good topic.
I needs to spend some time finding out much more
or working out more. Thanks for great info I was looking
for this information for my mission.
This article gives clear idea for the new people of blogging,
that really how to do blogging and site-building.
Today, while I was at work, my sister stole my apple ipad and tested
to see if it can survive a twenty five foot drop, just so she can be a youtube
sensation. My iPad is now broken and she has 83
views. I know this is entirely off topic but I had to share it with someone!
If you want to grow your know-how only keep visiting this site and be updated with the newest news update posted here.
Its like you learn my thoughts! You appear to understand so much about this, such as you wrote the book
in it or something. I think that you just could do with some % to power the message home
a little bit, however other than that, that is great blog.
An excellent read. I’ll definitely be back.
Excellent beat ! I would like to apprentice while you amend your site, how can i
subscribe for a blog site? The account aided me a acceptable deal.
I had been tiny bit acquainted of this your broadcast offered bright clear concept
I have read so many content about the blogger lovers however this article is genuinely a
fastidious paragraph, keep it up.
I am extremely impressed with your writing skills and also with the layout on your weblog.
Is this a paid theme or did you customize it yourself?
Anyway keep up the excellent quality writing, it is rare to see a great blog like this one today.
I’ve really noticed that credit score improvement activity needs to be conducted with techniques. If not, it’s possible you’ll find yourself destroying your rating. In order to grow into success fixing your credit history you have to confirm that from this time you pay your monthly dues promptly before their planned date. It is significant given that by never accomplishing this, all other methods that you will take to improve your credit rating will not be useful. Thanks for sharing your ideas.
Thanks for your suggestions. One thing we have noticed is the fact that banks along with financial institutions know the dimensions and spending habits of consumers while also understand that many people max outside their real credit cards around the getaways. They wisely take advantage of this kind of fact and begin flooding your current inbox plus snail-mail box together with hundreds of no-interest APR credit cards offers shortly after the holiday season concludes. Knowing that if you’re like 98 of American public, you’ll rush at the one opportunity to consolidate financial debt and transfer balances for 0 apr interest rates credit cards.
Great article, just what I wanted to find.
Having read this I thought it was extremely informative.
I appreciate you finding the time and energy to put this information together.
I once again find myself spending way too much time both reading and posting comments.
But so what, it was still worth it!
It’s hard to find educated people in this particular subject, however,
you seem like you know what you’re talking about!
Thanks
Thanks for sharing your thoughts on lombaqq. Regards
Thank you for the auspicious writeup. It in fact was a amusement
account it. Look advanced to more added agreeable from you!
By the way, how can we communicate?
Hello, I enjoy reading through your post. I like to write a
little comment to support you.
Wow that was unusual. I just wrote an extremely long comment but after I clicked submit my
comment didn’t appear. Grrrr… well I’m not writing all that over again. Anyhow, just wanted to say great blog!
Do you mind if I quote a few of your articles as long
as I provide credit and sources back to your webpage? My blog is in the very same area of interest as yours
and my visitors would really benefit from some of the information you present here.
Please let me know if this okay with you. Regards!
One other thing is that an online business administration training is designed for individuals to be able to well proceed to bachelor’s degree programs. The Ninety credit education meets the lower bachelor college degree requirements so when you earn your current associate of arts in BA online, you will possess access to the modern technologies on this field. Several reasons why students want to get their associate degree in business is because they’re interested in the field and want to have the general schooling necessary just before jumping right into a bachelor diploma program. Thx for the tips you provide as part of your blog.
Hi, after reading this amazing paragraph i am as well happy to share my know-how here with colleagues.
If you are going for finest contents like me, just
visit this site every day because it offers quality contents, thanks
Having read this I believed it was very informative.
I appreciate you taking the time and effort to put this short article together.
I once again find myself personally spending way too much time both reading and posting comments.
But so what, it was still worthwhile!
Howdy would you mind stating which blog platform you’re working with?
I’m going to start my own blog in the near future but I’m having a tough time choosing
between BlogEngine/Wordpress/B2evolution and
Drupal. The reason I ask is because your design and style seems different then most blogs and I’m looking for
something unique. P.S Sorry for
being off-topic but I had to ask!
Hey! I know this is somewhat off topic but I was wondering which blog
platform are you using for this site? I’m getting tired of WordPress because I’ve had problems
with hackers and I’m looking at alternatives for another
platform. I would be awesome if you could point me in the direction of
a good platform.
Awesome! Its really awesome paragraph, I have got much clear idea about
from this article.
Thanks for finally writing about > Fornix (crus)/ Stria
Terminalis – fxst | Whole Brain Protocol for Tractography with Empirical MRI
< Liked it!
Whats up very cool blog!! Guy .. Beautiful .. Amazing ..
I will bookmark your site and take the feeds additionally?
I am happy to find a lot of useful information right here within the submit, we want develop extra techniques in this regard, thank you
for sharing. . . . . .
It’s fantastic that you are getting thoughts from this post as well as from our discussion made at this time.
Hi there! This article could not be written any better!
Reading through this article reminds me of my previous roommate!
He continually kept talking about this. I’ll send this information to him.
Pretty sure he’ll have a good read. I appreciate you for sharing!
Heya are using WordPress for your blog platform?
I’m new to the blog world but I’m trying to get started and create
my own. Do you require any coding knowledge to make your own blog?
Any help would be greatly appreciated!
My programmer is trying to persuade me to move to .net from PHP.
I have always disliked the idea because of the costs.
But he’s tryiong none the less. I’ve been using WordPress on a variety of websites for about a
year and am anxious about switching to another platform.
I have heard very good things about blogengine.net.
Is there a way I can import all my wordpress content into it?
Any help would be greatly appreciated!
Hello there! I could have sworn I’ve been to this
site before but after browsing through some of the post I realized it’s new to me.
Anyhow, I’m definitely happy I found it and I’ll be bookmarking and checking back often!
fantastic issues altogether, you simply gained a brand new reader.
What may you suggest in regards to your publish that you
simply made some days ago? Any sure?
I know this if off topic but I’m looking into starting my own blog and
was wondering what all is required to get set up? I’m assuming having a blog like yours would cost
a pretty penny? I’m not very internet smart so I’m not 100% certain. Any suggestions or advice would be greatly
appreciated. Kudos
May I simply just say what a comfort to discover a
person that actually understands what they’re talking about on the web.
You actually realize how to bring a problem
to light and make it important. A lot more people need
to check this out and understand this side of the story.
It’s surprising you are not more popular because you
definitely have the gift.
Hello, Neat post. There’s a problem with your site in internet explorer, would check
this? IE still is the market chief and a large component of people will leave out your fantastic writing
because of this problem.
Hi there! This article could not be written much better!
Going through this article reminds me of my previous roommate!
He continually kept preaching about this. I will forward this information to him.
Pretty sure he’ll have a good read. Many thanks for sharing!
Piece of writing writing is also a excitement, if you be familiar with afterward you can write or else it is complicated to
write.
strelkaproject.com
그때부터 사람들이 모여들어 서로 소리를 질렀다.
It’s awesome in favor of me to have a website, which
is valuable in favor of my know-how. thanks admin
Simply wish to say your article is as surprising. The clarity in your publish is just spectacular and that i could assume
you’re a professional on this subject. Fine together with your permission let me to snatch your feed to keep up to date with
impending post. Thank you a million and please keep up
the gratifying work.
I am truly delighted to glance at this website posts which contains plenty of helpful information,
thanks for providing these data.
Have you ever considered publishing an e-book or guest authoring on other blogs?
I have a blog centered on the same ideas you discuss and
would love to have you share some stories/information.
I know my audience would enjoy your work. If you’re even remotely interested, feel free to send me an email.
My partner and I stumbled over here by a
different web page and thought I might as well check things
out. I like what I see so now i am following you. Look forward to exploring your
web page for a second time.
It is perfect time to make some plans for the longer term and it’s time to be happy.
I have learn this post and if I could I want to recommend you some fascinating things or suggestions.
Perhaps you could write subsequent articles relating
to this article. I wish to read even more things about it!
shopanho.com
진구지안의 창고에는 옷감이 짜여져 쌓여 있다.
I was wondering if you ever considered changing the layout of your site?
Its very well written; I love what youve got to say. But maybe you could a little more in the way of content so
people could connect with it better. Youve got an awful lot of text for only having
one or 2 pictures. Maybe you could space it out better?
I was suggested this blog by means of my cousin. I’m no longer sure whether or not this
publish is written by him as no one else understand such distinctive approximately my trouble.
You’re wonderful! Thank you!
Wow, wonderful blog structure! How long have you been running a blog for?
you make blogging glance easy. The full glance of your website is excellent,
as smartly as the content material!
https://gitlab.gnome.org/taisunwinbet
Great post.
These are genuinely impressive ideas in on the topic of blogging.
You have touched some pleasant factors here.
Any way keep up wrinting.
Thanks , I’ve recently been looking for info about this
subject for a long time and yours is the greatest I’ve came upon so far.
But, what concerning the conclusion? Are you sure about
the supply?
I all the time used to read piece of writing in news papers but now
as I am a user of web so from now I am using net for content, thanks
to web.
Hello, of course this article is genuinely pleasant and I have learned lot of things from it concerning blogging.
thanks.
Thank you for every other wonderful post. The place
else could anybody get that type of info in such an ideal manner
of writing? I’ve a presentation subsequent week, and I
am on the look for such information.
Hey I know this is off topic but I was wondering if you knew of any widgets
I could add to my blog that automatically tweet my newest twitter updates.
I’ve been looking for a plug-in like this for quite some time and was hoping maybe you would have
some experience with something like this. Please let me
know if you run into anything. I truly enjoy reading your blog and
I look forward to your new updates.
I like the valuable info you provide in your articles. I will bookmark your blog and check again here regularly. I am quite sure I will learn many new stuff right here! Good luck for the next!
It’s a shame you don’t have a donate button! I’d certainly donate to this superb blog! I guess for now i’ll settle for bookmarking and adding your RSS feed to my Google account. I look forward to brand new updates and will share this site with my Facebook group. Talk soon!
Admiring the persistence you put into your blog and detailed information you offer.
It’s good to come across a blog every once in a while that isn’t
the same out of date rehashed material. Fantastic
read! I’ve bookmarked your site and I’m adding your RSS feeds to
my Google account.
I think this is among the most important info for me. And
i am glad reading your article. But should remark on few general things,
The website style is ideal, the articles is really great : D.
Good job, cheers
I think this is one of the most vital info
for me. And i am glad reading your article. But wanna remark on some general things,
The web site style is ideal, the articles is really excellent :
D. Good job, cheers
Valuable info. Fortunate me I discovered your web site accidentally,
and I’m stunned why this coincidence did not took place earlier!
I bookmarked it.
Hi! I know this is kinda off topic but I was wondering which blog platform are
you using for this website? I’m getting fed up of WordPress because I’ve had
problems with hackers and I’m looking at options for another platform.
I would be awesome if you could point me in the
direction of a good platform.
Hello to every single one, it’s in fact a fastidious for me to pay a quick visit this web page,
it contains important Information.
I am extremely impressed with your writing skills as well
as with the layout on your blog. Is this a paid theme or did
you customize it yourself? Either way keep up the nice
quality writing, it’s rare to see a nice blog like this one today.
excellent put up, very informative. I wonder why the opposite specialists
of this sector don’t realize this. You should proceed your
writing. I am confident, you have a great readers’ base already!
Hi there colleagues, how is the whole thing, and what you would like to say on the topic of this
paragraph, in my view its genuinely awesome for me.
Saved as a favorite, I love your blog!
Heya i am for the first time here. I found this board and I
find It really useful & it helped me out a lot. I hope to
give something back and aid others like you helped me.
I am regular visitor, how are you everybody? This article posted at this site is in fact pleasant.
Hey there just wanted to give you a quick heads up and let you know a few of the pictures aren’t loading properly.
I’m not sure why but I think its a linking issue. I’ve tried it in two different web
browsers and both show the same outcome.
I think that is one of the so much vital info for me. And i am satisfied studying
your article. However should statement on few basic issues,
The web site taste is ideal, the articles is truly nice
: D. Excellent task, cheers
Appreciate it. Lots of material!
Hmm it seems like your site ate my first comment (it was extremely long) so I guess I’ll just sum
it up what I wrote and say, I’m thoroughly enjoying your blog.
I as well am an aspiring blog blogger but I’m
still new to everything. Do you have any recommendations for inexperienced
blog writers? I’d definitely appreciate it.
It’s not my first time to pay a visit this website, i am browsing this web site dailly and get fastidious information from here daily.
bookmarked!!, I really like your blog!
This is my first time pay a visit at here and i am in fact pleassant to read everthing at single
place.
Why users still use to read news papers when in this technological world everything is available on web?
Hi there friends, how is all, and what you desire to say regarding this post,
in my view its truly awesome for me.
Howdy! Someone in my Myspace group shared this site with us so I came to give it a look.
I’m definitely loving the information. I’m book-marking and will be
tweeting this to my followers! Fantastic blog and excellent
design.
If you want to improve your knowledge just keep visiting this site and be updated with the most recent gossip
posted here.
parrotsav.com
그러나 일단 이것이 축적되면 많은 구리 부족이 발생할 것입니다.
A fascinating discussion is definitely worth comment.
There’s no doubt that that you ought to write more about this topic, it
might not be a taboo matter but usually people do not talk about such subjects.
To the next! Best wishes!!
My wife and i were very ecstatic Ervin managed to finish up his analysis with the precious recommendations he grabbed while using the web pages. It is now and again perplexing to simply find yourself handing out techniques the others have been making money from. And we do understand we need the website owner to thank for that. The explanations you’ve made, the simple web site menu, the friendships you can help create – it is all superb, and it is leading our son in addition to our family do think that matter is entertaining, which is certainly wonderfully serious. Thanks for the whole lot!
It’s the best time to make some plans for the future and
it is time to be happy. I have read this post and if I
could I want to suggest you some interesting things or suggestions.
Perhaps you can write next articles referring to this article.
I desire to read more things about it!
I just couldn’t leave your web site before suggesting that I really loved the
usual information an individual provide for your guests?
Is going to be back ceaselessly in order to check up on new posts
It is the best time to make some plans for the future and it’s time to be happy.
I’ve read this post and if I could I wish to suggest you
few interesting things or tips. Perhaps you could write next articles referring
to this article. I want to read more things about it!
Hey! I know this is somewhat off-topic but I had
to ask. Does operating a well-established website such as yours require a massive amount work?
I am completely new to operating a blog however I do write in my diary
everyday. I’d like to start a blog so I can easily share my personal experience and views online.
Please let me know if you have any recommendations or tips for brand new aspiring blog owners.
Appreciate it!
When someone writes an paragraph he/she retains the
thought of a user in his/her brain that how a user can be aware of it.
Thus that’s why this post is outstdanding. Thanks!
I have been browsing online more than three hours as of late, yet I never discovered any interesting article like yours.
It’s lovely worth sufficient for me. Personally, if all site owners and bloggers made just right content as
you probably did, the net can be much more helpful than ever before.
Way cool! Some extremely valid points! I appreciate you penning this post plus the rest of the site is extremely good.
Hi just wanted to give you a quick heads up and let you know a few of the pictures
aren’t loading properly. I’m not sure why but I think its a
linking issue. I’ve tried it in two different internet browsers and
both show the same results.
Hi there, after reading this amazing article i am also glad to share my know-how
here with friends.
whoah this weblog is wonderful i like reading your posts.
Keep up the great work! You understand, a lot of persons are looking around for this info, you can help them greatly.
I am no longer sure where you’re getting your info, however good topic.
I must spend some time finding out much more or working out more.
Thank you for excellent information I used to be looking
for this info for my mission.
Hi my friend! I want to say that this post is amazing, great written and come with almost all vital infos.
I would like to see more posts like this .
You made some good points there. I looked on the internet for the subject matter and found most persons will approve with your blog.
Howdy! Do you know if they make any plugins to help with SEO? I’m trying to get my blog to rank for some targeted keywords but I’m not seeing very good gains. If you know of any please share. Many thanks!
Excellent post. I certainly love this website. Keep it up!
Keep this going please, great job!
Everything is very open with a very clear description of the challenges.
It was truly informative. Your site is extremely helpful.
Thanks for sharing!
What a stuff of un-ambiguity and preserveness
of valuable familiarity regarding unexpected feelings.
Undeniably consider that which you stated.
Your favourite justification seemed to be at
the internet the easiest factor to remember of.
I say to you, I certainly get annoyed even as other people
consider concerns that they just don’t know about. You managed
to hit the nail upon the highest and outlined out the
entire thing without having side effect , folks could take a signal.
Will probably be back to get more. Thank you
Hi there i am kavin, its my first occasion to commenting anyplace, when i read this paragraph i thought i could also create comment due to this good post.
I quite like reading a post that will make people think. Also,
thank you for allowing me to comment!
Hi! Would you mind if I share your blog with my zynga
group? There’s a lot of people that I think would really appreciate your content.
Please let me know. Many thanks
hello!,I love your writing so a lot! proportion we keep in touch extra approximately
your post on AOL? I need an expert on this house to unravel my
problem. Maybe that is you! Taking a look forward to see you.
What’s up to every , as I am genuinely eager of reading this webpage’s post to be updated regularly.
It includes fastidious stuff.
Hmm it looks like your website ate my first comment (it was extremely long) so
I guess I’ll just sum it up what I submitted and say,
I’m thoroughly enjoying your blog. I as well
am an aspiring blog writer but I’m still new to
everything. Do you have any suggestions for novice blog writers?
I’d definitely appreciate it.
Hey I know this is off topic but I was wondering if you knew of any widgets I could add to my blog
that automatically tweet my newest twitter updates.
I’ve been looking for a plug-in like this for quite some time and was hoping maybe you would have some experience with something like this.
Please let me know if you run into anything.
I truly enjoy reading your blog and I look forward to your new updates.
Hi there, just became aware of your blog through Google, and found that it’s really informative.
I’m gonna watch out for brussels. I will be grateful
if you continue this in future. Many people will
be benefited from your writing. Cheers!
Greetings! Very helpful advice in this particular post!
It is the little changes that produce the greatest
changes. Thanks a lot for sharing!
hello there and thank you for your information – I’ve
definitely picked up something new from right here.
I did however expertise some technical issues using this
site, as I experienced to reload the website a lot of times previous to I could get it
to load correctly. I had been wondering if your hosting is OK?
Not that I am complaining, but sluggish loading instances
times will sometimes affect your placement in google and could damage your high-quality score if ads and marketing
with Adwords. Anyway I am adding this RSS to my e-mail
and could look out for a lot more of your respective
intriguing content. Make sure you update this again soon.
Hi there to all, the contents present at this site
are really awesome for people experience, well, keep up the nice work fellows.
Excellent blog here! Also your site loads up very fast! What web host
are you using? Can I get your affiliate link to your host?
I wish my web site loaded up as quickly as yours lol
My brother suggested I may like this blog. He was totally right.
This publish actually made my day. You cann’t believe just how a lot time I had spent for this information! Thanks!
Great post. I was checking continuously this blog and I’m impressed!
Very helpful info specially the last part 🙂 I care for such info a lot.
I was looking for this particular information for a
long time. Thank you and best of luck.
My relatives every time say that I am killing my time here at net, but I know I am getting familiarity all the
time by reading such nice content.
Nice post. I used to be checking continuously this weblog and
I am impressed! Extremely helpful information particularly the closing phase 🙂 I care
for such info a lot. I was looking for this particular
information for a very long time. Thanks and good luck.
Hi there! This is my 1st comment here so I
just wanted to give a quick shout out and tell you I genuinely enjoy reading
your posts. Can you suggest any other blogs/websites/forums that cover the
same topics? Thank you so much!
Everything is very open with a very clear clarification of the
challenges. It was definitely informative. Your website is useful.
Many thanks for sharing!
Today, I went to the beachfront with my kids. I found a sea shell and gave it to my 4 year old daughter and said “You can hear the ocean if you put this to your ear.” She
put the shell to her ear and screamed. There was a hermit crab inside and it pinched her ear.
She never wants to go back! LoL I know this is totally off topic but I had to tell
someone!
Hi there to every body, it’s my first visit of this web site; this webpage consists of remarkable and actually fine material in support of readers.
http://dukeanddusty.biz/__media__/js/netsoltrademark.php?d=hottelecom.biz/hi/
delta 8 moonrocks review
Hello, the whole thing is going fine here and ofcourse every one is sharing information, that’s actually excellent, keep up writing.
http://maps.google.bs/url?q=https://hottelecom.biz/id/
WOW just what I was searching for. Came here by searching
for growth matrix scam
Having read this I thought it was extremely informative.
I appreciate you spending some time and energy to put this short article
together. I once again find myself personally spending way too much time both reading and posting comments.
But so what, it was still worth it!
Wow, that’s what I was looking for, what a material!
existing here at this webpage, thanks admin of this site.
wonderful publish, very informative. I wonder why the other specialists of this sector do not understand this.
You must continue your writing. I’m sure, you’ve a huge readers’ base already!
Hello Dear, are you genuinely visiting this website regularly, if so after that you will definitely take fastidious knowledge.
We stumbled over here by a different web page and thought I should check things out.
I like what I see so now i am following you. Look forward
to looking into your web page for a second time.
It is not my first time to pay a visit this site, i am visiting this
site dailly and get good data from here everyday.
Thanks a lot for sharing this with all folks you actually know
what you’re speaking about! Bookmarked. Kindly also seek advice from my website =).
We will have a hyperlink change contract between us
Howdy just wanted to give you a quick heads up.
The text in your post seem to be running off the screen in Safari.
I’m not sure if this is a formatting issue or something to do
with internet browser compatibility but I figured I’d post to let you know.
The design and style look great though! Hope you get the issue fixed soon. Cheers
Very good information. Lucky me I discovered your site by chance (stumbleupon).
I have saved as a favorite for later!
Thanks on your marvelous posting! I seriously enjoyed reading it,
you will be a great author. I will always bookmark
your blog and will come back from now on. I want to encourage you to definitely
continue your great work, have a nice evening!
Hello there! This article could not be written any better! Reading through this article reminds me of my previous roommate! He continually kept preaching about this. I will send this information to him. Pretty sure he will have a great read. I appreciate you for sharing!
Wow, fantastic blog layout! How long have you been blogging for?
you made blogging look easy. The overall look of your web
site is wonderful, let alone the content!
I’m amazed, I have to admit. Seldom do I encounter a blog that’s both equally educative and entertaining,
and let me tell you, you’ve hit the nail on the head.
The issue is something too few men and women are speaking intelligently about.
I’m very happy that I stumbled across this in my hunt for something regarding
this.
I visited many sites except the audio feature for audio songs existing at this web
page is really superb.
Very nice post. I simply stumbled upon your blog and wanted to mention that I’ve truly loved surfing
around your weblog posts. After all I’ll be subscribing on your rss
feed and I’m hoping you write once more soon!
I’m amazed, I must say. Seldom do I encounter a blog that’s both
educative and amusing, and without a doubt, you’ve hit the nail on the head.
The issue is something not enough men and women are speaking intelligently about.
I am very happy that I came across this in my search for something concerning this.
Good post but I was wanting to know if you could write a litte
more on this topic? I’d be very grateful if you could elaborate a little bit more.
Kudos!
Amazing issues here. I’m very happy to see your post.
Thanks a lot and I am taking a look ahead
to contact you. Will you kindly drop me a e-mail?
Excellent post. I am experiencing some of these issues as well..
It’s remarkable to pay a visit this web site and reading the views of all
friends on the topic of this post, while I am also eager of getting knowledge.
smcasino7.com
Fang Jifan은 만족스럽게 생각했습니다. Fang Jifan이 효도하지 않는다고 말하지 마십시오.
When I initially commented I seem to have clicked the -Notify me when new comments are added- checkbox and now whenever
a comment is added I get four emails with the exact same
comment. There has to be an easy method you are able to remove me from that
service? Many thanks!
Hi, I think your blog might be having browser compatibility issues.
When I look at your website in Opera, it looks fine but when opening in Internet Explorer, it has some overlapping.
I just wanted to give you a quick heads up! Other then that, awesome
blog!
Hey there! This is kind of off topic but I need some advice from an established blog.
Is it tough to set up your own blog? I’m not very techincal but I can figure things out pretty
quick. I’m thinking about creating my own but I’m not sure
where to start. Do you have any points or suggestions?
Thank you
It’s difficult to find experienced people for this topic, however, you sound like you
know what you’re talking about! Thanks
Nice blog here! Also your website loads up very fast!
What host are you using? Can I get your affiliate link to your host?
I wish my web site loaded up as quickly as yours lol
I’m not sure why but this blog is loading extremely slow for me.
Is anyone else having this problem or is it a problem on my end?
I’ll check back later and see if the problem still exists.
I’m gone to inform my little brother, that he should also pay a
visit this web site on regular basis to get updated from most
up-to-date gossip.
Attractive part of content. I just stumbled upon your weblog and in accession capital to assert that I get actually enjoyed account your blog posts.
Any way I will be subscribing for your augment or even I success
you get admission to persistently fast.
With havin so much content do you ever run into any problems of
plagorism or copyright violation? My blog has a lot of
exclusive content I’ve either written myself or outsourced but
it appears a lot of it is popping it up all over the internet without my agreement.
Do you know any techniques to help prevent content from
being stolen? I’d truly appreciate it.
m88
Highly descriptive article, I liked that a lot. Will there be a part 2?
Hey there are using WordPress for your blog platform? I’m new to the blog world but I’m trying to get started and set up my own. Do you require any html coding expertise to make your own blog? Any help would be greatly appreciated!
You actually make it seem so easy with your presentation but I find this matter to
be really something that I think I would never understand.
It seems too complicated and very broad for me. I’m looking forward for
your next post, I’ll try to get the hang of it!
Very nice blog post. I certainly love this website.
Thanks!
Thank you a lot for sharing this with all
of us you actually know what you’re talking about! Bookmarked.
Kindly also seek advice from my web site =). We will have a
link trade contract between us
I have to thank you for the efforts you’ve put in penning this website.
I am hoping to check out the same high-grade blog posts from you in the future as well.
In truth, your creative writing abilities has motivated me to get my own blog now
😉
I want to to thank you for this very good read!!
I definitely loved every little bit of it. I’ve
got you bookmarked to look at new things you post…
Helpful info. Fortunate me I found your site unintentionally, and
I am shocked why this coincidence didn’t came about earlier!
I bookmarked it.
You are so awesome! I don’t think I’ve read through anything like this before.
So nice to find another person with a few original thoughts on this issue.
Seriously.. thanks for starting this up.
This web site is one thing that’s needed on the
web, someone with some originality!
I’m not sure where you are getting your info, but good topic.
I must spend some time studying much more or understanding
more. Thanks for wonderful info I used to
be in search of this information for my mission.
Aw, this was an incredibly nice post. Taking the time and actual effort to create a great article… but what can I say… I procrastinate a lot and never
manage to get nearly anything done.
Fabulous, what a webpage it is! This webpage gives helpful data
to us, keep it up.
I do accept as true with all the ideas you’ve offered to your post.
They’re very convincing and will definitely work.
Nonetheless, the posts are too quick for newbies.
May just you please extend them a bit from subsequent time?
Thank you for the post.
Howdy! I’m at work surfing around your blog from my new iphone 4!
Just wanted to say I love reading your blog and look
forward to all your posts! Carry on the superb work!
Hi there! This post could not be written much better! Looking at this article
reminds me of my previous roommate! He continually kept preaching
about this. I will send this post to him. Fairly certain he will have a great read.
Thank you for sharing!
Someone necessarily lend a hand to make severely articles I might state.
This is the very first time I frequented your website page and thus far?
I surprised with the research you made to make this particular post extraordinary.
Wonderful process!
Hey! I know this is somewhat off topic but I was wondering which blog
platform are you using for this site? I’m getting fed up of WordPress because I’ve had problems with hackers and I’m
looking at alternatives for another platform.
I would be awesome if you could point me
in the direction of a good platform.
What i do not realize is in fact how you are not really much
more smartly-preferred than you may be now.
You are so intelligent. You know thus significantly on the
subject of this matter, produced me in my view imagine it
from so many numerous angles. Its like women and men are
not involved except it’s something to accomplish with Girl gaga!
Your personal stuffs nice. At all times deal with it up!
Hi there, its nice piece of writing regarding media
print, we all understand media is a wonderful source of information.
Great site you have got here.. It’s hard to find high quality writing like yours nowadays.
I seriously appreciate people like you! Take care!!
Keep on working, great job!
Definitely believe that which you said. Your favorite reason appeared to be on the internet the simplest thing to be aware of.
I say to you, I definitely get irked while people think about worries
that they just do not know about. You managed to
hit the nail upon the top as well as defined out the whole thing without having
side-effects , people can take a signal. Will probably be back to get more.
Thanks
Whoa! This blog looks exactly like my old one! It’s on a
entirely different subject but it has pretty much the same layout and design. Great choice of colors!
s9vako02
Thanks for the marvelous posting! I really enjoyed reading it, you happen to be a great author.I will remember to bookmark your blog and will come back very soon. I want to encourage yourself to continue your great posts, have a nice weekend!
It’s truly a great and useful piece of info. I’m happy that you shared this helpful information with us.
Please stay us up to date like this. Thanks for sharing.
Tremendous things here. I am very satisfied to look your article.
Thanks so much and I am having a look forward to touch you.
Will you kindly drop me a e-mail?
Amazing! This blog looks just like my old one! It’s on a completely different topic but it has pretty much the same page layout and
design. Great choice of colors!
We’re a group of volunteers and opening a brand new scheme in our community.
Your website provided us with helpful info to work on. You’ve done an impressive
activity and our entire group will likely be thankful to you.
Helpful info. Lucky me I found your website by chance, and I am stunned why this coincidence didn’t happened in advance!
I bookmarked it.
An outstanding share! I’ve just forwarded this onto a coworker who
has been conducting a little homework on this. And he
in fact bought me breakfast due to the fact that I
stumbled upon it for him… lol. So allow me to reword this….
Thank YOU for the meal!! But yeah, thanx for spending time to discuss this subject here on your website.
What’s up, this weekend is fastidious in favor
of me, for the reason that this occasion i am reading this wonderful informative article here at my home.
I do not even know how I ended up here, but I thought this
post was great. I do not know who you are but certainly you are going
to a famous blogger if you aren’t already 😉 Cheers!
An interesting discussion is definitely worth comment.
I think that you need to publish more on this subject matter, it might not be a taboo matter but usually people do not talk about these subjects.
To the next! Best wishes!!
I’m very happy to uncover this great site. I wanted to thank you for your time for this
wonderful read!! I definitely loved every little bit of it and i also have you book marked to see new things in your website.
I’m pretty pleased to uncover this page. I need to to thank you
for ones time just for this fantastic read!! I definitely appreciated every little
bit of it and I have you book-marked to check out new things in your site.
I’ve read several just right stuff here. Certainly price bookmarking
for revisiting. I surprise how much attempt you put to make
this kind of magnificent informative web site.
Great post.
I believe everything composed was very reasonable.
But, what about this? what if you added a
little information? I am not saying your information isn’t solid, however suppose you added a title that makes people desire more?
I mean Fornix (crus)/ Stria Terminalis –
fxst | Whole Brain Protocol for Tractography with Empirical
MRI is kinda plain. You ought to peek at Yahoo’s front page and watch how they
create news headlines to grab people interested.
You might try adding a video or a pic or two to get readers interested about everything’ve written. In my opinion,
it might bring your blog a little livelier.
I am really enjoying the theme/design of your site.
Do you ever run into any web browser compatibility issues?
A few of my blog visitors have complained about my site not operating correctly in Explorer but looks great in Firefox.
Do you have any tips to help fix this problem?
This blog was… how do you say it? Relevant!!
Finally I’ve found something which helped me. Cheers!
Unquestionably believe that which you said. Your favorite justification seemed to be on the internet the simplest thing to be aware of.
I say to you, I definitely get annoyed while people think about
worries that they plainly do not know about.
You managed to hit the nail upon the top as well as defined out the
whole thing without having side-effects , people could take a signal.
Will likely be back to get more. Thanks
Woah! I’m really enjoying the template/theme of this website.
It’s simple, yet effective. A lot of times it’s difficult to get
that “perfect balance” between usability and appearance.
I must say you’ve done a excellent job with this.
Also, the blog loads extremely fast for me on Internet explorer.
Superb Blog!
May I simply just say what a comfort to discover an individual who genuinely knows what they’re discussing on the
internet. You certainly know how to bring a problem to light and make it important.
More people need to look at this and understand this side of your story.
It’s surprising you are not more popular given that you most certainly possess the gift.
I loved as much as you’ll receive carried out
right here. The sketch is attractive, your authored material stylish.
nonetheless, you command get bought an impatience over that you wish be delivering the following.
unwell unquestionably come more formerly again since exactly the same nearly a
lot often inside case you shield this increase.
Asking questions are actually good thing if you are not understanding something entirely, except this paragraph gives
pleasant understanding even.
This blog was… how do I say it? Relevant!! Finally I’ve
found something which helped me. Many thanks!
I have noticed that car insurance companies know the autos which are vulnerable to accidents and various risks. Additionally, these people know what form of cars are given to higher risk and also the higher risk they have got the higher the actual premium rate. Understanding the simple basics with car insurance will assist you to choose the right form of insurance policy that will take care of your needs in case you get involved in any accident. Thanks for sharing the actual ideas on your own blog.
I could not refrain from commenting. Perfectly written!
Hello i am kavin, its my first time to commenting anyplace,
when i read this paragraph i thought i could also create comment due to this brilliant post.
Thanks – Enjoyed this post, can I set it up so I get an email sent to me whenever there is a fresh post?
Hi, I do believe this is an excellent site. I stumbledupon it 😉
I may come back once again since I book-marked it. Money and freedom is the greatest way to change, may you be rich and continue to help others.
Very nice post. I just stumbled upon your weblog and wanted
to mention that I’ve really enjoyed browsing your blog posts.
In any case I’ll be subscribing on your feed and I’m hoping you write once more soon!
There’s definately a great deal to learn about
this topic. I like all of the points you have made.
Very nice post. I just stumbled upon your weblog and
wanted to say that I have truly enjoyed surfing around your
blog posts. After all I will be subscribing to your feed and
I hope you write again soon!
I think this is among the most vital info for me. And i am
glad reading your article. But should remark on few general
things, The site style is perfect, the articles is really great :
D. Good job, cheers
Hello there! This is my first visit to your blog! We are a
collection of volunteers and starting a new project in a community in the
same niche. Your blog provided us useful information to work on. You
have done a marvellous job!
When I initially commented I seem to have clicked on the -Notify me when new comments are added- checkbox and now each time a comment is added I recieve 4 emails with the exact same comment. There has to be a way you can remove me from that service? Kudos!
Terrific article! This is the kind of info that
are supposed to be shared around the net. Shame on Google for not positioning this
put up higher! Come on over and discuss with my site .
Thanks =)
Hello There. I discovered your blog the use of msn. That is a really well written article.
I’ll be sure to bookmark it and return to learn extra of your helpful information. Thanks for
the post. I will certainly return.
Wow, amazing blog structure! How long have you ever been blogging for?
you made blogging glance easy. The entire look of
your site is fantastic, as well as the content
material!
I blog often and I truly appreciate your content.
This article has truly peaked my interest. I will book mark your site and keep
checking for new information about once a week. I
opted in for your RSS feed too.
Hello! I just wanted to ask if you ever have any issues with hackers?
My last blog (wordpress) was hacked and I ended up losing a few
months of hard work due to no data backup. Do you have
any solutions to prevent hackers?
Great article! That is the type of info that are meant to be shared across the net.
Shame on the seek engines for no longer positioning this post
upper! Come on over and seek advice from my web site .
Thanks =)
Hi there, just wanted to say, I loved this
article. It was inspiring. Keep on posting!
I’m not that much of a online reader to be honest but your blogs really nice,
keep it up! I’ll go ahead and bookmark your site to come back later on. Cheers
Hello to all, the contents existing at this web site are
genuinely awesome for people experience, well, keep up the good work fellows.
Oh my goodness! Impressive article dude! Thank you so much, However I am going through difficulties
with your RSS. I don’t know why I can’t subscribe to it.
Is there anybody getting similar RSS problems? Anyone who knows the answer can you kindly respond?
Thanx!!
I think the admin of this site is genuinely working hard in support of his web
site, because here every data is quality based material.
It’s hard to come by knowledgeable people in this particular subject, however,
you seem like you know what you’re talking about! Thanks
Awesome blog! Do you have any hints for aspiring writers?
I’m hoping to start my own blog soon but I’m a little lost on everything.
Would you advise starting with a free platform like WordPress
or go for a paid option? There are so many options out there that I’m totally overwhelmed ..
Any tips? Many thanks!
Howdy! This is my first visit to your blog! We are a collection of volunteers and starting a new project in a
community in the same niche. Your blog provided us beneficial information to work on. You have done a
wonderful job!
This website was… how do I say it? Relevant!! Finally I’ve found something which helped me.
Cheers!
My family members every time say that I am killing my time
here at net, however I know I am getting know-how daily by reading such pleasant content.
My blog – drinking dart board
Every weekend i used to pay a visit this site, as i wish for enjoyment, since this this website conations actually nice funny material too.
Pretty element of content. I simply stumbled upon your weblog and in accession capital to claim that I get in fact loved account your weblog posts. Any way I will be subscribing for your augment and even I achievement you get admission to constantly quickly.
RybelsusRybelsusRybelsusRybelsusRybelsus
Actually no matter if someone doesn’t understand
after that its up to other visitors that they will help, so here it takes place.
This is really fascinating, You’re an overly professional
blogger. I’ve joined your feed and stay up for in quest of more of
your wonderful post. Also, I’ve shared your website in my social networks
I’ve been exploring for a little bit for any high quality articles or blog posts
on this kind of space . Exploring in Yahoo I ultimately stumbled upon this web site.
Studying this information So i am satisfied to convey that I have an incredibly excellent
uncanny feeling I came upon just what I needed. I most no doubt will make sure to do
not fail to remember this website and provides it a glance regularly.
Nice blog! Is your theme custom made or did you download it from somewhere?
A theme like yours with a few simple tweeks would really make my
blog stand out. Please let me know where you
got your theme. Thanks a lot
Saved as a favorite, I really like your web site!
Hi there, i read your blog from time to time and i own a
similar one and i was just wondering if you get a lot
of spam comments? If so how do you protect against it, any plugin or anything
you can advise? I get so much lately it’s driving me mad so any support is very much appreciated.
I saw similar here: Sklep internetowy
I have read so many articles regarding the blogger lovers however this article is genuinely a fastidious
piece of writing, keep it up.
I all the time used to study post in news papers but now as I am a
user of net therefore from now I am using net for posts,
thanks to web.
A person necessarily lend a hand to make seriously posts I might state.
That is the first time I frequented your web page and to this point?
I surprised with the analysis you made to make this particular
publish incredible. Great task!
Ahaa, its fastidious conversation concerning this piece of writing here at this web
site, I have read all that, so now me also commenting here.
Currently it sounds like Movable Type is the preferred blogging platform out there right now. (from what I’ve read) Is that what you’re using on your blog?
Hey very cool website!! Man .. Excellent .. Wonderful ..
I will bookmark your blog and take the feeds also? I’m happy to seek out
so many helpful information here in the submit, we’d like develop
extra techniques in this regard, thanks for sharing.
. . . . .
This is a topic that is close to my heart… Many thanks!
Where are your contact details though?
Greetings from Idaho! I’m bored to death at work so I decided to browse your blog
on my iphone during lunch break. I enjoy the knowledge you
provide here and can’t wait to take a look when I get home.
I’m surprised at how fast your blog loaded on my phone ..
I’m not even using WIFI, just 3G .. Anyways, wonderful blog!
I could not refrain from commenting. Well written!
Hello to every body, it’s my first pay a visit of this webpage; this website consists of amazing and actually excellent stuff designed for visitors.
Greetings I am so happy I found your website, I really found you by error, while I was searching on Yahoo for something else, Nonetheless I am here now and would just like to say many
thanks for a fantastic post and a all round interesting blog (I also love the theme/design), I don’t have time to read through it all at the minute but I have book-marked it and
also added in your RSS feeds, so when I have time I will be
back to read a lot more, Please do keep up
the great b.
We stumbled over here coming from a different web page
and thought I might as well check things out. I like what I see so i
am just following you. Look forward to looking at
your web page yet again.
Hmm it looks like your website ate my first comment (it was extremely long) so I
guess I’ll just sum it up what I wrote and say, I’m thoroughly enjoying your blog.
I as well am an aspiring blog blogger but I’m still new to
everything. Do you have any recommendations for first-time blog writers?
I’d certainly appreciate it.
Amazing blog! Is your theme custom made or did you download it from somewhere?
A theme like yours with a few simple tweeks would really make my blog jump out.
Please let me know where you got your design. With
thanks
What’s Taking place i am new to this, I stumbled upon this
I’ve discovered It absolutely useful and it has helped me out loads.
I’m hoping to contribute & assist different users like
its aided me. Good job.
I am actually grateful to the holder of this website who has shared
this fantastic paragraph at at this time.
Very nice article, just what I was looking for.
If some one desires to be updated with most up-to-date technologies therefore he must
be pay a visit this web page and be up to date daily.
Hey I know this is off topic but I was wondering if you knew of any widgets I could add to my blog that automatically tweet my newest
twitter updates. I’ve been looking for a plug-in like
this for quite some time and was hoping maybe you
would have some experience with something like this.
Please let me know if you run into anything. I truly enjoy reading your blog and I look forward to your new updates.
You actually make it appear so easy along with your presentation however I in finding this matter to be actually
something that I think I would by no means understand.
It kind of feels too complicated and very vast for me. I’m having a look forward
in your subsequent submit, I will try to get the hold of it!
I’ve been browsing online more than three hours nowadays,
yet I by no means found any fascinating article like yours.
It is pretty worth sufficient for me. In my view, if all website owners and bloggers made just right content
material as you probably did, the internet will probably be much more
useful than ever before.
Your means of explaining all in this article is actually nice, every one be capable of easily be aware of it, Thanks
a lot.
It’s an amazing piece of writing designed for all the web visitors;
they will get benefit from it I am sure.
Greetings! I know this is kinda off topic nevertheless I’d figured I’d ask.
Would you be interested in exchanging links or maybe guest writing a
blog article or vice-versa? My site goes over a lot of the same subjects as yours and I
feel we could greatly benefit from each other. If you’re interested feel free to send me an e-mail.
I look forward to hearing from you! Awesome blog by the way!
Good web site you’ve got here.. It’s difficult to find quality writing like yours
these days. I really appreciate individuals like
you! Take care!!
I every time emailed this web site post page
to all my associates, as if like to read it then my contacts will too.
Hello to every one, the contents present at this web page are genuinely remarkable for people knowledge, well, keep up the good work fellows.
Hey there! Someone in my Facebook group shared
this site with us so I came to look it over. I’m definitely enjoying the
information. I’m bookmarking and will be tweeting this
to my followers! Exceptional blog and outstanding design.
When I initially commented I clicked the
“Notify me when new comments are added” checkbox and
now each time a comment is added I get four e-mails with the same comment.
Is there any way you can remove me from that service?
Thanks!
You made some really good points there. I checked on the web for more information about
the issue and found most people will go along with your views on this site.
I’d like to thank you for the efforts you have put
in writing this site. I’m hoping to check out the same high-grade content from you later on as well.
In truth, your creative writing abilities has motivated me to
get my very own site now 😉
Fantastic post however I was wondering if you could write a litte more on this
topic? I’d be very thankful if you could elaborate a little bit further.
Bless you!
Someone necessarily lend a hand to make critically posts I would state.
This is the first time I frequented your website page and so far?
I amazed with the research you made to make this particular put up incredible.
Fantastic activity!
You can certainly see your expertise in the article
you write. The sector hopes for even more passionate writers like you who aren’t afraid to mention how they believe.
At all times go after your heart.
Anti-Forensics is the use of tools and techniques to frustrate a computer examination.
Wonderful blog you have here but I was curious about if you knew of
any user discussion forums that cover the same topics discussed here?
I’d really like to be a part of online community where I can get comments from other experienced people that share the same interest.
If you have any suggestions, please let me know.
Thank you!
Good day! Do you use Twitter? I’d like to follow you if
that would be ok. I’m definitely enjoying your blog and look forward to new updates.
I truly love your website.. Great colors & theme.
Did you build this site yourself? Please reply back as I’m wanting to create my own personal blog and
would like to learn where you got this from or exactly what the theme is
named. Many thanks!
Magnificent beat ! I would like to apprentice while you amend your web site, how can i subscribe
for a blog site? The account aided me a acceptable deal. I had been a little bit acquainted
of this your broadcast offered bright clear idea
First off I want to say fantastic blog! I had a quick question which I’d like to ask if
you do not mind. I was curious to find out how you center yourself and clear your thoughts before writing.
I’ve had a difficult time clearing my mind in getting
my ideas out. I truly do take pleasure in writing however it just seems
like the first 10 to 15 minutes are wasted simply
just trying to figure out how to begin. Any recommendations or hints?
Appreciate it!
It is perfect time to make some plans for the long run and it is
time to be happy. I have read this post and if I may
just I desire to counsel you some fascinating things or
advice. Perhaps you can write next articles referring to this article.
I desire to learn more things approximately it!
WOW just what I was looking for. Came here by searching for inground pool installer
Hey there! Do you know if they make any plugins to protect against hackers?
I’m kinda paranoid about losing everything I’ve worked hard on. Any recommendations?
A fascinating discussion is definitely worth comment.
I believe that you ought to publish more on this
subject, it might not be a taboo matter but generally people don’t discuss such issues.
To the next! Many thanks!!
qiyezp.com
Hongzhi 황제는 음침한 얼굴로 말했다. “나는 그들을보고 싶지 않습니다. 가자, 차에 타십시오.”
Hi there colleagues, how is all, and what you desire to say about this post, in my view its
genuinely remarkable for me.
I think this is among the most vital information for me. And i am glad reading your article.
But wanna remark on few general things, The site
style is wonderful, the articles is really nice : D.
Good job, cheers
of course like your web-site but you need to take a look at the spelling on quite
a few of your posts. Several of them are rife with spelling issues and I find
it very troublesome to tell the reality nevertheless I’ll
surely come again again.
When I originally commented I clicked the “Notify me when new comments are added” checkbox and now each
time a comment is added I get several e-mails with the same comment.
Is there any way you can remove people from that service?
Thank you!
Hello! Do you know if they make any plugins to
assist with Search Engine Optimization? I’m trying to get my blog to rank for some targeted keywords but I’m not seeing very good gains.
If you know of any please share. Many thanks! You can read similar blog here:
Dobry sklep
Fantastic post but I was wanting to know if you could write a litte more on this subject?
I’d be very grateful if you could elaborate a little bit more.
Kudos!
Greetings, I do believe your website could possibly be having browser compatibility problems.
When I take a look at your site in Safari, it looks fine however,
when opening in I.E., it’s got some overlapping issues.
I just wanted to provide you with a quick heads up!
Aside from that, excellent site!
Pretty nice post. I just stumbled upon your weblog and wished to say that I’ve truly enjoyed
surfing around your blog posts. After all I will be subscribing to
your feed and I hope you write again very soon!
We are a gaggle of volunteers and opening a brand new scheme in our community.
Your site provided us with useful information to work on.
You’ve done an impressive task and our entire community might be thankful to you.
Hello, all is going perfectly here and ofcourse every
one is sharing facts, that’s in fact good, keep up writing.
bunn programmable coffee maker
large coffee brewer
welding classes in md
troy bilt pony 42 oil type
professional dart throwing
tropical bonsai trees
prune avocado trees
zinc in dark chocolate
indoor ficus
heat vision glasses
Have a look at my web site … atn viper
It’s very interesting! If you need help, look here: ARA Agency
I pay a visit everyday some websites and sites to read content, however this web site gives quality based writing.
Awesome! Its actually remarkable piece of writing, I have got much clear idea regarding from this post.
my page; tracfone coupon
Hello to all, it’s really a good for me to visit this web page, it
consists of useful Information.
That is very fascinating, You are an excessively skilled blogger.
I’ve joined your rss feed and look ahead to looking for more of your wonderful post.
Additionally, I’ve shared your web site in my social networks
Wow, this piece of writing is good, my younger sister is analyzing these things,
thus I am going to let know her.
Hey there! I could have sworn I’ve been to this site before
but after reading through some of the post I realized it’s new to
me. Nonetheless, I’m definitely delighted I found it and I’ll
be book-marking and checking back often!
My developer is trying to convince me to move to .net from PHP.
I have always disliked the idea because of the costs.
But he’s tryiong none the less. I’ve been using WordPress on a variety of websites for about a year
and am concerned about switching to another platform.
I have heard great things about blogengine.net. Is there a
way I can transfer all my wordpress posts into it? Any help would
be greatly appreciated!
Hello every one, here every one is sharing these kinds of know-how, so it’s fastidious to read this
weblog, and I used to pay a quick visit this webpage every day.
I visited many sites but the audio quality for audio songs present at this site
is in fact excellent.
Ahaa, its good dialogue concerning this article here at this web site, I have read all that,
so at this time me also commenting at this place.
I constantly emailed this website post page to all my associates, since if like to
read it afterward my friends will too.
Hey there great website! Does running a blog such as
this take a lot of work? I’ve no expertise in computer programming but I was
hoping to start my own blog in the near future. Anyhow, if you have any suggestions or techniques for
new blog owners please share. I understand this is off topic
however I just wanted to ask. Thank you!
The meal went without incident although a couple of times she had to avert her gaze from Dan for fear of bursting out laughing. Dan went up to his room saying he wanted to listen to music, he laid on his bed playing with himself waiting for his father to leave. He thought about the events of the last two weeks it had been the craziest time of his life.
He put it all down to Mary Harris, she had been his girlfriend for two years, he liked her a lot, she was pretty and intelligent two things that rarely combined in his experience, the problem was she steadfastly refused to let him put his cock inside her, he had thought he was getting somewhere when he took his cock out that evening in the cinema when she had finally allowed him to get his hand inside her panties, it was the first time she had allowed him to do any more than play with her tits. He had taken her hand and placed it on his erection, initially he had been pleased to hear her sharp intake of breath when she felt the size of him and he was encouraged when she began stroking him.
It was later that the problem started, he had her in the car park, they were kissing, her sweater was pushed up together with her bra and he was sucking on her nipples. He managed to get her laid across the front of one of the cars, his hand up her skirt, trying to pull down her panties. That was when she had stopped him.
https://rentry.org/52ycki8u
https://chyoa.com/user/darii1976
https://vg4s1966.diary.ru/
https://www.hentai-foundry.com/user/llamadrama1984/profile
https://permacultureglobal.org/users/55608-sherry-reid
https://rentry.org/ppb5hmer
https://chyoa.com/user/drombrus1990
https://imageevent.com/nightlite196619601979
https://dragontry1963.bandcamp.com/album/my-first-erotic-story
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3186538
With havin so much written content do you ever run into any issues of plagorism or copyright violation? My blog has a lot of unique content I’ve
either created myself or outsourced but it appears
a lot of it is popping it up all over the internet without my agreement.
Do you know any ways to help prevent content from being stolen? I’d truly appreciate it.
sandyterrace.com
“단서가 있습니다.”Fang Jifan이 갑자기 “사실인가요? “라고 물었습니다.
Hmm it appears like your blog ate my first comment (it was extremely long) so I guess I’ll just sum it up what I had
written and say, I’m thoroughly enjoying your blog.
I too am an aspiring blog blogger but I’m still new to
the whole thing. Do you have any suggestions for inexperienced blog
writers? I’d genuinely appreciate it.
Anti-Forensics involves the use to tools and techniques used to frustrate a digital forensics investigation.
Hello There. I found your blog the use of msn. This is a very neatly written article.
I will be sure to bookmark it and come back to read more of your useful info.
Thanks for the post. I’ll definitely return.
This article will assist the internet viewers for creating new weblog or even a
blog from start to end.
Magnificent goods from you, man. I’ve have in mind your
stuff previous to and you are simply extremely great.
I actually like what you’ve got here, really like what you’re stating
and the best way wherein you are saying it. You are making it enjoyable and you still care
for to keep it wise. I can not wait to read much more
from you. This is actually a tremendous website.
Cool blog! Is your theme custom made or did you download it from
somewhere? A theme like yours with a few simple adjustements
would really make my blog shine. Please let me know
where you got your design. Many thanks
whoah this weblog is great i like studying
your articles. Stay up the great work! You recognize, many persons are hunting around for this info, you
can help them greatly.
I do accept as true with all of the ideas you have introduced
for your post. They are very convincing and can definitely
work. Still, the posts are very quick for beginners.
May you please extend them a little from next time? Thank
you for the post.
Hurrah! After all I got a webpage from where I be able to in fact obtain valuable information concerning
my study and knowledge.
Superb blog you have here but I was wondering if you knew of
any forums that cover the same topics discussed
here? I’d really like to be a part of online community where I can get advice from other knowledgeable individuals
that share the same interest. If you have any recommendations, please let me know.
Cheers!
Thanks very interesting blog!
I’m amazed, I must say. Seldom do I come across a blog that’s
both equally educative and engaging, and let me tell you, you’ve hit the nail on the head.
The problem is something that too few folks are speaking intelligently about.
I am very happy I found this in my search for something relating to this.
I loved as much as you’ll receive carried out right here.
The sketch is attractive, your authored material stylish.
nonetheless, you command get bought an shakiness over
that you wish be delivering the following. unwell unquestionably come more formerly again since exactly the same nearly
a lot often inside case you shield this increase.
It’s actually a cool and useful piece of info. I am happy that you shared this helpful information with us.
Please stay us informed like this. Thanks for sharing.
Hello there! I could have sworn I’ve visited this web site before but after looking at many of the posts
I realized it’s new to me. Nonetheless, I’m definitely delighted I found it and
I’ll be bookmarking it and checking back regularly!
This design is spectacular! You definitely know how to keep a
reader entertained. Between your wit and your videos, I was almost moved to start my own blog
(well, almost…HaHa!) Great job. I really loved what you
had to say, and more than that, how you presented it.
Too cool!
Asking questions are genuinely nice thing if you are not understanding anything entirely, however
this article offers fastidious understanding yet.
I always spent my half an hour to read this weblog’s articles or reviews every day along with
a mug of coffee.
always i used to read smaller articles or reviews that as
well clear their motive, and that is also happening with this post which I
am reading here.
First of all I want to say terrific blog! I had a quick question in which I’d like
to ask if you do not mind. I was curious to know how you center yourself and clear your head prior to writing.
I’ve had difficulty clearing my thoughts in getting my thoughts out
there. I truly do enjoy writing however it just seems like the
first 10 to 15 minutes are usually lost simply just trying to figure out how to begin.
Any recommendations or hints? Thank you!
What i do not realize is in fact how you’re not actually a lot more neatly-liked than you may be right now.
You’re very intelligent. You already know therefore considerably when it comes to this
matter, produced me for my part believe it from numerous various
angles. Its like women and men are not fascinated until
it’s something to do with Lady gaga! Your own stuffs outstanding.
All the time maintain it up!
Great beat ! I would like to apprentice while you amend your website, how could
i subscribe for a blog site? The account aided me a acceptable deal.
I had been a little bit acquainted of this your broadcast provided bright clear concept
I truly love your site.. Excellent colors & theme. Did you build this web site yourself?
Please reply back as I’m looking to create my very own site and want
to find out where you got this from or just what the theme is named.
Thanks!
An interesting discussion is worth comment. There’s no
doubt that that you need to publish more on this subject, it may not be a taboo subject but typically people don’t speak about these
issues. To the next! Kind regards!!
What i don’t understood is actually how you are now not really a lot more well-liked than you
may be right now. You’re very intelligent. You realize therefore significantly with regards to this
topic, made me individually consider it from a lot of varied angles.
Its like women and men are not fascinated until it is something to accomplish with Lady gaga!
Your individual stuffs excellent. Always maintain it up!
Hi, I do think this is a great blog. I stumbledupon it 😉 I’m going to return once again since i have book-marked it.
Money and freedom is the greatest way to change, may you be rich and
continue to help others.
Hello there! This is my first visit to your
blog! We are a collection of volunteers and starting a new initiative in a community in the same niche.
Your blog provided us beneficial information to work on. You have done
a wonderful job!
Hello my friend! I want to say that this post is amazing,
great written and come with almost all important
infos. I would like to see extra posts like this .
Good day! I know this is kinda off topic however I’d figured I’d ask.
Would you be interested in trading links or maybe guest writing a blog article or vice-versa?
My blog discusses a lot of the same subjects as yours and I think we
could greatly benefit from each other. If you happen to be
interested feel free to send me an e-mail.
I look forward to hearing from you! Great blog by the way!
Wonderful blog! Do you have any tips for aspiring writers?
I’m planning to start my own site soon but I’m a little lost on everything.
Would you recommend starting with a free platform like WordPress
or go for a paid option? There are so many options out there
that I’m totally confused .. Any tips? Bless you!
https://ddnews.co.kr/blog/2021/08/23/카카오톡-pc버전-다운로드/
I simply couldn’t leave your web site prior to suggesting that I really
loved the standard information a person supply for your guests?
Is going to be back incessantly in order to investigate cross-check new posts
I was recommended this website by my cousin. I’m
not sure whether this post is written by him as nobody else know such detailed about my trouble.
You are wonderful! Thanks!
Hello, I enjoy reading through your article post. I like to write a little
comment to support you.
Fantastic beat ! I would like to apprentice while you amend your
site, how can i subscribe for a blog web site? The account helped me a acceptable deal.
I had been a little bit acquainted of this your broadcast provided
bright clear idea
I really like what you guys tend to be up too. This kind of clever work and coverage!
Keep up the amazing works guys I’ve included you guys to my own blogroll.
Hi there would you mind sharing which blog platform you’re working with?
I’m planning to start my own blog soon but I’m having a tough time making
a decision between BlogEngine/Wordpress/B2evolution and Drupal.
The reason I ask is because your layout seems
different then most blogs and I’m looking for something completely unique.
P.S My apologies for being off-topic but I had to ask!
영등포안마살롱
Hi! Do you know if they make any plugins to protect against hackers?
I’m kinda paranoid about losing everything I’ve worked hard on. Any recommendations?
qiyezp.com
Fang Jifan은 예의 바르지 않고 바늘을 꼬집고 팔을 세게 찔렀습니다.
I get pleasure from, cause I found just what I was looking for.
You’ve ended my four day long hunt! God Bless you man. Have a great day.
Bye
After looking into a handful of the blog posts on your blog, I really like your way of
writing a blog. I book marked it to my bookmark site list and will be checking back soon.
Please check out my website as well and let me know your opinion.
I’m impressed, I have to admit. Rarely do I encounter a blog that’s both equally educative and entertaining, and without
a doubt, you’ve hit the nail on the head.
The issue is something not enough folks are speaking intelligently about.
I am very happy that I came across this in my hunt for something
relating to this.
That is a great tip particularly to those fresh to the blogosphere.
Simple but very precise information… Thank you for sharing this one.
A must read article!
Hello, just wanted to mention, I loved this article.
It was practical. Keep on posting!
This is really interesting, You’re a very skilled blogger.
I’ve joined your rss feed and look forward to seeking more of your wonderful post.
Also, I’ve shared your site in my social networks!
I’m really impressed with your writing skills as well as with the layout on your blog.
Is this a paid theme or did you customize it yourself?
Anyway keep up the nice quality writing, it’s rare to see a nice blog like this one these days.
Great post! We are linking to this great content on our site.
Keep up the great writing.
Greetings! Very useful advice within this post! It’s the little changes that produce the
greatest changes. Thanks for sharing!
Hi, I do believe your blog could possibly be having internet browser compatibility issues.
When I look at your web site in Safari, it looks fine however,
if opening in IE, it’s got some overlapping issues.
I merely wanted to give you a quick heads up!
Other than that, excellent website!
Hey just wanted to give you a brief heads up and let you know a few
of the pictures aren’t loading properly. I’m not sure why but I think its a linking issue.
I’ve tried it in two different browsers and both show the same outcome.
Great post. I used to be checking constantly this weblog and I’m impressed!
Very helpful information specially the final part 🙂 I deal with such information a lot.
I was seeking this certain info for a long time. Thank you
and best of luck.
An interesting discussion is definitely worth comment.
I think that you should publish more about this topic, it may not be a taboo subject but typically people do not talk about such topics.
To the next! Best wishes!!
Hello, i read your blog occasionally and i own a similar one and i was just wondering
if you get a lot of spam comments? If so how do you prevent it, any plugin or anything you can suggest?
I get so much lately it’s driving me insane so any support is
very much appreciated.
Simply want to say your article is as amazing. The clarity in your post is simply excellent and i could assume you
are an expert on this subject. Fine with your permission allow me to grab your feed to keep up to date
with forthcoming post. Thanks a million and please keep up the enjoyable work.
Heya! I know this is somewhat off-topic however I had to ask.
Does managing a well-established blog such as yours require a large amount of work?
I’m brand new to operating a blog however I do write in my diary daily.
I’d like to start a blog so I can easily share my own experience and
feelings online. Please let me know if you have any ideas or tips for brand new aspiring bloggers.
Thankyou!
이태원스웨디시안마게이클럽
I have been surfing on-line more than 3 hours as of late, but I by no
means discovered any attention-grabbing article like yours.
It is beautiful value sufficient for me. Personally, if all
web owners and bloggers made good content material
as you did, the web shall be a lot more useful than ever before.
I blog often and I truly thank you for your information. This great article has
truly peaked my interest. I will bookmark your
website and keep checking for new details
about once per week. I opted in for your RSS feed as
well.
What’s up to every body, it’s my first pay a quick visit of this weblog; this website consists of remarkable and in fact good
material for readers.
Thank you for the good writeup. It in reality was
a amusement account it. Glance advanced to far added agreeable from you!
By the way, how could we keep up a correspondence?
Hello my friend! I want to say that this post is awesome, great written and
come with almost all vital infos. I’d like to see extra
posts like this .
Magnificent beat ! I would like to apprentice while you amend your website, how could i subscribe for a blog site?
The account aided me a acceptable deal. I had been a little bit acquainted of this your broadcast provided bright clear concept
I believe one of your adverts caused my browser to resize, you might want to put that on your blacklist.
We stumbled over here by a different website and thought I
might as well check things out. I like what I see so now i
am following you. Look forward to looking over your web page yet again.
Hi, Neat post. There’s an issue with your web site in internet
explorer, might check this? IE nonetheless is the market chief
and a huge part of other people will omit your excellent writing due to this problem.
A. 카카오택시 앱을 실행한 후 ‘예약’ 탭을 누르시면 됩니다. 이후 출발 위치, 도착 위치, 탑승 시간 등을 입력하고 ‘예약하기’를 누르시면 예약이 완료됩니다. 다음 사이트에서 섹스 비디오를 시청하세요 강남출장섹스마사지
Hi there I am so thrilled I found your web site, I really found you by error,
while I was browsing on Yahoo for something else, Anyhow I am here now and would just like
to say thank you for a tremendous post and a all round thrilling blog (I also love the theme/design), I don’t have time to
look over it all at the minute but I have saved it and also added in your RSS feeds, so
when I have time I will be back to read more, Please
do keep up the excellent work.
What’s Happening i am new to this, I stumbled upon this I have found It absolutely helpful and it
has helped me out loads. I am hoping to contribute & help different customers like its aided me.
Good job.
Howdy! Someone in my Myspace group shared this site with us so I came
to give it a look. I’m definitely enjoying the information. I’m bookmarking and will be tweeting this to my followers!
Superb blog and wonderful style and design.
Hi, I do think this is an excellent site. I stumbledupon it ;
) I am going to return yet again since i have book-marked it.
Money and freedom is the best way to change, may you be rich and
continue to guide other people.
Do you mind if I quote a few of your posts as long
as I provide credit and sources back to your webpage? My website is in the exact same area
of interest as yours and my visitors would genuinely benefit from a lot of the
information you provide here. Please let me know if this alright with you.
Thank you!
Hay!
Having read this I thought it was rather informative. I appreciate you finding the time and effort to
put this short article together. I once again find myself spending a lot of time both reading and leaving
comments. But so what, it was still worth it!
Useful information. Fortunate me I found your web site by chance, and I’m shocked
why this coincidence did not came about earlier! I bookmarked it.
Hey there exceptional website! Does running a blog like this require a great deal of work?
I’ve very little expertise in computer programming
however I was hoping to start my own blog soon. Anyhow, should you have any recommendations or techniques for new blog owners please share.
I understand this is off subject nevertheless I simply needed to ask.
Thanks!
First of all I want to say excellent blog! I had a quick question in which
I’d like to ask if you do not mind. I was interested to know how you center
yourself and clear your head prior to writing. I have had trouble clearing my thoughts in getting
my ideas out. I do enjoy writing but it just seems like the first 10 to 15 minutes are usually lost simply
just trying to figure out how to begin. Any ideas or hints?
Thank you!
Thanks for sharing your thoughts about mpo777 link. Regards
My brother recommended I might like this web site.
He was totally right. This post truly made my day.
You can not imagine just how much time I had spent for this information! Thanks!
여행지
I for all time emailed this website post page to all my contacts, for the reason that
if like to read it then my friends will too.
Fantastic goods from you, man. I have be mindful your stuff previous to and you are simply too fantastic. I really like what you have received right here, really like what you’re stating and the best way wherein you assert it. You are making it entertaining and you continue to care for to stay it wise. I can not wait to learn far more from you. This is really a terrific web site.
Hello! This post couldn’t be written any better! Reading this post reminds me of my old room mate! He always kept talking about this. I will forward this article to him. Fairly certain he will have a good read. Many thanks for sharing!
Hi there! Quick question that’s entirely off topic. Do you know how to make your site mobile friendly?
My web site looks weird when viewing from my
iphone4. I’m trying to find a template or plugin that might be able to
resolve this issue. If you have any recommendations, please
share. Thanks!
It’s awesome to pay a visit this site and reading the views of all friends concerning this piece of writing, while I am also zealous of getting
experience.
The meal went without incident although a couple of times she had to avert her gaze from Dan for fear of bursting out laughing. Dan went up to his room saying he wanted to listen to music, he laid on his bed playing with himself waiting for his father to leave. He thought about the events of the last two weeks it had been the craziest time of his life.
He put it all down to Mary Harris, she had been his girlfriend for two years, he liked her a lot, she was pretty and intelligent two things that rarely combined in his experience, the problem was she steadfastly refused to let him put his cock inside her, he had thought he was getting somewhere when he took his cock out that evening in the cinema when she had finally allowed him to get his hand inside her panties, it was the first time she had allowed him to do any more than play with her tits. He had taken her hand and placed it on his erection, initially he had been pleased to hear her sharp intake of breath when she felt the size of him and he was encouraged when she began stroking him.
It was later that the problem started, he had her in the car park, they were kissing, her sweater was pushed up together with her bra and he was sucking on her nipples. He managed to get her laid across the front of one of the cars, his hand up her skirt, trying to pull down her panties. That was when she had stopped him.
https://haveagood.holiday/users/337739
https://www.brollopsguiden.se/medlemspresentation/86275
https://www.brollopsguiden.se/medlemspresentation/86064
https://replay7771965.bandcamp.com/album/guilty-pleasures
https://www.brollopsguiden.se/medlemspresentation/86269
https://anotepad.com/notes/madgtm92
https://chyoa.com/user/lowncrew1966
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3188667
https://www.quia.com/profiles/ribowers
https://launchpad.net/~caesarj19771
Howdy this is somewhat of off topic but I was wondering if blogs use WYSIWYG editors or if you have to manually code with HTML. I’m starting a blog soon but have no coding experience so I wanted to get guidance from someone with experience. Any help would be enormously appreciated!
usdt casino erfahrungen
You ought to take part in a contest for one of the highest quality sites on the web.
I most certainly will recommend this website!
hey there and thank you for your information – I have certainly picked
up something new from right here. I did however expertise a few technical
issues using this website, since I experienced
to reload the web site many times previous to I could
get it to load properly. I had been wondering if your hosting
is OK? Not that I’m complaining, but slow loading
instances times will often affect your placement in google
and can damage your quality score if advertising and marketing with Adwords.
Well I am adding this RSS to my email and can look out for
much more of your respective interesting content.
Ensure that you update this again very soon.
The other day, while I was at work, my cousin stole my apple ipad and tested to see if it can survive a 25 foot drop, just so she can be a youtube sensation. My apple ipad is now broken and she has 83 views. I know this is completely off topic but I had to share it with someone!
artrolux koch
수원출장샵
It’s remarkable to pay a visit this site and reading the views of all mates regarding this post, while I am also zealous of getting familiarity.
depanten allegro
That is really attention-grabbing, You are an excessively professional blogger. I’ve joined your rss feed and look forward to in quest of more of your great post. Additionally, I’ve shared your site in my social networks
depanten crema
울산콜걸
Hi to every one, the contents existing at this web site are truly amazing for people knowledge, well, keep up the nice work fellows.
ripper casino no deposit bonus codes 2024
Excellent post but I was wanting to know if you could
write a litte more on this subject? I’d be very grateful if you could
elaborate a little bit further. Thanks!
I do not even know how I ended up here, but I thought this post was
good. I don’t know who you are but definitely you are going to a famous blogger if you aren’t already 😉 Cheers!
At this time it seems like Drupal is the preferred blogging platform out there right now. (from what I’ve read) Is that what you’re using on your blog?
beep beep casino 20
I’m really enjoying the theme/design of your weblog.
Do you ever run into any internet browser compatibility issues?
A number of my blog audience have complained
about my site not operating correctly in Explorer but looks great in Chrome.
Do you have any recommendations to help fix this problem?
Amazing things here. I am very satisfied to see your article.
Thanks so much and I am looking ahead to touch you. Will you please drop me a e-mail?
Definitely believe that which you said. Your favorite justification appeared to be on the net the easiest thing to be aware of.
I say to you, I definitely get irked while people
consider worries that they plainly do not know about.
You managed to hit the nail upon the top as well
as defined out the whole thing without having side effect , people can take a signal.
Will probably be back to get more. Thanks
Hello I am so excited I found your website,
I really found you by mistake, while I was browsing on Yahoo for something else, Regardless
I am here now and would just like to say thank
you for a remarkable post and a all round entertaining blog (I also love the
theme/design), I don’t have time to read through it all at the moment
but I have saved it and also included your RSS feeds, so when I have time I will be back to read more,
Please do keep up the excellent b.
Very rapidly this site will be famous among all blogging viewers, due to it’s nice articles
거제출장안마
It’s truly very complex in this full of activity life to listen news on Television, therefore I just use web for that purpose, and obtain the hottest information.
Hi there, i read your blog from time to time and i own a similar one and i
was just curious if you get a lot of spam comments?
If so how do you protect against it, any plugin or anything you can advise?
I get so much lately it’s driving me mad so
any assistance is very much appreciated.
Hello there, There’s no doubt that your website
may be having browser compatibility problems. Whenever I take a
look at your blog in Safari, it looks fine but when opening in Internet Explorer, it
has some overlapping issues. I simply wanted to give you a quick heads up!
Apart from that, excellent blog!
Now I am going away to do my breakfast, once having my breakfast coming over again to read
additional news.
Gama Casino
Gama Casino
Just desire to say your article is as surprising.
The clearness for your post is just cool and i can assume
you’re knowledgeable on this subject. Well along with your permission allow
me to grab your feed to stay up to date
with coming near near post. Thank you one million and please keep up the rewarding
work.
Hi! I just wanted to ask if you ever have any trouble with hackers? My last blog (wordpress) was hacked and I ended up losing many months of hard work due to no data backup. Do you have any methods to prevent hackers?
This is a topic that is near to my heart… Best wishes!
Where are your contact details though?
아름다운스웨디시업소
Отечественный производитель продает разборные гантели https://gantel-razbornaya.ru/ – у нас найдете замечательный объем предложений. Сборные утяжелители дают продуктивно выполнять силовые тренировки в любом месте. Изделия для спорта отличаются функциональностью, универсальностью в эксплуатации. Предприятие эффективно испытывает и внедряет свежие идеи, чтобы исполнить спортивные цели постоянных клиентов. В изготовлении качественного инвентаря реализуются высококлассные марки металла. Достойный каталог интернет-магазина снарядов позволяет получить разборные отягощения для продуктивной программы тренировок. Для домашних тренировок – это приятный инвентарь с маленькими размерами и большой фунциональности.
No matter if some one searches for his necessary thing,
therefore he/she wants to be available that in detail, so that
thing is maintained over here.
I couldn’t refrain from commenting. Perfectly written!
Greetings from Carolina! I’m bored to tears at work so I decided to browse your website on my
iphone during lunch break. I really like the information you provide here and can’t wait to take a look when I get home.
I’m amazed at how fast your blog loaded on my phone ..
I’m not even using WIFI, just 3G .. Anyways, wonderful
site!
https://kakaotaxi.dasgno.com/kakao-pay
Hello there! I could have sworn I’ve been to this blog before but after browsing through some of
the articles I realized it’s new to me. Anyhow, I’m certainly pleased I came across it and I’ll be bookmarking it and checking back regularly!
May I simply say what a comfort to uncover someone that truly understands what they’re talking about on the web.
You definitely realize how to bring a problem to light and make it important.
More people must check this out and understand this side
of the story. I was surprised that you aren’t
more popular because you certainly have the gift.
I got this site from my pal who told me concerning this web page and now this time
I am visiting this web page and reading very informative content here.
Hi i am kavin, its my first occasion to commenting anywhere, when i read this
article i thought i could also create comment due to this good post.
하동동해출장만남 소자본 창업
Link exchange is nothing else however it is just placing the other person’s web site
link on your page at appropriate place and other person will
also do same in favor of you.
Hey there! I could have sworn I’ve been to this website before
but after browsing through some of the post I realized it’s new to me.
Anyhow, I’m definitely happy I found it and I’ll be bookmarking and checking back
often!
Heya i’m for the first time here. I found this board and I find It truly useful & it helped me out much.
I hope to give something back and help others like
you helped me.
For hottest news you have to visit internet and on world-wide-web I found
this site as a most excellent web page for most
up-to-date updates.
Nice post. I learn something new and challenging on blogs I
stumbleupon every day. It will always be useful to read content from other authors and use a little something from other websites.
Hi there to every one, the contents present at this web site are actually awesome for people knowledge, well, keep up the
nice work fellows.
Great site you have got here.. It’s hard to find quality writing like yours
nowadays. I really appreciate individuals like you!
Take care!!
A fascinating discussion is worth comment. I do think that you should publish more
on this subject matter, it might not be a taboo subject but generally people do not speak about such issues.
To the next! Best wishes!!
With havin so much content and articles do you ever run into any issues of plagorism or copyright infringement? My blog has a lot of unique content I’ve either written myself or outsourced but it looks like a lot of it is popping it up all over the internet without my authorization. Do you know any methods to help prevent content from being stolen? I’d really appreciate it.
Do you have any video of that? I’d like to find out some additional information.
I needed to thank you for this good read!!
I absolutely enjoyed every bit of it. I have got you bookmarked
to check out new things you post…
Hey I know this is off topic but I was wondering if you knew of any widgets I could
add to my blog that automatically tweet my newest twitter updates.
I’ve been looking for a plug-in like this for quite some time and was hoping maybe you would have some experience with something like this.
Please let me know if you run into anything. I truly enjoy reading your blog
and I look forward to your new updates.
Создаваемые отечественной компанией тренажеры для кинезитерапии https://trenazhery-dlya-kineziterapii.ru и специально разработаны для восстановления после травм. Конструкции имеют интересное предложение цены и качества.
Продаем очень недорого блочную раму с усиленной конструкцией. В каталоге для кинезитерапии всегда в реализации модели грузоблочного и нагружаемого типа.
Выпускаемые тренажеры для реабилитации обеспечивают мягкую и безопасную тренировку, что особенно важно для тренирующихся пациентов в процессе восстановления.
Устройства обладают регулируемым сопротивлением и уровнями нагрузки, что дает возможность индивидуализировать силовые тренировки в соответствии с задачами каждого пациента.
Все тренажеры актуальны для ЛФК по методике врача Сергея Бубновского. Оборудованы рукоятками для комфортного осуществления тяг сидя или лежа.
Hello, I enjoy reading through your article post. I like to write a little comment to support you.
Hello there, You’ve done an incredible job. I will certainly digg it and personally suggest to my friends.
I am sure they’ll be benefited from this web site.
Hey I know this is off topic but I was wondering if you
knew of any widgets I could add to my blog that automatically tweet my newest twitter updates.
I’ve been looking for a plug-in like this for quite some time and was hoping maybe you would have some experience with something like this.
Please let me know if you run into anything. I truly enjoy reading your blog and I look
forward to your new updates.
Howdy! This is my 1st comment here so I just wanted to give a quick shout out and say I genuinely enjoy reading your articles.
Can you suggest any other blogs/websites/forums that deal
with the same subjects? Thanks for your time!
My brother recommended I may like this website. He was once entirely right.
This post truly made my day. You can not consider simply
how a lot time I had spent for this info! Thank you!
I always spent my half an hour to read this website’s posts all the time along with a mug of coffee.
What’s up it’s me, I am also visiting this web page regularly, this site is
in fact fastidious and the people are in fact sharing pleasant thoughts.
hi!,I love your writing very so much! percentage we keep up a
correspondence more approximately your article on AOL?
I require a specialist in this area to solve my problem.
May be that’s you! Looking forward to look you.
I’m amazed, I have to admit. Seldom do I encounter a blog
that’s both equally educative and entertaining, and
let me tell you, you’ve hit the nail on the head.
The issue is something which too few folks are speaking intelligently about.
Now i’m very happy that I stumbled across this in my search for something relating to this.
I think this is one of the most vital information for
me. And i am glad reading your article. But want to remark
on few general things, The site style is great, the articles is really nice : D.
Good job, cheers
Hi there, its pleasant paragraph concerning media print, we all know media is
a fantastic source of facts.
Very good post. I will be going through a few of these issues as well..
I want to to thank you for this good read!! I definitely loved every little bit of it.
I have you saved as a favorite to check out new stuff you post…
Good day! Do you use Twitter? I’d like to follow you if that would be ok.
I’m undoubtedly enjoying your blog and look forward to new
posts.
Fantastic site you have here but I was curious about if you knew of any forums that cover the same topics talked about in this article? I’d really like to be a part of community where I can get advice from other experienced individuals that share the same interest. If you have any suggestions, please let me know. Many thanks!
http://www.ku.undong.net/bbs/board.php?bo_table=free&wr_id=75541
https://ad.infocloud.co.kr/bbs/board.php?bo_table=free&wr_id=1215095
http://sinbiromall.hubweb.net/bbs/board.php?bo_table=free&wr_id=367823
http://lstelecom.co.kr/bbs/board.php?bo_table=free&wr_id=69849
http://old.remain.co.kr/bbs/board.php?bo_table=free&wr_id=1101080
Hey I know this is off topic but I was wondering if you knew of any widgets I could add to my blog that automatically tweet my newest
twitter updates. I’ve been looking for a plug-in like this for quite some time and was hoping maybe you would
have some experience with something like this.
Please let me know if you run into anything. I truly enjoy reading your
blog and I look forward to your new updates.
I really like what you guys tend to be up too. Such clever work and coverage!
Keep up the very good works guys I’ve included you guys to blogroll.
I am regular visitor, how are you everybody? This piece of
writing posted at this web site is really good.
Hello, I enjoy reading through your article. I like to write a little comment to support you.
Great goods from you, man. I have understand your stuff previous to and you
are just extremely excellent. I really like what you’ve acquired here, really like what you are saying
and the way in which you say it. You make it enjoyable and you still take care of to
keep it smart. I can’t wait to read far more from you.
This is actually a great website.
my web blog … چت سکسی
Hi Dear, are you really visiting this site daily, if so afterward you will without doubt obtain good knowledge.
강남콜걸
Hmm it looks like your site ate my first comment (it was extremely long) so I guess
I’ll just sum it up what I wrote and say, I’m thoroughly enjoying your blog.
I too am an aspiring blog blogger but I’m still new to everything.
Do you have any helpful hints for newbie blog writers?
I’d genuinely appreciate it.
After looking into a handful of the blog posts on your site,
I truly like your way of writing a blog. I bookmarked it to my bookmark webpage list and will be checking back in the near future.
Take a look at my website as well and tell me what you think.
Hello very cool website!! Guy .. Excellent .. Superb .. I’ll bookmark your blog and take the feeds also? I am satisfied to find a lot of useful info here within the publish, we want develop more techniques in this regard, thank you for sharing. . . . . .
keramin krem benu gyogyszertar
Good way of describing, and nice piece of writing to obtain facts concerning
my presentation focus, which i am going to deliver
in college.
Right here is the perfect site for anyone who wishes to find out about this topic. You understand a whole lot its almost hard to argue with you (not that I really would want to…HaHa). You definitely put a brand new spin on a topic that has been written about for decades. Wonderful stuff, just excellent!
keramin crema funghi
Pretty section of content. I just stumbled upon your blog and in accession capital to assert that I get in fact enjoyed account your blog posts. Anyway I will be subscribing to your augment and even I achievement you access consistently quickly.
dissertation literature review writing services
WOW just what I was looking for. Came here by searching for https://linkdomino168.fans/
Howdy, i read your blog from time to time and i own a similar one and i
was just curious if you get a lot of spam responses?
If so how do you stop it, any plugin or anything you
can advise? I get so much lately it’s driving me crazy so
any help is very much appreciated.
Hi it’s me, I am also visiting this website on a regular basis, this website is in fact nice
and the people are in fact sharing pleasant thoughts.
The meal went without incident although a couple of times she had to avert her gaze from Dan for fear of bursting out laughing. Dan went up to his room saying he wanted to listen to music, he laid on his bed playing with himself waiting for his father to leave. He thought about the events of the last two weeks it had been the craziest time of his life.
He put it all down to Mary Harris, she had been his girlfriend for two years, he liked her a lot, she was pretty and intelligent two things that rarely combined in his experience, the problem was she steadfastly refused to let him put his cock inside her, he had thought he was getting somewhere when he took his cock out that evening in the cinema when she had finally allowed him to get his hand inside her panties, it was the first time she had allowed him to do any more than play with her tits. He had taken her hand and placed it on his erection, initially he had been pleased to hear her sharp intake of breath when she felt the size of him and he was encouraged when she began stroking him.
It was later that the problem started, he had her in the car park, they were kissing, her sweater was pushed up together with her bra and he was sucking on her nipples. He managed to get her laid across the front of one of the cars, his hand up her skirt, trying to pull down her panties. That was when she had stopped him.
https://okwave.jp/profile/u3105891.html
https://launchpad.net/~absconcier19931
https://www.hentai-foundry.com/user/liglaz1984/profile
http://www.nfomedia.com/profile?uid=rOiShiF
https://rentry.org/ze2tbrte
https://anotepad.com/notes/kgxpj6gf
https://chyoa.com/user/rao1961
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3187283
https://www.divephotoguide.com/user/sepiatone1963
https://launchpad.net/~erebus19751
When some one searches for his vital thing,
so he/she desires to be available that in detail, thus that thing is
maintained over here.
I’m really loving the theme/design of your website.
Do you ever run into any web browser compatibility problems?
A number of my blog readers have complained about my website not operating correctly in Explorer but
looks great in Opera. Do you have any tips to
help fix this problem?
Good day! I could have sworn I’ve visited this blog before but after
browsing through some of the articles I realized it’s new to
me. Anyways, I’m definitely pleased I discovered it and I’ll be
bookmarking it and checking back often!
수원출장샵
An outstanding share! I’ve just forwarded this onto a co-worker who
had been doing a little research on this. And he actually bought me lunch due to the fact
that I stumbled upon it for him… lol. So let me reword this….
Thanks for the meal!! But yeah, thanx for spending time to discuss this topic here on your blog.
Hey guys, let’s talk about the brain! Did you know our brain has special parts that help us remember things and deal with our feelings? One important part is called the fornix. It’s like a highway that carries messages between our memory center, called the hippocampus, and other parts of the brain. So, when you’re trying to remember something, like your favorite song lyrics or what you learned in class, thank your fornix!
Now, let’s talk about another part called the stria terminalis. This one helps us handle emotions, especially when we’re stressed or scared. It’s like our brain’s alarm system! When something scary happens, the stria terminalis helps our brain figure out how to react. Pretty cool, right?
Understanding these parts of our brain can help scientists learn more about how our minds work and how to help people who might have problems with memory or emotions. So, next time you’re learning about the brain, remember the fornix and stria terminalis!
Attractive section of content. I just stumbled upon your website and in accession capital to assert
that I acquire actually enjoyed account your blog posts.
Any way I will be subscribing to your feeds and even I achievement you access consistently quickly.
Amazing! Its actually amazing post, I have got much clear idea regarding from this paragraph.
#best#links#
virtueel nummer voor altijd
Woah! I’m really enjoying the template/theme of this site.
It’s simple, yet effective. A lot of times it’s difficult to get that “perfect balance” between usability and visual appearance.
I must say you have done a amazing job with this.
In addition, the blog loads extremely quick for me on Opera.
Superb Blog!
Good day! Do you know if they make any plugins to protect against hackers?
I’m kinda paranoid about losing everything I’ve worked hard
on. Any tips?
This is very fascinating, You’re a very professional blogger. I have joined your feed and look ahead to searching for more of your excellent post. Also, I’ve shared your site in my social networks
http://www.webmast.kr/bbs/board.php?bo_table=free&wr_id=112952
https://hosimkig.gwangju.ac.kr/bbs/board.php?bo_table=free&wr_id=567284
http://lstelecom.co.kr/bbs/board.php?bo_table=free&wr_id=69908
http://trademaker.co.kr/yc5/bbs/board.php?bo_table=free&wr_id=2467764
http://121.254.254.30/bbs/board.php?bo_table=free&wr_id=990557
This article gives clear idea in support of the new users of blogging,
that in fact how to do blogging and site-building.
Thanks , I’ve recently been looking for information about this topic for a while and
yours is the greatest I’ve came upon till now. But,
what concerning the conclusion? Are you positive in regards to the supply?
https://www.pepperboy.kr/2022/06/How-to-use-Greek-yogurt.html
Appreciating the dedication you put into your blog and detailed information you provide. It’s great to come across a blog every once in a while that isn’t the same old rehashed information. Great read! I’ve saved your site and I’m including your RSS feeds to my Google account.
http://www.glat.kr/bbs/board.php?bo_table=free&wr_id=481219
I constantly emailed this web site post page to all my contacts, since
if like to read it after that my friends will too.
Greetings! This is my first visit to your blog!
We are a group of volunteers and starting a new project in a community in the same niche.
Your blog provided us useful information to work on. You have done a extraordinary job!
Woah! I’m really enjoying the template/theme of this website.
It’s simple, yet effective. A lot of times it’s very hard to get that
“perfect balance” between user friendliness and visual appearance.
I must say you’ve done a great job with this. In addition, the blog loads very quick for me on Opera.
Outstanding Blog!
I do not know if it’s just me or if everyone else encountering issues with
your blog. It appears as if some of the text in your posts are running off the screen. Can somebody else please
comment and let me know if this is happening
to them too? This could be a issue with my
internet browser because I’ve had this happen previously.
Cheers
대전나이트클럽
Great article! We are linking to this particularly great article on our website.
Keep up the great writing.
I do not even understand how I ended up right here, but
I thought this post was good. I don’t understand who you’re
however definitely you are going to a well-known blogger if you aren’t already.
Cheers!
Hi, Neat post. There is an issue together with your site in internet explorer, could check this?IE nonetheless is the market leader and a good part of people will leave out your wonderful writing due to this problem.
There is clearly a bundle to realize about this. I feel you made certain good points in features also.
Appreciating the hard work you put into your site and in depth information you offer. It’s nice to come across a blog every once in a while that isn’t the same old rehashed information. Great read! I’ve bookmarked your site and I’m including your RSS feeds to my Google account.
Thank you for another informative site. The place else could I am getting that type of information written in such an ideal method?
I’ve a mission that I’m just now operating on, and I have been at the look out
for such information.
Very descriptive post, I enjoyed that bit. Will there be
a part 2?
Please let me know if you’re looking for a writer for your site.
You have some really good posts and I think I would be a good asset.
If you ever want to take some of the load off, I’d love to write some content for your
blog in exchange for a link back to mine. Please blast me an e-mail if
interested. Many thanks!
My coder is trying to persuade me to move to .net from PHP.
I have always disliked the idea because of the expenses.
But he’s tryiong none the less. I’ve been using WordPress
on several websites for about a year and am anxious about switching to another platform.
I have heard fantastic things about blogengine.net.
Is there a way I can import all my wordpress posts into it?
Any help would be really appreciated!
Hi there, i read your blog from time to time and i own a similar one and i was just wondering if you get a lot of spam comments?
If so how do you stop it, any plugin or anything you
can recommend? I get so much lately it’s driving me insane so any help is very much appreciated.
영등포안마살롱
Excellent web site. Lots of helpful information here.
I am sending it to a few pals ans also sharing in delicious.
And naturally, thanks in your sweat!
exprimegranada.com
このブログはいつも私に多くを教えてくれます。感謝しています。
Hi there, all the time i used to check webpage posts here early in the morning,
since i love to gain knowledge of more and more.
Wonderful beat ! I would like to apprentice while you amend your site, how can i subscribe for a blog website?
The account aided me a acceptable deal. I had been a little bit acquainted of this your broadcast provided bright
clear concept
Thanks for sharing excellent informations. Your web-site is very cool. I am impressed by the details that you have on this blog. It reveals how nicely you understand this subject. Bookmarked this web page, will come back for extra articles. You, my pal, ROCK! I found simply the information I already searched all over the place and just couldn’t come across. What a great web site.
Remarkable! Its actually awesome piece of writing, I have got much clear idea on the topic of from this piece of writing.
Гама казино
Just desire to say your article is as astounding. The clarity in your submit is simply spectacular and i could assume you’re a professional in this subject. Fine along with your permission allow me to seize your RSS feed to stay up to date with impending post. Thank you a million and please carry on the gratifying work.
Ginkgo Biloba для мозга и нервной системы 130 mg
Superb website you have here but I was wondering if
you knew of any forums that cover the same topics discussed in this article?
I’d really love to be a part of group where I can get
suggestions from other experienced individuals that share the same
interest. If you have any suggestions, please let me know.
Thank you!
Do you have a spam problem on this website; I also am a blogger, and I was wanting
to know your situation; we have developed some
nice practices and we are looking to swap techniques with other folks,
please shoot me an e-mail if interested.
https://acase.co.kr/35
이태원스웨디시안마게이클럽
After looking over a handful of the blog articles on your web site, I truly appreciate
your way of writing a blog. I bookmarked it to my bookmark website list and will be checking back in the near future.
Please check out my web site too and tell me how you
feel.
It’s very easy to find out any topic on net as compared to books, as I found this post
at this site.
Terrific article! That is the kind of info that are meant to be shared across the net.
Shame on Google for not positioning this post higher!
Come on over and visit my site . Thank you =)
Great blog! Do you have any recommendations for aspiring writers?
I’m hoping to start my own site soon but I’m a little lost on everything.
Would you propose starting with a free platform like WordPress or go for a paid option? There are so many choices out
there that I’m completely overwhelmed .. Any recommendations?
Thank you!
https://info.channel.seoul.kr/news/3859
Hi, Neat post. There is an issue with your site in internet explorer, could check this?
IE nonetheless is the marketplace chief and a big section of other folks
will pass over your great writing due to this problem.
Hi there! Do you know if they make any plugins to assist with SEO?
I’m trying to get my blog to rank for some targeted keywords but I’m not seeing
very good success. If you know of any please share.
Many thanks! I saw similar art here: Link Building
https://dday.tistory.com/263
Good day! I know this is somewhat off topic but I was wondering which blog
platform are you using for this site? I’m getting fed up of WordPress because
I’ve had problems with hackers and I’m looking at options for
another platform. I would be fantastic if you could point me in the direction of a good platform.
Hey I am so grateful I found your site, I really found you by accident,
while I was researching on Google for something else, Anyways I am here now and
would just like to say cheers for a fantastic post and a
all round exciting blog (I also love the theme/design),
I don’t have time to look over it all at the moment but I have
bookmarked it and also added your RSS feeds, so when I have time I will be back to read much more,
Please do keep up the fantastic job.
It is in point of fact a nice and useful piece of information. I’m happy that you simply shared this helpful
info with us. Please stay us up to date like
this. Thanks for sharing.
I’m amazed, I must say. Rarely do I encounter a blog that’s both educative and entertaining, and without a doubt, you’ve hit the
nail on the head. The problem is an issue that too few men and women are speaking intelligently about.
Now i’m very happy I came across this in my search for something concerning this.
I like reading through an article that can make men and women think.
Also, thank you for allowing me to comment!
Hello! Do you know if they make any plugins to assist with SEO?
I’m trying to get my website to rank for some targeted keywords but I’m
not seeing very good results. If you know of any please share.
Thank you! You can read similar art here: AA List
https://nicesongtoyou.com/job/byeolugsijang-newspaper
This is my first time go to see at here and i am actually pleassant to read all at alone place.
Hi i am kavin, its my first occasion to commenting anyplace, when i read this article
i thought i could also create comment due to this good article.
https://download-forum.co.kr/entry/넥슨-v4-다운로드
https://download-install.com/tag/네이트온20기업사용
Wow that was strange. I just wrote an incredibly long comment but after I clicked submit my comment didn’t appear.
Grrrr… well I’m not writing all that over again.
Anyway, just wanted to say wonderful blog!
Казино VODKA онлайн – играть в автоматы на деньги
Узнайте о захватывающем мире казино VODKA, где современный дизайн, разнообразие игровых автоматов и щедрые бонусы ждут каждого игрока. Погрузитесь в атмосферу слотов на деньги с казино VODKA.
Казино VODKA: Погружение в мир азартных развлечений
В мире азартных игр существует огромное количество казино, каждое из которых стремится привлечь внимание игроков своими уникальными предложениями и атмосферой. Одним из таких заведений является казино VODKA, которое предлагает своим посетителям захватывающие игровые возможности и неповторимый опыт азартных развлечений.
Виртуальное пространство казино VODKA
Казино VODKA водка бет казино играть предлагает своим клиентам широкий спектр азартных игр, доступных в виртуальном пространстве. От классических игровых автоматов до настольных игр, таких как рулетка, блэкджек и покер – здесь каждый игрок сможет найти что-то по своему вкусу. Современный дизайн и удобный интерфейс позволяют наслаждаться игровым процессом без каких-либо проблем или задержек.
Бонусы и акции
Одним из способов привлечения новых игроков и поощрения постоянных являются бонусы и акции. Казино VODKA не остается в стороне и предлагает своим клиентам различные бонусы за регистрацию, первые депозиты или участие в акциях. Эти бонусы могут значительно увеличить шансы на победу и сделать игровой процесс еще более увлекательным.
Безопасность и поддержка
Важным аспектом любого казино является обеспечение безопасности игроков и защита их личной информации. Казино VODKA придает этому особое внимание, используя передовые технологии шифрования данных и обеспечивая конфиденциальность всех транзакций. Кроме того, круглосуточная служба поддержки готова ответить на любые вопросы и помочь в решении возникающих проблем.
Заключение
Казино VODKA – это место, где каждый азартный игрок найдет что-то по своему вкусу. Богатый выбор игр, интересные бонусы и высокий уровень безопасности делают его привлекательным вариантом для тех, кто хочет испытать удачу и получить незабываемые эмоции от азартных развлечений. Сделайте свой первый шаг в мир азарта и испытайте удачу в казино VODKA уже сегодня!
https://film35.tistory.com/72
bookmarked!!, I like your web site!
https://kakaotaxi.dasgno.com/kakao-taxi2
https://www.pornhub.com/view_video.php?viewkey=ph58c466aa61bc5
https://hipsterlibertarian.com/6791
I have read so many articles or reviews on the topic of the blogger lovers however this article is really a
good piece of writing, keep it up.
Nice post. I was checking continuously this blog and I’m impressed!
Extremely useful information particularly the last part 🙂 I care for such info much.
I was seeking this particular information for a very long time.
Thank you and best of luck.
Every weekend i used to go to see this website, as i wish for enjoyment, as this this
web site conations genuinely nice funny data too.
I am not sure where you’re getting your info, however great topic.
I must spend some time studying much more or understanding more.
Thanks for magnificent information I used to be in search of this information for my mission.
This is very fascinating, You are an excessively skilled blogger.
I’ve joined your feed and sit up for seeking more of your wonderful
post. Also, I have shared your web site in my social networks
Hey are using WordPress for your site platform?
I’m new to the blog world but I’m trying to get started and set up my own. Do you require any
coding expertise to make your own blog? Any help would be really appreciated!
My partner and I stumbled over here coming from a different website and thought I should check things out.
I like what I see so i am just following you. Look forward to looking at
your web page again.
magnificent issues altogether, you simply won a emblem new reader.
What could you suggest about your post that you just made some
days in the past? Any sure?
I read this article completely on the topic of the comparison of most recent and preceding technologies,
it’s remarkable article.
Hmm is anyone else experiencing problems with the images on this blog loading?
I’m trying to determine if its a problem on my end or if it’s the
blog. Any feedback would be greatly appreciated.
Hi there, just wanted to mention, I liked this article.
It was helpful. Keep on posting!
Hi to all, it’s genuinely a good for me to pay a visit this web site, it consists of helpful Information.
Superb website you have here but I was wanting to know if you knew of any discussion boards that cover the same topics
discussed in this article? I’d really love to be a part of group
where I can get feed-back from other experienced individuals that share the same interest.
If you have any suggestions, please let me know.
Bless you!
https://tinyurl.com/2gc753le
https://new-app.download/windows/tving/
Woah! I’m really loving the template/theme of this site.
It’s simple, yet effective. A lot of times it’s tough to get that “perfect balance” between user friendliness and appearance.
I must say you have done a very good job with this. In addition, the blog loads super quick for me on Chrome.
Excellent Blog!
Good way of describing, and good paragraph to obtain information regarding my presentation subject, which i am going to convey in institution of
higher education.
Зайдите на сайт https://pecom.asia/ и вы сможете воспользоваться услугами транспортной компании по доставке грузов из Китая в Россию. Доставка товаров из Китая осуществляется в максимально сжатые сроки. Мы оказываем большой перечень услуг по грузоперевозкам из Китая, страхованию, таможенному оформлению. Грузоперевозки и доставка товаров из Китая по выгодным ценам. Подробнее на сайте.
Spot on with this write-up, I actually believe this web site needs a
lot more attention. I’ll probably be back again to read through more, thanks for the info!
This post is worth everyone’s attention. How can I find
out more?
Hi, I do think this is a great blog. I stumbledupon it 😉 I am going to revisit once again since i
have book-marked it. Money and freedom is the best way
to change, may you be rich and continue to help other people.
https://www.pornhub.com/view_video.php?viewkey=ph5fa4d22a641bd
The meal went without incident although a couple of times she had to avert her gaze from Dan for fear of bursting out laughing. Dan went up to his room saying he wanted to listen to music, he laid on his bed playing with himself waiting for his father to leave. He thought about the events of the last two weeks it had been the craziest time of his life.
He put it all down to Mary Harris, she had been his girlfriend for two years, he liked her a lot, she was pretty and intelligent two things that rarely combined in his experience, the problem was she steadfastly refused to let him put his cock inside her, he had thought he was getting somewhere when he took his cock out that evening in the cinema when she had finally allowed him to get his hand inside her panties, it was the first time she had allowed him to do any more than play with her tits. He had taken her hand and placed it on his erection, initially he had been pleased to hear her sharp intake of breath when she felt the size of him and he was encouraged when she began stroking him.
It was later that the problem started, he had her in the car park, they were kissing, her sweater was pushed up together with her bra and he was sucking on her nipples. He managed to get her laid across the front of one of the cars, his hand up her skirt, trying to pull down her panties. That was when she had stopped him.
https://anotepad.com/notes/rbqcqpt3
https://www.hentai-foundry.com/user/moonlighter1956/profile
https://www.haikudeck.com/presentations/O4yPvAbtmj
https://www.haikudeck.com/presentations/GGOBElXupc
https://www.haikudeck.com/presentations/varIxv1C6r
https://rentry.org/wdpfemqa
https://launchpad.net/~lusterbunny199919851
https://anotepad.com/notes/d6d537g4
https://www.quia.com/profiles/s356roderick
https://haveagood.holiday/users/339779
Great goods from you, man. I have understand your stuff previous
to and you are just extremely great. I really like what you have acquired here, certainly like what you’re stating and the way in which you say it.
You make it enjoyable and you still care for to keep it smart.
I cant wait to read much more from you. This
is really a tremendous web site.
https://king.creaming.net/on/8504
Всероссийская благотворительная школьная ярмарка предлагает вам разместить собственные товары на ней или купить за образовательную валюту бенефиты и услуги в приложении VK Умникоины без какой-либо регистрации и скачивания. https://start.shkolnayayarmarka.ru/ – сайт, где есть возможность в качестве пожертвования разместить личные товары и услуги. Также здесь вы подробнее ознакомитесь с единицей обмена, которую применяют для осуществления сделок между покупателем и продавцом на благотворительных ярмарках. Пишите нам уже сейчас.
I’m really inspired along with your writing talents as well as with the structure to
your blog. Is that this a paid subject or did you customize it your self?
Anyway keep up the excellent quality writing,
it is rare to see a nice weblog like this one nowadays..
Hi to every , as I am truly keen of reading this blog’s post to be updated regularly. It contains good material.
http://www.wonkhouse.co.kr/bbs/board.php?bo_table=free&wr_id=1460022
I know this web site presents quality based content and additional data,
is there any other site which gives these things in quality?
https://dday.tistory.com/988
https://new-software.download/windows/sandboxie/
Hey everyone, I just wanted to share this cool blog I found that covers everything Artificial Intelligence! Whether you’re a total beginner or a seasoned AI enthusiast, there’s something for everyone here. Check it out: https://everyminutedeals.com/category/artificial-intelligence/
Heya are using WordPress for your blog platform?
I’m new to the blog world but I’m trying to get started
and create my own. Do you need any coding expertise to make your own blog?
Any help would be really appreciated!
I don’t know if it’s just me or if perhaps everybody else encountering issues with your blog.
It appears as if some of the written text within your content are running off the screen.
Can somebody else please comment and let me know if this is happening to
them too? This may be a issue with my browser because I’ve had this happen previously.
Thanks
Spot on with this write-up, I honestly believe this website needs far more attention.
I’ll probably be returning to see more, thanks for the information!
Thanks very nice blog!
Great post! We will be linking to this great post on our website.
Keep up the good writing.
Thank you for the auspicious writeup. It in fact was a amusement account it.
Look advanced to far added agreeable from you! However, how could we communicate?
Very nice post. I just stumbled upon your blog and wanted to say that I’ve truly enjoyed
browsing your blog posts. After all I will be subscribing
on your rss feed and I’m hoping you write once more soon!
Hi, its pleasant paragraph about media print,
we all know media is a fantastic source of facts.
THCA Flower Indoor and Green House Cheap Ounces at THCA KING
One other thing to point out is that an online business administration course is designed for learners to be able to efficiently proceed to bachelor degree education. The 90 credit college degree meets the lower bachelor college degree requirements and once you earn your own associate of arts in BA online, you’ll have access to the most up-to-date technologies with this field. Some reasons why students are able to get their associate degree in business is because they are interested in this area and want to have the general education and learning necessary just before jumping in a bachelor education program. Many thanks for the tips you really provide within your blog.
Magnificent beat ! I would like to apprentice while you amend your site, how can i subscribe for a blog website?
The account helped me a acceptable deal. I had been a little bit
acquainted of this your broadcast provided bright clear idea
Hi there to every one, the contents present at this web site are really remarkable for people knowledge, well,
keep up the good work fellows.
Hmm is anyone else encountering problems with the pictures on this blog
loading? I’m trying to figure out if its a problem on my end or if it’s the blog.
Any suggestions would be greatly appreciated.
Asking questions are truly nice thing if you are not understanding something completely, however
this post offers fastidious understanding yet.
I’m really enjoying the design and layout of your blog.
It’s a very easy on the eyes which makes it much more pleasant for me to come here and visit more often. Did you hire out a designer to create your theme?
Superb work!
How do I switch some blog posts from one of my blogs, onto a different blog. Both on Blogger?
Greetings! Very helpful advice within this article!
It is the little changes that will make the largest changes.
Thanks a lot for sharing!
Way cool! Some very valid points! I appreciate you writing this post plus the rest of the website is very good.
nice article
Appreciating the commitment you put into your blog and
in depth information you present. It’s nice to
come across a blog every once in a while that isn’t the
same out of date rehashed information. Great read! I’ve saved your site and I’m adding your RSS feeds to my
Google account.
First off I would like to say wonderful blog! I had a quick question which I’d like to ask if you do not mind.
I was curious to find out how you center yourself and clear
your mind before writing. I’ve had difficulty clearing my thoughts in getting
my thoughts out. I do enjoy writing however it just seems like the first 10 to
15 minutes are wasted just trying to figure out how to begin. Any recommendations
or hints? Kudos!
Hello there, just became aware of your blog through Google,
and found that it is truly informative. I am going
to watch out for brussels. I will be grateful if you continue this in future.
A lot of people will be benefited from your writing.
Cheers!
What i do not understood is if truth be told how you’re now not
actually a lot more smartly-preferred than you might be right now.
You are so intelligent. You know thus considerably relating to this subject, made me personally believe it
from numerous numerous angles. Its like men and women aren’t involved until
it is something to accomplish with Lady gaga!
Your personal stuffs outstanding. All the time handle
it up!
I’m not sure why but this web site is loading extremely slow for me. Is anyone else having this problem or is it a problem on my end? I’ll check back later on and see if the problem still exists.
I have been browsing online greater than 3 hours nowadays, yet I never discovered any fascinating article like yours.
It is beautiful value sufficient for me. In my view, if all
webmasters and bloggers made excellent content as you did,
the net shall be much more helpful than ever before.
https://download-install.com/entry/알약-무료-다운로드-설치방법-공개용-기업용
That is very interesting, You are an excessively skilled blogger.
I have joined your rss feed and look ahead to in the hunt
for extra of your fantastic post. Also, I have shared your website in my social networks
Hi there very nice web site!! Man .. Excellent .. Amazing ..
I’ll bookmark your site and take the feeds also? I’m satisfied to find numerous useful
information right here within the submit, we want develop more strategies in this regard,
thanks for sharing. . . . . .
https://bystarlight.tistory.com/765
На сайте https://lemon-car.ru/ укажите номер телефона для того, чтобы воспользоваться услугами проверенного, надежного автосервиса, который работает на выгодных условиях. На все услуги даются гарантии, потому как в компании работают лучшие и компетентные специалисты, знающие свою работу. Прямо сейчас вы сможете записаться на интересующую услугу. Для вас доступен кузовной, слесарный ремонт, двигателя, а также АКПП. Оказывается полный комплекс услуг. В первую очередь, осуществляется диагностика автомобиля, а потом определяется стоимость ремонта.
Greate post. Keep posting such kind of info on your blog.
Im really impressed by your blog.
Hi there, You have performed an incredible job. I’ll certainly digg it and in my view recommend to my friends.
I am confident they’ll be benefited from this site.
It’s really very complex in this active life to listen news on Television, thus
I just use the web for that purpose, and obtain the
newest information.
Hey there, I think your site might be having browser compatibility issues.
When I look at your blog site in Opera, it looks fine but when opening in Internet Explorer,
it has some overlapping. I just wanted to give you a quick heads up!
Other then that, superb blog!
https://dday.tistory.com/1210
You should be a part of a contest for one of the finest blogs online.
I am going to recommend this website!
I keep listening to the news broadcast speak about receiving free online grant applications so I have been looking around for the most excellent site to get one. Could you tell me please, where could i get some?
hi!,I like your writing so much! share we communicate more about your post on AOL? I need a specialist on this area to solve my problem. Maybe that’s you! Looking forward to see you.
My brother suggested I might like this website. He used to be totally right. This post truly made my day. You cann’t believe just how a lot time I had spent for this info! Thank you!
Good day! This is my first comment here so I just wanted to give a quick shout out and say I genuinely enjoy reading through your posts. Can you suggest any other blogs/websites/forums that cover the same topics? Thanks a lot!
I do accept as true with all of the ideas you have offered for your post.
They’re really convincing and can definitely work. Nonetheless, the posts are too quick for beginners.
May just you please lengthen them a bit from subsequent time?
Thanks for the post.
Howdy this is kinda of off topic but I was wanting to know if blogs use WYSIWYG editors or if you have to manually code with HTML. I’m starting a blog soon but have no coding skills so I wanted to get guidance from someone with experience. Any help would be enormously appreciated!
https://king.creaming.net/on/2597
Sex
https://ddnews.co.kr/4th-fund/
На сайте http://www.tribal-tattoo.ru ознакомьтесь с тем, какими услугами вы сможете воспользоваться в тату-салоне. Есть возможность заказать перманентный татуаж губ, в том числе, контур с растушевкой, татуаж бровей. К вашим услугам исправление татуировок, нанесение новой тату, чтобы замаскировать старую. Также доступно перекрытие шрамов, рубцов. Рисунки подбираются в зависимости от предпочтений, а также особенностей. При необходимости мастер выберет для вас оптимальный вариант. Услуга абсолютно безопасная.
My brother recommended I may like this blog. He was
entirely right. This submit actually made my day. You
cann’t believe simply how a lot time I had spent for this info!
Thank you!
nice article visite https://braceletwithpictureinside.store/ now
I relish, lead to I discovered just what I used to be having a look for.
You’ve ended my 4 day long hunt! God Bless you man. Have a nice
day. Bye
I think that everything typed made a bunch of sense. But, consider this, what if you were to create a killer headline?
I ain’t suggesting your content is not solid, however what if you added something that makes people
desire more? I mean Fornix (crus)/ Stria Terminalis – fxst |
Whole Brain Protocol for Tractography with Empirical MRI is kinda
boring. You should glance at Yahoo’s home page and note how
they create article titles to grab viewers to open the links.
You might add a related video or a related picture or two to
get people excited about everything’ve written. Just my opinion,
it would make your posts a little livelier.
Hmm is anyone else having problems with the images on this blog loading?
I’m trying to figure out if its a problem on my end or if it’s the blog.
Any suggestions would be greatly appreciated.
Cool blog! Is your theme custom made or did you download
it from somewhere? A theme like yours with a few simple adjustements would really make my blog stand out.
Please let me know where you got your theme.
With thanks
Great article, totally what I was looking for.
I always used to study post in news papers but now as I am a
user of net thus from now I am using net for articles, thanks to web.
sandyterrace.com
지금… 적어도 상황은 그렇게 나쁘지는 않습니다.
Woah! I’m really digging the template/theme of this blog. It’s simple, yet effective. A lot of times it’s very hard to get that “perfect balance” between user friendliness and visual appearance. I must say you have done a very good job with this. Also, the blog loads extremely fast for me on Internet explorer. Superb Blog!
I was recommended this blog by way of my cousin. I
am not certain whether this post is written by him as nobody else understand such precise about my difficulty.
You’re incredible! Thank you!
nice article visite https://rainbowhighdolls.store/ now
Howdy! I know this is kinda off topic however I’d figured
I’d ask. Would you be interested in exchanging links or maybe guest writing a blog article or vice-versa?
My website covers a lot of the same subjects as yours
and I believe we could greatly benefit from each other.
If you might be interested feel free to send me an email.
I look forward to hearing from you! Awesome blog by the way!
each time i used to read smaller posts that as well clear their motive, and that is also happening with this paragraph which I am reading now.
Scam
Hi everyone, it’s my first pay a quick visit at this site,
and piece of writing is actually fruitful in support of me,
keep up posting such content.
Thanks on your marvelous posting! I definitely enjoyed reading it, you
could be a great author.I will be sure to bookmark your blog and will come
back later on. I want to encourage continue your great posts, have
a nice afternoon!
https://honeytipit.tistory.com/tag/넷마블20사천성20바로가기
https://pornmaster.fun/hd/chilldarn-mp4kamwali-bai-down-blouse-showing-bobs
One thing I’d like to say is car insurance termination is a horrible experience and if you’re doing the appropriate things like a driver you simply won’t get one. Some individuals do obtain the notice that they’ve been officially dumped by their particular insurance company they have to fight to get further insurance after having a cancellation. Inexpensive auto insurance rates tend to be hard to get from a cancellation. Knowing the main reasons with regard to auto insurance cancellations can help car owners prevent losing one of the most important privileges readily available. Thanks for the suggestions shared by means of your blog.
Asking questions are truly fastidious thing if you are not understanding something fully,
but this paragraph presents pleasant understanding even.
На сайте http://kinomanhd720.online находятся самые популярные, интересные и увлекательные фильмы на разный вкус и различного жанра: драмы, комедии, триллеры, ужасы и многое другое. Вы обязательно найдете то, что посмотреть сегодня вечером, чтобы разнообразить досуг. Все они представлены в отличном качестве и с хорошим звуком. Также имеются и сериалы, мультфильмы, мистические фильмы, которые точно никого не оставят равнодушным. Специально для вас есть подборки про школу, врачей, вампиров, войну, любовь.
I have realized that in old digital cameras, unique receptors help to maintain focus automatically. Those kind of sensors of some video cameras change in in the area of contrast, while others employ a beam involving infra-red (IR) light, particularly in low lumination. Higher standards cameras sometimes use a blend of both systems and will often have Face Priority AF where the digicam can ‘See’ a new face and concentrate only on that. Thanks for sharing your ideas on this weblog.
Hey there! I’m at work browsing your blog from my new iphone 4!
Just wanted to say I love reading through your blog and
look forward to all your posts! Keep up the fantastic work!
It’s difficult to find well-informed people about this subject, but you seem like you know
what you’re talking about! Thanks
I could not resist commenting. Very well written!
This text is worth everyone’s attention. Where can I find out more?
Would you be thinking about exchanging links?
It is not my first time to visit this website, i am browsing this site
dailly and obtain fastidious data from here all the time.
What’s up i am kavin, its my first occasion to commenting anywhere, when i read
this paragraph i thought i could also create comment due to this sensible paragraph.
nice article visite https://motionsensorlightbulb.store/ now
shop gifts onhttps://motionsensorlightbulb.store/ now
game1kb.com
이 신사들은 비오는 날을 계획하고 있다는 말입니까?
The meal went without incident although a couple of times she had to avert her gaze from Dan for fear of bursting out laughing. Dan went up to his room saying he wanted to listen to music, he laid on his bed playing with himself waiting for his father to leave. He thought about the events of the last two weeks it had been the craziest time of his life.
He put it all down to Mary Harris, she had been his girlfriend for two years, he liked her a lot, she was pretty and intelligent two things that rarely combined in his experience, the problem was she steadfastly refused to let him put his cock inside her, he had thought he was getting somewhere when he took his cock out that evening in the cinema when she had finally allowed him to get his hand inside her panties, it was the first time she had allowed him to do any more than play with her tits. He had taken her hand and placed it on his erection, initially he had been pleased to hear her sharp intake of breath when she felt the size of him and he was encouraged when she began stroking him.
It was later that the problem started, he had her in the car park, they were kissing, her sweater was pushed up together with her bra and he was sucking on her nipples. He managed to get her laid across the front of one of the cars, his hand up her skirt, trying to pull down her panties. That was when she had stopped him.
https://okwave.jp/profile/u3107642.html
http://www.nfomedia.com/profile?uid=rOiThgG
https://chyoa.com/user/alure6661968
https://www.brollopsguiden.se/medlemspresentation/86094
https://onedio.ru/profile/xkiskax-195-1
https://www.quia.com/profiles/chspading
https://launchpad.net/~spielhur19641
https://rentry.org/x77tbxye
https://www.cossa.ru/profile/?ID=238973
https://www.divephotoguide.com/user/seeside1988
I like the valuable info you supply on your articles.
I will bookmark your blog and check once more right here frequently.
I am relatively certain I’ll be informed many new stuff proper right
here! Best of luck for the following!
What a material of un-ambiguity and preserveness of valuable know-how on the topic
of unpredicted feelings.
Hay
Excellent blog here! Also your site loads up fast! What web host are
you using? Can I get your affiliate link to your host? I wish my
site loaded up as fast as yours lol
Thanks so much for giving everyone an extremely marvellous possiblity to read from this blog. It can be so great and as well , stuffed with a great time for me and my office peers to search your blog particularly 3 times in 7 days to find out the new guides you have. Not to mention, we are usually amazed for the surprising principles you serve. Some two facts in this posting are in fact the most efficient we’ve ever had.
The very root of your writing while appearing agreeable originally, did not really sit properly with me after some time. Somewhere within the paragraphs you were able to make me a believer unfortunately only for a short while. I however have a problem with your leaps in assumptions and you would do nicely to fill in all those breaks. When you can accomplish that, I would surely be impressed.
Link exchange is nothing else except it is simply placing the other person’s website link
on your page at proper place and other person will also do same
in support of you.
Hi there, its fastidious post about media print, we all
understand media is a enormous source of data.
Very shortly this web page will be famous among all blogging
and site-building visitors, due to it’s nice content
https://itmanual.net/EC9791EC8580-ED858DEC8AA4ED8AB8-EB8298EB8884EAB8B0-EC9791EC8580-EC8580-EBB684ED95A0-EBB0A9EBB295/
“비아그라(Viagra)”와 “시알리스(Cialis)”는 모두 남성의 발기 기능 장애(ED, 남성 성기능 부전)를 치료하는 데에 사용되는 약물이지만, 그들 사이에는 몇 가지 차이점이 있습니다.
https://www.vianafil.com
Inspiring story there. What occurred after?
Thanks!
https://mintfin.tistory.com/tag/넷마블20장기20바로가기
I really like what you guys are usually up too.
This sort of clever work and exposure! Keep up the excellent works guys I’ve incorporated you guys to blogroll.
Amazing! Its really amazing paragraph, I have got much clear idea regarding from
this piece of writing.
It’s going to be ending of mine day, however before finish I am
reading this enormous paragraph to improve my knowledge.
Wow, wonderful blog layout! How long have you been blogging for?
you made blogging look easy. The overall look of your website is great, as well as the content!
This is my first time go to see at here and i am genuinely impressed to read
everthing at alone place.
https://chotiple.tistory.com/tag/nPDF20유틸20도구
https://nicesongtoyou.com/welfare/infants/
Very great post. I simply stumbled upon your weblog and wished
to say that I’ve truly loved surfing around your blog posts.
After all I’ll be subscribing in your feed and I am hoping you write once more soon!
Today, I went to the beach front with my kids.
I found a sea shell and gave it to my 4 year old daughter and said “You can hear the ocean if you put this to your ear.” She placed the shell to
her ear and screamed. There was a hermit crab inside and it pinched her ear.
She never wants to go back! LoL I know this is totally off topic but I had to tell someone!
Marvelous, what a web site it is! This webpage provides useful facts to us, keep it
up.
We are a group of volunteers and starting a new scheme in our
community. Your site offered us with valuable information to work on. You have done
a formidable job and our whole community will be thankful to you.
Appreciation to my father who told me concerning this weblog, this website is in fact remarkable.
Hello! I could have sworn I’ve been to this web site before but
after browsing through a few of the posts I realized
it’s new to me. Nonetheless, I’m certainly happy I discovered it and I’ll be book-marking it and checking back often!
This text is invaluable. When can I find out more?
Hurrah, that’s what I was looking for, what a information! existing here at this webpage,
thanks admin of this site.
This post is worth everyone’s attention. Where can I find out more?
After exploring a handful of the articles on your web site,
I truly like your way of writing a blog. I saved
it to my bookmark website list and will be checking back
in the near future. Please check out my website as well and let me know how you feel.
I know this website gives quality depending posts and extra information,
is there any other web site which provides these kinds of data in quality?
Very energetic blog, I loved that bit. Will there be a part 2?
Hi i am kavin, its my first occasion to commenting anyplace,
when i read this paragraph i thought i could also create comment due to this good
article.
На сайте https://bdalogistic.ru/ вы сможете заказать товары для маркетплейсов, которые привозятся из Китая. Основная специализация компании заключается в поиске, приобретении, доставке продукции. Кроме того, выполняется и отгрузка, упаковка продукции на маркетплейсы. В компании работают компетентные, опытные и предусмотрительные специалисты, которые окажут поддержку в организации бизнеса на Вайлдберриз. Рассчитать цену услуги получится прямо сейчас. В любом случае она будет доступной.
I read this piece of writing fully on the topic of
the comparison of latest and previous technologies, it’s
awesome article.
https://kimyunjeong.com/629
With havin so much content and articles do you ever run into any problems of plagorism or
copyright violation? My website has a lot of exclusive content I’ve either authored myself or outsourced but it seems a lot of it
is popping it up all over the web without my agreement. Do you know any techniques to help reduce
content from being stolen? I’d truly appreciate it.
We are a group of volunteers and starting a new scheme in our community. Your web site offered us with valuable information to paintings on. You’ve done an impressive task and our entire community can be thankful to you.
Right now it appears like Expression Engine is the preferred blogging platform available right now.
(from what I’ve read) Is that what you’re using on your blog?
Thanks for the tips on credit repair on your web-site. Things i would tell people would be to give up a mentality that they’ll buy at this point and fork out later. Being a society we tend to do that for many things. This includes family vacations, furniture, plus items we want. However, you need to separate your current wants from all the needs. As long as you’re working to raise your credit ranking score you really have to make some sacrifices. For example you possibly can shop online to economize or you can go to second hand shops instead of high priced department stores regarding clothing.
Its such as you read my thoughts! You seem to know a lot approximately this, such
as you wrote the ebook in it or something. I feel that you just can do with a few
percent to drive the message house a bit, however instead of that, this is fantastic blog.
A fantastic read. I will certainly be back.
Fabulous, what a blog it is! This webpage presents helpful data
to us, keep it up.
This paragraph will assist the internet
users for setting up new blog or even a weblog from start to end.
Hey there! This post could not be written any better! Reading through this post reminds me of my previous room mate!
He always kept chatting about this. I will forward this page to him.
Pretty sure he will have a good read. Thank you for
sharing!
Российский производитель предлагает блины на ресурсе https://diski-dlya-shtang.ru для напряженной эксплуатации в коммерческих тренировочных залах и в домашних условиях. Завод из России производит тренировочные диски разного посадочного диаметра и любого востребованного веса для разборных штанг. Рекомендуем к покупке обрезиненные тренировочные блины для силовых тренировок. Они не скользят, не гремят и более безопасны. Производимые изделия не нуждаются в постоянном обслуживании и ориентированы на длительную эксплуатацию в клубах. Предлагаем внушительный каталог бамперных дисков с разным видом защитного покрытия. Оформите веса с необходимой массой и посадочным диаметром по недорогим ценам напрямую у производителя.
Great website! I am loving it!! Will be back later to read some more. I am taking your feeds also.
I have read so many content regarding the blogger
lovers but this article is truly a pleasant paragraph, keep it up.
Found an article that is worth reading – it’s really interesting! http://colonia.rusff.me/viewtopic.php?id=152#p3047
I got this site from my friend who shared with me concerning this site and at
the moment this time I am browsing this website and
reading very informative content at this time.
I have been absent for some time, but now I remember why I used to love this site. Thanks , I will try and check back more often. How frequently you update your site?
Please let me know if you’re looking for a article writer for your blog. You have some really good posts and I feel I would be a good asset. If you ever want to take some of the load off, I’d really like to write some content for your blog in exchange for a link back to mine. Please blast me an email if interested. Thank you!
“Watchers” introduces us to Travis Cornell, a former Delta Force operator who has become disillusioned with life. He spends his days in seclusion in the canyons of California, finding solace only in the occasional company of his faithful dog. His solitary existence is shattered one day by an unexpected encounter with a remarkably intelligent golden retriever — a dog that seems to possess an uncanny understanding of the dangers lurking in the shadows.
I read this piece of writing completely concerning the resemblance of most up-to-date and preceding technologies, it’s awesome article.
In Dean Koontz Intensity We meet Chyna Shepherd, a young woman with a vibrant spirit and a resilience forged by a difficult past. Seeking respite from the pressures of her life, she decides to spend a weekend relaxing at her friend’s isolated farmhouse in the picturesque Napa Valley. Unbeknownst to Chyna, this idyllic setting will become a claustrophobic battleground where she must confront the darkest corners of human nature.
Российский изготовитель предлагает тренировочные диски в интернет-магазине https://diski-dlya-shtang.ru для усиленной работы в коммерческих тренировочных центрах и в домашних условиях. Отечественный завод производит тренировочные блины разного посадочного диаметра и любого востребованного веса для сборных штанг и гантелей. Советуем к приобретению прорезиненные тренировочные блины для силовых тренировок. Они не скользят, не гремят и более безопасны. Производимые изделия не нуждаются в постоянном обслуживании и ориентированы на длительную работу в центрах. Реализуем большой ассортимент любительских дисков с разным видом защитного покрытия. Закажите веса с нужной массой и посадочным диаметром по недорогим ценам напрямую у производителя.
Your way of describing everything in this article is actually pleasant,
all can without difficulty understand it, Thanks a
lot.
hello there and thank you for your information – I have certainly picked up something new from right here.
I did however expertise several technical issues using this web site,
as I experienced to reload the web site lots of times previous
to I could get it to load properly. I had been wondering if your web host is OK?
Not that I’m complaining, but sluggish loading instances
times will often affect your placement in google and can damage your quality score if advertising
and marketing with Adwords. Well I’m adding this RSS to my email and can look out for
a lot more of your respective fascinating content. Ensure that you update this again very soon.
If some one wishes to be updated with hottest technologies after
that he must be pay a quick visit this site and be up to date everyday.
Teaches the fundamentals of self-hypnosis: The book provides clear guidance on the techniques and principles involved in effective self-hypnosis.
Heya i am for the first time here. I found this board and I find It really useful & it helped me out much.
I hope to give something back and help others like you helped me.
Wow, this article is pleasant, my sister is analyzing these kinds of
things, so I am going to inform her.
Hi! I could have sworn I’ve been to this site
before but after going through many of the articles I realized
it’s new to me. Anyhow, I’m certainly pleased I found it and I’ll
be book-marking it and checking back regularly!
Thanks for a marvelous posting! I genuinely enjoyed
reading it, you’re a great author. I will always bookmark your
blog and may come back someday. I want to encourage you to
continue your great writing, have a nice holiday
weekend!
Hmm it seems like your website ate my first comment (it
was super long) so I guess I’ll just sum it up what I submitted and say, I’m thoroughly enjoying your blog.
I as well am an aspiring blog writer but I’m still
new to the whole thing. Do you have any suggestions for novice blog writers?
I’d definitely appreciate it.
I’m gone to tell my little brother, that he should also pay a visit this blog on regular
basis to get updated from most recent news.
First of all I would like to say awesome blog! I had a
quick question that I’d like to ask if you don’t mind. I was curious
to know how you center yourself and clear your thoughts before writing.
I’ve had a difficult time clearing my mind in getting my thoughts out.
I truly do enjoy writing but it just seems like the first 10 to 15 minutes are generally wasted just trying
to figure out how to begin. Any ideas or hints? Thank you!
You’re so awesome! I do not believe I’ve truly read anything
like this before. So wonderful to find someone with some genuine thoughts on this subject.
Seriously.. thank you for starting this up. This website is
one thing that is required on the web, someone with some originality!
Hey there, I think your website might be having browser compatibility issues.
When I look at your blog in Chrome, it looks fine but when opening in Internet
Explorer, it has some overlapping. I just wanted to give you a quick heads up!
Other then that, amazing blog!
The central theme of Atomic Habits is that massive success doesn’t come from huge leaps, but rather from small, consistent improvements. Just as atoms are the building blocks of matter, seemingly insignificant “atomic habits” have the power to compound into extraordinary outcomes.
qiyezp.com
이 재무 보고서는 Fang Jifan의 여동생 Fang Xiaofan이 계산했습니다.
you’re truly a excellent webmaster. The website loading pace is incredible.
It sort of feels that you are doing any distinctive trick.
Furthermore, The contents are masterwork. you have done a magnificent process in this topic!
What’s up, everything is going well here and ofcourse every one is sharing information, that’s in fact excellent, keep
up writing.
Thanks very interesting blog!
Valuable information. Lucky me I discovered your site accidentally, and I’m shocked why
this accident did not took place earlier! I bookmarked it.
Spot on with this write-up, I actually feel this web site needs a great deal more
attention. I’ll probably be back again to read more, thanks for the information!
It is not my first time to pay a visit this web site, i am browsing this web
page dailly and obtain pleasant facts from here every day.
I have read so many posts on the topic of the blogger lovers except this article is actually a pleasant paragraph, keep it up.
Fantastic items from you, man. I have have in mind your stuff prior to and you are just too excellent.
I actually like what you have obtained right here, really like what you’re stating and
the best way during which you say it. You’re
making it enjoyable and you continue to care for to stay it sensible.
I can’t wait to read much more from you. This is actually
a terrific web site.
https://honeytiplabs.com/아이폰-중복-사진/
Sweet blog! I found it while searching on Yahoo News. Do you have any suggestions on how to
get listed in Yahoo News? I’ve been trying for a while but
I never seem to get there! Appreciate it
I simply couldn’t go away your web site prior to suggesting that I actually loved the
usual information a person provide to your visitors?
Is gonna be back continuously in order to inspect new posts
Hi there, after reading this remarkable piece of writing i am
as well delighted to share my experience here with friends.
Admiring the hard work you put into your blog and detailed information you provide.
It’s good to come across a blog every once in a while that isn’t the same unwanted rehashed information. Great read!
I’ve bookmarked your site and I’m adding your RSS feeds to my Google account.
The meal went without incident although a couple of times she had to avert her gaze from Dan for fear of bursting out laughing. Dan went up to his room saying he wanted to listen to music, he laid on his bed playing with himself waiting for his father to leave. He thought about the events of the last two weeks it had been the craziest time of his life.
He put it all down to Mary Harris, she had been his girlfriend for two years, he liked her a lot, she was pretty and intelligent two things that rarely combined in his experience, the problem was she steadfastly refused to let him put his cock inside her, he had thought he was getting somewhere when he took his cock out that evening in the cinema when she had finally allowed him to get his hand inside her panties, it was the first time she had allowed him to do any more than play with her tits. He had taken her hand and placed it on his erection, initially he had been pleased to hear her sharp intake of breath when she felt the size of him and he was encouraged when she began stroking him.
It was later that the problem started, he had her in the car park, they were kissing, her sweater was pushed up together with her bra and he was sucking on her nipples. He managed to get her laid across the front of one of the cars, his hand up her skirt, trying to pull down her panties. That was when she had stopped him.
https://quern1987.diary.ru/
https://www.divephotoguide.com/user/tsunaesi1963
https://www.quia.com/profiles/ribowers
https://kozo4ka1969.diary.ru/
https://launchpad.net/~dions19801
https://cannabis.net/user/146915
https://chyoa.com/user/kreig971974
https://www.divephotoguide.com/user/bizy19621999
https://bellboy1995.micro.blog/about/
https://www.metal-archives.com/users/morello19831982
It’s going to be end of mine day, however before
finish I am reading this impressive article to increase my know-how.
Hi, yup this article is really fastidious and I
have learned lot of things from it concerning blogging. thanks.
Howdy! This is my first visit to your blog! We are a
collection of volunteers and starting a new project in a community in the same niche.
Your blog provided us valuable information to work on. You have done a marvellous job!
I quite like reading a post that will make men and
women think. Also, thank you for allowing for me to comment!
Добрый день всем!
Приобретите диплом университета России без предоплаты и с гарантией качества доставки в любую точку страны!
http://saksx-attestats.ru/
Приобретите российский диплом с гарантией качества и доставкой в любую точку страны без предварительной оплаты!
Наши услуги помогут вам приобрести диплом ВУЗа с доставкой по всей России без предварительной оплаты – быстро и надежно!
Discovered an article that might interest you Р don’t miss it! https://www.cardiachill.com/users/cchatruletka
With havin so much content do you ever run into any
problems of plagorism or copyright infringement?
My blog has a lot of unique content I’ve either authored myself or outsourced but it
appears a lot of it is popping it up all over the internet without my authorization. Do you know any
techniques to help reduce content from being ripped off?
I’d certainly appreciate it.
https://sharply.tistory.com/844
When some one searches for his vital thing, so he/she needs to be available that
in detail, therefore that thing is maintained over here.
Thanks , I have just been looking for info about this topic for a long time and
yours is the best I’ve found out so far. However, what in regards
to the bottom line? Are you sure concerning the source?
Link exchange is nothing else however it is only placing the other
person’s website link on your page at suitable place and other person will also do same
in favor of you.
Write more, thats all I have to say. Literally, it seems as though you relied on the video to make your point. You obviously know what youre talking about, why throw away your intelligence on just posting videos to your blog when you could be giving us something enlightening to read?
It’s actually a great and useful piece of info. I am glad that you shared
this helpful information with us. Please keep us informed like this.
Thanks for sharing.
Thankfulness to my father who informed me concerning this blog, this web site is genuinely awesome.
Thanks for finally talking about > Fornix (crus)/ Stria Terminalis
– fxst | Whole Brain Protocol for Tractography with
Empirical MRI < Liked it!
When I originally commented I clicked the “Notify me when new comments are added” checkbox and now each time
a comment is added I get several e-mails with the same comment.
Is there any way you can remove people from that service?
Bless you!
Spot on with this write-up, I really suppose this website needs rather more consideration. I抣l in all probability be again to learn far more, thanks for that info.
Great goods from you, man. I have understand your stuff previous to and
you’re just too magnificent. I actually like what you’ve acquired here,
certainly like what you are saying and the way in which
you say it. You make it enjoyable and you still care for to keep it wise.
I can not wait to read much more from you. This is
actually a tremendous site.
Hi there I am so glad I found your web site, I really found you by accident, while I was browsing on Bing for something else, Nonetheless I am here now and would just like to say thanks a lot for a fantastic post and a all round exciting blog (I also love the theme/design), I don抰 have time to browse it all at the minute but I have book-marked it and also added your RSS feeds, so when I have time I will be back to read much more, Please do keep up the excellent job.
Anti-Forensics involves the use to tools and techniques used to frustrate a digital forensics investigation.
Hi! I could have sworn I’ve been to this blog before but after browsing
through a few of the posts I realized it’s new to me.
Anyways, I’m definitely pleased I discovered it and I’ll be book-marking it and
checking back often!
Hello there, just became alert to your blog through Google, and found that it is truly informative.
I am going to watch out for brussels. I’ll appreciate if you continue
this in future. Numerous people will be benefited from your writing.
Cheers!
Wonderful blog! I found it while surfing around on Yahoo News.
Do you have any tips on how to get listed in Yahoo News?
I’ve been trying for a while but I never seem to
get there! Cheers
비아그라 * 시알리스 * 레비트라
찾으시는 제품 모두 현재 1+1으로 하나 더
서비스 비아그라 * 시알리스 * 레비트라 * 중 한가지 4정 더
▶ 1+1 최저가 사러가기 클릭
비아그라구매
You really make it seem so easy with your presentation but I find this matter to be actually something which I
think I would never understand. It seems too complex and very broad for me.
I’m looking forward for your next post, I will try to
get the hang of it!
I am genuinely grateful to the owner of this site who has shared this
impressive piece of writing at at this time.
nice article visite https://necklacewithpictureinside.com/ now
https://www.monkfishjowls.com/피자스쿨-영업시간-정확한-시간
Its such as you learn my thoughts! You seem to understand a lot approximately this, like you wrote the ebook in it or something. I believe that you could do with some p.c. to pressure the message home a little bit, but other than that, that is magnificent blog. An excellent read. I will definitely be back.
nice article visite https://mensjewelrybox.store/ now
Together with every little thing which appears to be building throughout this subject matter, all your perspectives are generally fairly refreshing. However, I beg your pardon, but I do not subscribe to your whole theory, all be it radical none the less. It seems to everybody that your commentary are generally not totally validated and in fact you are generally your self not really wholly convinced of your point. In any event I did appreciate looking at it.
Generally I don’t learn post on blogs, however I would like to say that this write-up very forced me to take a look at and do it!
Your writing style has been surprised me. Thanks, very nice article.
nice article visite https://gpsdogfence.store/ now
shop gifts on https://beanbagbed.store/ now
Hey there! Do you know if they make any plugins to protect against hackers?
I’m kinda paranoid about losing everything I’ve worked hard on. Any tips?
Добрый день всем!
Приобретите диплом Гознака у нас по специальной цене с доставкой в любой регион России без предоплаты.
купить аттестат
Приобретите диплом колледжа недорого с доставкой по РФ и гарантией качества без предварительной оплаты – просто и надежно!
Закажите диплом Вуза с гарантией качества и доставкой по всей России без предоплаты.
A. 카카오택시 앱을 실행한 후 ‘예약’ 탭을 누르시면 됩니다. 이후 출발 위치, 도착 위치, 탑승 시간 등을 입력하고 ‘예약하기’를 누르시면 예약이 완료됩니다. 다음 사이트에서 섹스 비디오를 시청하세요 강남출장섹스마사지
Nice blog right here! Also your site loads up fast!
What host are you the use of? Can I am getting
your affiliate link on your host? I wish my website
loaded up as fast as yours lol
Добрый день!
Диплом продолжает быть моим вызовом, но теперь у меня есть все необходимые ресурсы для его успешного завершения.
Выберите и приобретите диплом ВУЗа России недорого, качественно и с отправкой почтой!
Желаю вам всем отличных оценок!
где купить диплом
купить диплом в орле
купить диплом в химках
купить диплом сварщика
купить диплом повара-кондитера
купить диплом в миассе
Привет всем!
Написание диплома по ночам ухудшило мое самочувствие, но я не сдаюсь перед лицом трудностей.
Хотите приобрести диплом ВУЗа недорого без предоплаты на нашем сайте? Доставляем в любую точку России.
Желаю всем отличных оценок!
https://rudiplomirovana.com
купить диплом в архангельске
купить диплом в петропавловске-камчатском
купить диплом о высшем образовании
купить диплом в старом осколе
купить диплом в тольятти
Приветики, дорогие мои!!
Изучение найденных мною материалов для диплома рекомендую всем, кто сталкивается с аналогичными трудностями.
На нашем сайте вы можете недорого заказать и купить диплом с гарантией доставки в любой регион РФ.
https://premialnie-diplomiki.com/
Желаю для каждого положительных оценок!
купить диплом в альметьевске
купить диплом в березниках
купить диплом в шахтах
куплю диплом о высшем образовании
купить диплом в хабаровске
Доброго дня!
Диплом заставил меня работать ночами, что угрожало моему здоровью.
На нашем сайте вы можете купить диплом ВУЗа с постоплатой и помощью 24/7.
купить диплом фармацевта
Желаю всем отличных оценок!
купить диплом физика
купить диплом в бердске
купить диплом электромонтажника
купить диплом в кстово
купить диплом в нефтекамске
I’m not sure exactly why but this web site is loading incredibly slow for me.
Is anyone else having this issue or is it a issue on my end?
I’ll check back later on and see if the problem still exists.
Awesome article.
I cling on to listening to the news lecture about getting free online grant applications so I have been looking around for the best site to get one. Could you tell me please, where could i get some?
Привет всем!
Диплом перестал казаться непреодолимым благодаря помощи, полученной из интернета.
Предлагаем всем желающим приобрести диплом университета России по выгодной цене с доставкой “под ключ”.
Желаю всем честных оценок!
купить диплом техникума
купить диплом в хасавюрте
куплю диплом о высшем образовании
купить диплом юриста
купить диплом воспитателя
купить диплом в твери
Привет всем!
Диплом продолжает развиваться благодаря различным онлайн-ресурсам.
Выберите и заказывайте диплом ВУЗа России качественно и с возможностью отправки почтой!
Желаю для каждого отличных оценок!
https://eonline-diplomy.com
купить диплом в губкине
купить диплом в барнауле
купить диплом штукатура
купить диплом юриста
купить диплом в старом осколе
The practice of timestomping involves the deliberate alteration of these timestamps, reshaping the perceived chronology of actions and potentially obscuring the true sequence of events.
Do you mind if I quote a couple of your posts as long as I provide credit and sources back to your site? My blog is in the exact same niche as yours and my users would truly benefit from some of the information you present here. Please let me know if this alright with you. Appreciate it!
It’s genuinely very difficult in this full of activity life to listen news
on TV, so I only use internet for that reason, and
get the latest information.
Wow, incredible weblog structure! How long have you ever been blogging for?
you made blogging glance easy. The overall look of your web
site is great, as smartly as the content!
You can see similar here sklep online
nice article visite https://bluetoothhearingaids.store/ now
Здравствуйте!
Поиск ресурсов для диплома в интернете значительно улучшил мою работу и облегчил задачу.
Недорого и без проблем приобрести диплом института, техникума, колледжа выгоднее всего у нас, с доставкой по РФ.
Желаю всем отличных оценок!
купить диплом о высшем образовании
купить диплом в дзержинске
купить диплом медбрата
купить диплом в каменске-уральском
купить диплом в нижнекамске
купить диплом медсестры
Доброго дня!
Диплом медленно, но верно, становится менее затруднительным с помощью интернета.
Желаете приобрести диплом ВУЗа недорого и без предоплаты на нашем сайте? Мы доставим его в любую точку России.
Желаю вам всем отличных оценок!
купить аттестат за 11 класс
куплю диплом кандидата наук
купить диплом в верхней пышме
купить диплом в череповце
купить диплом в благовещенске
купить диплом программиста
Всем хорошего дня!
Диплом перестал казаться непреодолимым благодаря помощи, полученной из интернета.
У нас в компании вы можете купить диплом Гознак со скидкой, гарантией и доставкой в любой город РФ.
https://adiplom-russian.com/
Желаю вам всем пятерочных) оценок!
купить диплом в кинешме
купить диплом в ишимбае
купить диплом в пензе
купить диплом хореографа
купить диплом в петрозаводске
Добрый день!
Прогресс в дипломе обеспечен благодаря использованию онлайн-ресурсов.
Закажите диплом у нас и мы доставим его вам в любой регион России без предоплаты, с гарантированной конфиденциальностью.
https://adiplom-russian.com/
Желаю каждому не двоешных) оценок!
купить диплом машиниста
купить диплом в киселевске
купить диплом дизайнера
купить диплом в подольске
купить диплом в волгодонске
The central theme of Atomic Habits is that massive success doesn’t come from huge leaps, but rather from small, consistent improvements.
“Self-sabotage is what happens when we refuse to consciously meet our innermost needs, often because we do not believe we are capable of handling them.”
David Goggins begins his story in the depths of suffering. His childhood was one marked by extreme abuse, a perpetually absent father, and a mother working under dire conditions to keep the family afloat.
I don’t even understand how I finished up right here, but I
assumed this publish was good. I do not realize
who you might be however certainly you’re going to a well-known blogger for those who aren’t
already. Cheers!
asdf
Woah! I’m really digging the template/theme of this blog.
It’s simple, yet effective. A lot of times it’s hard to
get that “perfect balance” between user friendliness and appearance.
I must say that you’ve done a excellent job
with this. Also, the blog loads very quick for me on Opera.
Excellent Blog!
Hey there just wanted to give you a quick heads up. The
text in your content seem to be running off the screen in Internet explorer.
I’m not sure if this is a format issue or something to do with
web browser compatibility but I figured I’d post
to let you know. The design and style look great though!
Hope you get the issue solved soon. Thanks
great publish, very informative. I wonder why the opposite experts of this sector don’t
understand this. You must continue your writing.
I’m confident, you have a great readers’ base already!
Hmm is anyone else experiencing problems with the pictures on this blog loading?
I’m trying to find out if its a problem on my end or if it’s the
blog. Any suggestions would be greatly appreciated.
I’m curious to find out what blog platform you happen to be working with?
I’m having some minor security issues with my latest blog and I would like
to find something more secure. Do you have any
recommendations?
https://king.creaming.net/on/555
На сайте https://dom-otmostka.ru/ оставьте заявку для того, чтобы заказать такую популярную услугу, как монтаж отмостки из брусчатки, бетона, плитки, камня. Кроме того, оказываются услуги, связанные с обустройством ливневой канализации, а также дренажом. В компании работают компетентные сотрудники, которые произведут монтаж дорожек, детских, парковочных площадок. На любые работы действуют гарантии 3 года. Воспользуйтесь и вы компетентными работами лучших сотрудников. На все услуги установлены привлекательные расценки.
I like it when people come together and share views.
Great website, keep it up!
Today, while I was at work, my cousin stole my iPad and tested to see if it can survive a thirty foot drop, just so she can be a youtube sensation. My iPad is now broken and she has 83 views. I know this is entirely off topic but I had to share it with someone!
Thank you for the auspicious writeup. It if truth be told was once a enjoyment account it. Glance advanced to more added agreeable from you! By the way, how can we be in contact?
Hi there, after reading this awesome article i am also cheerful to share my familiarity here
with mates.
Generally I don’t learn article on blogs, but I wish to say that this write-up very forced me to check out and do it! Your writing style has been amazed me. Thank you, quite great article.
My spouse and I stumbled over here different web address and thought I might as well check things out.
I like what I see so now i’m following you. Look forward
to looking into your web page again.
Greetings! This is my first visit to your blog! We are a team of
volunteers and starting a new project in a community in the same
niche. Your blog provided us useful information to
work on. You have done a outstanding job!
https://mythings.tistory.com/85
My partner and I absolutely love your blog and find a lot of your post’s to be
precisely what I’m looking for. Does one offer guest writers to write content for you
personally? I wouldn’t mind publishing a post or elaborating on a few of
the subjects you write concerning here. Again, awesome site!
Today, while I was at work, my cousin stole my apple ipad and tested to see if it can survive a
30 foot drop, just so she can be a youtube sensation. My iPad is now destroyed and she has 83 views.
I know this is totally off topic but I had to share it with someone!
Just desire to say your article is as astonishing.
The clearness in your post is just nice and i can think you are an expert in this subject.
Well along with your permission allow me to grab your feed to
stay updated with forthcoming post. Thank you 1,000,000
and please continue the gratifying work.
I have been exploring for a bit for any high quality articles or weblog posts on this kind of house .
Exploring in Yahoo I eventually stumbled upon this web site.
Reading this information So i’m satisfied to convey that
I’ve a very just right uncanny feeling I came upon exactly what I needed.
I so much without a doubt will make certain to do not forget
this site and give it a look regularly.
Hello there! This post couldn’t be written any better!
Reading through this post reminds me of my good old room mate!
He always kept chatting about this. I will forward
this article to him. Pretty sure he will have a good read.
Thanks for sharing!
Valuable info. Lucky me I discovered your website accidentally, and
I am surprised why this accident did not took place in advance!
I bookmarked it.
Hey just wanted to give you a quick heads up. The words in your content seem to be running off the screen in Firefox.
I’m not sure if this is a format issue or something to do with web browser compatibility but I thought I’d post to let you know.
The design look great though! Hope you get the issue
resolved soon. Kudos
На сайте https://shemi-otopleniya.ru/ ознакомьтесь с техническими характеристиками, особенностями, нюансами гибкой подводки, выполненной из нержавеющей стали. Так вы сможете ознакомиться с размерами, диаметром, а также различными модификациями. Есть варианты небольших размеров, а также более внушительных. Также можно ознакомиться и с информацией, которая посвящена шаровому крану. Регулярно публикуются любопытные и интересные новости на данную тематику. Обязательно ознакомьтесь и с фотографиями.
На сайте https://www.kapatel.ru/ вы почерпнете много интересной, познавательной информации, которая касается дачи, сада, а также огорода. Вы узнаете о том, как правильно обустроить дачу, огород так, чтобы они были функциональными, вместительными и радовали богатым урожаем. Имеются данные о строительстве, ремонте и различных отделочных работах. Опубликованы материалы о вредителях, различных заболеваниях, о самых цветущих растениях. Вы можете задавать вопросы и присылать сообщения для уточнения некоторых моментов.
Thanks for the auspicious writeup. It if truth be told was once a enjoyment
account it. Look complicated to more delivered agreeable from you!
By the way, how can we keep up a correspondence?
Hi I am so excited I found your web site, I really found you by
error, while I was looking on Bing for something else, Anyhow
I am here now and would just like to say cheers for a fantastic post and a all
round interesting blog (I also love the theme/design), I don’t have time to go
through it all at the moment but I have book-marked it and also added in your RSS feeds, so
when I have time I will be back to read much more, Please do keep up
the great b.
Excellent blog here! Also your web site loads up very fast!
What web host are you using? Can I get your affiliate link to your host?
I wish my website loaded up as quickly as
yours lol
I am really glad to read this blog posts which contains lots of helpful data, thanks for providing these information.
Nice blog here! Additionally your site so much up fast!
What host are you the use of? Can I am getting your associate link to your host?
I want my website loaded up as quickly as yours lol
На сайте https://1-line.su/ воспользуйтесь помощью квалифицированных, компетентных сотрудников популярного предприятия «Первая линия». Они окажут необходимую поддержку в вопросах оптимизации, автоматизации различных бизнес-процессов. Компания сделает все возможное для того, чтобы вы работали наиболее эффективно и продуктивно. Воспользуйтесь полным набором работающих инструментов, которые точно вам пригодятся для построения бизнеса. Для того чтобы начать сотрудничество, просто напишите на электронную почту.
Spot on with this write-up, I actually think this website wants rather more consideration. I抣l probably be once more to learn way more, thanks for that info.
Thanks for your write-up on the travel industry. I’d personally also like to include that if you are a senior thinking of traveling, its absolutely vital that you buy traveling insurance for elderly people. When traveling, golden-agers are at biggest risk of getting a health care emergency. Obtaining right insurance policies package in your age group can protect your health and give you peace of mind.
Thanks for the interesting things you have discovered in your blog post. One thing I would like to touch upon is that FSBO interactions are built over time. By presenting yourself to owners the first weekend their FSBO can be announced, ahead of the masses get started calling on Thursday, you build a good interconnection. By giving them resources, educational resources, free reviews, and forms, you become a good ally. By subtracting a personal desire for them as well as their situation, you generate a solid interconnection that, in many cases, pays off in the event the owners opt with a broker they know and also trust – preferably you.
Thanks a lot for sharing this with all of us you really
recognize what you’re talking about! Bookmarked.
Please also visit my site =). We may have a hyperlink exchange
arrangement among us
Please let me know if you’re looking for a writer for your site.
You have some really good posts and I feel I would be a good asset.
If you ever want to take some of the load off, I’d love to write
some articles for your blog in exchange for a link back to mine.
Please send me an email if interested. Thanks!
Thanks for a marvelous posting! I genuinely enjoyed reading it, you can be
a great author.I will be sure to bookmark your blog and will eventually come back sometime soon. I want to encourage you to continue your
great posts, have a nice morning!
https://gorgopage.com/현대카드-결제일별-이용기간-분실-재발급-해지-고객/
На сайте https://td1000.ru/ находится огромное количество интересных, познавательных статей, которые касаются того, как развить социальные сети и простимулировать количество подписчиков. Также имеются данные о турах по Ямалу. Опубликован материал и о вместительных гардеробных и о том, как подобрать мебель. Есть контент, посвященный различным игровым онлайн-заведениям. Полезные статьи откроют вам глаза на многочисленные вопросы. По этой причине вы узнаете много нового и интересного.
На сайте https://climat.best приобретите качественные, функциональные кондиционеры, сплит-системы, тепловые насосы, а также чиллеры. Вся продукция именитых, проверенных и надежных брендов, а потому техника прослужит долго без потери технических характеристик, возможностей. А если необходимо подобрать что-то определенное, то воспользуйтесь специальным поиском. Перед вами есть и самые популярные позиции, которые пользуются особенным спросом среди покупателей. В интернет-магазине вы отыщите как настенные, так и напольно-потолочные конструкции, кассетные.
You are a very intelligent person!
Привет всем!
Диплом вызвал у меня серьезные сложности из-за пропущенных сроков.
Приобретайте диплом у нас и получите качественный документ без предоплаты с доставкой в любую точку России.
Желаю всем прекрасных отметок!
купить диплом вуза
купить диплом в кунгуре
купить диплом в белорецке
купить диплом моториста
купить диплом в каспийске
купить диплом средне техническое
Доброго дня!
Для диплома интернет стал источником полезных ресурсов, облегчивших задачу.
Мы предлагаем вам купить диплом университета недорого с доставкой в любую точку России, оплата после получения.
Желаю всем честных оценок!
купить диплом специалиста
купить диплом архитектора
купить диплом в коврове
купить аттестат за 11 класс
купить диплом преподавателя
купить аттестат за классов
Приветики, дорогие мои!!
Диплом стал легче благодаря ресурсам, которые я нашел в интернете, ускоряя процесс написания.
Приобретите диплом университета у нас и получите его с доставкой в любую точку России с гарантией качества.
https://rusd-diploms.com/
Желаю всем пятерочных) оценок!
купить диплом техникума
купить диплом менеджера по туризму
купить диплом в майкопе
купить диплом в махачкале
купить диплом в клинцах
Outstanding story there. What occurred after? Take care!
Всем хорошего дня!
Интернет стал незаменимым помощником в написании диплома, обнаружив множество полезных ресурсов.
Закажите диплом у нас – это легко! Без предоплаты, с гарантией качества и доставкой по всей России.
https://rusd-diploms.com/
Желаю всем честных отметок!
купить диплом медицинского училища
купить диплом в междуреченске
купить диплом учителя
купить диплом медсестры
купить диплом судоводителя
I am actually glad to read this website posts which carries
plenty of helpful facts, thanks for providing these kinds of information.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/piter1954/about_me/
https://chyoa.com/user/adirael1952
https://www.hentai-foundry.com/user/allur31r1973/profile
https://rentry.org/ghh3uv55
https://niblers1966.bandcamp.com/album/letting-him-in
Всем хорошего дня!
Несмотря на срыв сроков и угрозу для здоровья, я нашел в интернете ресурсы, которые помогают мне быть более продуктивным.
Закажите диплом у нас легко! Без предоплаты, с гарантией качества и доставкой по всей России.
Желаю всем честных оценок!
купить диплом бакалавра
купить диплом в саратове
купить диплом в находке
где купить диплом среднем
купить диплом в черкесске
купить диплом в азове
Приветики, дорогие мои!!
Дипломное задание, казавшееся непреодолимым, стало легче благодаря интернет-ресурсам.
Приобретите диплом университета с гарантированной доставкой в любой город России с оплатой после получения.
Желаю вам всем не двоешных) оценок!
https://aurus-diplomas.com/
купить диплом в коврове
купить диплом во владивостоке
купить аттестат за 9 классов
купить диплом математика
купить диплом в новоуральске
Hey I am so happy I found your webpage, I really found you by accident, while
I was browsing on Askjeeve for something else,
Nonetheless I am here now and would just like to say cheers for a tremendous
post and a all round interesting blog (I also love the theme/design), I don’t have time to read it
all at the moment but I have book-marked it and also
included your RSS feeds, so when I have time I will be back to read a
lot more, Please do keep up the fantastic job.
https://dday.tistory.com/808
Discovered an article that will definitely interest you – don’t miss the chance to familiarize yourself http://mangorpp.getbb.ru/viewtopic.php?f=3&t=314
Hi, yup this paragraph is really pleasant and I have learned lot of
things from it concerning blogging. thanks.
I know this web page gives quality dependent content and extra material,
is there any other site which provides these kinds
of stuff in quality?
whoah this blog is fantastic i love reading your articles. Keep up the good work! You know, a lot of people are hunting around for this info, you can help them greatly.
I’m impressed, I have to admit. Rarely do I come
across a blog that’s both equally educative and engaging, and
without a doubt, you have hit the nail on the head. The problem is something that not
enough people are speaking intelligently about. I am very
happy I stumbled across this during my search for something
regarding this.
Hi my friend! I wish to say that this post is amazing, nice written and come with almost all
vital infos. I would like to look extra posts like this .
I like the valuable information you provide in your articles.
I’ll bookmark your blog and check again here regularly.
I’m quite sure I will learn plenty of new stuff right here!
Good luck for the next!
Hi it’s me, I am also visiting this website on a regular basis,
this web site is truly good and the people are genuinely sharing fastidious thoughts.
Доброго дня!
Диплом вынудил меня перейти на ночные смены работы, что угрожало моему благополучию.
Приобретайте диплом у нас и получите качественный документ без предоплаты, с доставкой в любую точку России.
Желаю вам всем пятерочных) оценок!
https://diplomys-asx.com
купить диплом старого образца
купить диплом в белогорске
купить диплом в киселевске
купить диплом в каменске-шахтинском
купить дипломы о высшем с занесением
Всем хорошего дня!
Однако, нахождение ресурсов в интернете сделало мою работу над дипломом более эффективной.
Выберите и заказывайте диплом ВУЗа России качественно и с возможностью отправки почтой!
Желаю каждому честных оценок!
https://diplomys-asx.com
старые дипломы купить
купить диплом в горно-алтайске
купить диплом в минеральных водах
купить диплом педагога
купить диплом в южно-сахалинске
I have been browsing online more than 2 hours today,
yet I never found any interesting article
like yours. It is pretty worth enough for me.
In my opinion, if all site owners and bloggers made good content as you did, the internet
will be a lot more useful than ever before.
Hi there friends, how is everything, and what you wish for to say
regarding this paragraph, in my view its really remarkable in favor of me.
Hello there, just became alert to your blog through Google,
and found that it is truly informative. I’m going to watch out for brussels.
I will be grateful if you continue this in future. Numerous people
will be benefited from your writing. Cheers!
Excellent goods from you, man. I have understand
your stuff previous to and you’re just too great.
I really like what you’ve acquired here, really like what you’re saying and the way in which you say it.
You make it enjoyable and you still care for to keep it sensible.
I can not wait to read much more from you. This is really a tremendous
web site.
I’m still learning from you, while I’m trying to achieve my goals. I definitely love reading everything that is posted on your site.Keep the information coming. I loved it!
What’s up to every one, the contents present at this site are actually amazing for people experience, well, keep
up the good work fellows.
Hey there, I think your website might be having browser compatibility issues.
When I look at your website in Opera, it looks fine but when opening in Internet Explorer,
it has some overlapping. I just wanted to give you a quick heads up!
Other then that, awesome blog!
https://kakaotaxi.dasgno.com/kakao-fee
Normally I do not learn article on blogs, however I wish to say that this write-up very compelled me to take a look at and do it! Your writing style has been surprised me. Thank you, quite great article.
Excellent blog! Do you have any suggestions for aspiring writers? I’m hoping to start my own site soon but I’m a little lost on everything. Would you suggest starting with a free platform like WordPress or go for a paid option? There are so many choices out there that I’m totally overwhelmed .. Any suggestions? Kudos!
Hmm it appears like your website ate my first comment (it was extremely long) so
I guess I’ll just sum it up what I wrote and say, I’m thoroughly enjoying your blog.
I too am an aspiring blog writer but I’m still new to the whole thing.
Do you have any points for first-time blog writers? I’d really appreciate
it.
Thank you, I have just been looking for information approximately this
topic for a while and yours is the greatest I’ve found out so far.
However, what concerning the conclusion? Are you sure in regards
to the source?
What a material of un-ambiguity and preserveness
of valuable familiarity about unexpected emotions.
Howdy! Would you mind if I share your blog with my facebook group? There’s a lot of folks that I think would really appreciate your content. Please let me know. Thanks
Приветики, дорогие мои!!
Изучение найденных мною материалов для диплома рекомендую всем, кто сталкивается с аналогичными трудностями.
Приобретите диплом университета у нас и получите его с доставкой в любую точку России с гарантией качества.
Желаю вам всем пятерочных) оценок!
https://russiany-diplomas.com
купить диплом в великом новгороде
купить диплом в новоалтайске
купить диплом в смоленске
купить диплом в новороссийске
купить диплом в костроме
Здравствуйте!
Интернет предложил инструменты для диплома, делая написание более простым и доступным.
Приобретайте диплом университета без предоплаты с доставкой по всей России.
Желаю всем честных оценок!
https://russiany-diplomas.com
купить диплом для иностранцев
купить диплом в буйнакске
купить диплом в первоуральске
купить диплом охранника
купить диплом в ухте
Доброго дня!
Интернет предложил инструменты для диплома, делая написание более простым и доступным.
Приобретение диплома института, техникума или колледжа у нас – это просто, с доставкой по РФ.
Желаю всем не двоешных) отметок!
купить аттестат за 9 класс
купить диплом в люберцах
купить свидетельство о заключении брака
купить диплом в прокопьевске
купить диплом в новокуйбышевске
купить диплом в обнинске
Приветики, дорогие мои!!
Диплом стал для меня огромной проблемой, ведь я сильно задержался со сроками его сдачи.
Мы предлагаем вам купить диплом университета без предоплаты, с гарантированной доставкой по всей России.
Желаю для каждого прекрасных отметок!
где купить диплом
купить диплом мастера маникюра и педикюра
купить диплом в новочеркасске
купить диплом в бору
купить диплом в обнинске
купить диплом в губкине
Right here is the perfect site for anyone who wishes to understand this topic.
You understand a whole lot its almost hard to argue
with you (not that I actually would want to…HaHa).
You certainly put a fresh spin on a topic which has been written about for years.
Excellent stuff, just excellent!
I am really enjoying the theme/design of your web site. Do you ever run into any browser compatibility issues? A small number of my blog visitors have complained about my site not working correctly in Explorer but looks great in Opera. Do you have any solutions to help fix this issue?
Just about all of the things you assert happens to be supprisingly precise and it makes me wonder the reason why I hadn’t looked at this with this light previously. This particular piece really did switch the light on for me as far as this particular issue goes. However there is one particular point I am not too comfy with and while I attempt to reconcile that with the central theme of your point, allow me see just what the rest of your visitors have to point out.Nicely done.
hello there and thank you for your info ?I抳e certainly picked up anything new from right here. I did however expertise some technical issues using this website, as I experienced to reload the web site lots of times previous to I could get it to load correctly. I had been wondering if your web hosting is OK? Not that I am complaining, but sluggish loading instances times will very frequently affect your placement in google and can damage your high-quality score if ads and marketing with Adwords. Anyway I抦 adding this RSS to my email and could look out for a lot more of your respective exciting content. Make sure you update this again soon..
Hey there would you mind letting me know which web host you’re using?
I’ve loaded your blog in 3 different web browsers and
I must say this blog loads a lot quicker then most.
Can you suggest a good web hosting provider at a
reasonable price? Kudos, I appreciate it!
Добрый день!
В интернете я нашел ресурсы для диплома, которые помогли мне ускорить написание.
На нашем сайте вы можете купить диплом ВУЗа, недорого, с оплатой после получения, поддержка 24/7.
Желаю вам всем прекрасных отметок!
https://plands-diplomy.com
купить диплом в братске
купить диплом в междуреченске
купить диплом в ишиме
купить диплом в саранске
купить диплом в хасавюрте
One thing I’ve noticed is there are plenty of misguided beliefs regarding the banking companies intentions whenever talking about property foreclosures. One fairy tale in particular is the bank needs to have your house. The financial institution wants your hard earned cash, not the home. They want the bucks they lent you along with interest. Steering clear of the bank will draw a foreclosed conclusion. Thanks for your write-up.
Всем хорошего дня!
Моя работа над дипломом стала намного легче после того, как я обнаружил полезные онлайн-ресурсы.
Мы поможем вам купить диплом университета без предоплаты и доставим его в любой город России с гарантированной безопасностью.
Желаю вам всем прекрасных оценок!
https://plands-diplomy.com
купить диплом в саранске
купить диплом в красноярске
купить диплом в пятигорске
купить дипломы о высшем цены
купить диплом энергетика
Thanks for the tips you write about through your blog. In addition, a lot of young women who become pregnant do not even seek to get medical insurance because they fear they would not qualify. Although a few states now require that insurers produce coverage irrespective of the pre-existing conditions. Prices on these types of guaranteed programs are usually bigger, but when with the high cost of medical care it may be the safer strategy to use to protect your financial future.
Доброго дня!
Эффективность моей работы над дипломом возросла благодаря доступу к онлайн-материалам.
Заказывайте диплом у нас и получите его быстро и безопасно, оплата по факту доставки, отправка в любой регион России.
Желаю каждому отличных отметок!
купить диплом о высшем образовании
купить диплом специалиста
купить диплом матроса
купить диплом в нижневартовске
купить диплом в чапаевске
купить диплом в балаково
Found captivating reading that I’d like to offer you – you won’t regret it https://?????????.??/showthread.php?tid=34260
Доброго дня!
Инструменты из интернета значительно упростили мой процесс работы над дипломом.
Поможем выбрать, сделать заказ и купить диплом любого учебного заведения в любом населенном пункте по самым низким ценам.
Желаю для каждого прекрасных отметок!
https://frees-diplomy.com
купить диплом бурильщика
купить диплом в улан-удэ
купить диплом в альметьевске
купить диплом в первоуральске
купить диплом инженера
Thanks , I have recently been searching for info approximately this topic for ages and yours is the best I have discovered so far. However, what about the bottom line? Are you certain about the supply?
Доброго дня!
В интернете я нашел ресурсы для диплома, которые помогли мне ускорить написание.
Наш интернет-магазин предлагает купить российский диплом по выгодной цене, с гарантией проверенного качества.
Желаю всем не двоешных) оценок!
купить аттестат за 9 класс
купить диплом о среднем специальном
купить диплом в нальчике
купить диплом в ейске
купить диплом в бору
купить диплом в глазове
One important thing is that when you are searching for a student loan you may find that you will need a cosigner. There are many cases where this is correct because you may find that you do not have a past credit standing so the mortgage lender will require you have someone cosign the credit for you. Great post.
Excellent blog here! Also your site loads up very fast!
What host are you using? Can I get your affiliate link to your
host? I wish my site loaded up as quickly as yours lol
Приветики, дорогие мои!!
Использование онлайн-материалов сделало диплом более доступным для выполнения.
Мы поможем вам купить диплом университета без предоплаты и доставим его в любой город России с гарантированной безопасностью.
Желаю каждому честных отметок!
https://diplom-originalniy.com
купить диплом в тамбове
купить диплом в волжском
купить диплом ветеринара
где купить диплом образование
купить диплом слесаря
Hello there! Would you mind if I share your blog with my twitter group? There’s a lot of people that I think would really enjoy your content. Please let me know. Thanks
I like what you guys are up too. Such smart work and reporting! Carry on the excellent works guys I抳e incorporated you guys to my blogroll. I think it’ll improve the value of my website 🙂
Thanks for your thoughts. One thing I have noticed is that often banks in addition to financial institutions really know the spending routines of consumers and as well understand that the majority of people max out their cards around the breaks. They sensibly take advantage of this particular fact and commence flooding ones inbox and also snail-mail box along with hundreds of Zero APR card offers soon after the holiday season ends. Knowing that in case you are like 98 of American public, you’ll rush at the possiblity to consolidate financial debt and switch balances for 0 apr interest rates credit cards.
I have been examinating out many of your articles and i must say nice stuff. I will surely bookmark your blog.
I have realized some new elements from your web page about pc’s. Another thing I’ve always presumed is that computer systems have become a specific thing that each family must have for most reasons. They offer convenient ways to organize homes, pay bills, go shopping, study, listen to music and in some cases watch tv shows. An innovative strategy to complete these tasks is to use a mobile computer. These computer systems are mobile ones, small, powerful and convenient.
I was recommended this blog by my cousin. I’m not sure whether this post is written by him as no one else know such detailed about my problem. You are incredible! Thanks!
I have observed that of all kinds of insurance, medical health insurance is the most debatable because of the clash between the insurance cover company’s need to remain profitable and the user’s need to have insurance plan. Insurance companies’ income on wellness plans are certainly low, so some businesses struggle to make money. Thanks for the tips you discuss through this website.
Thanks for the auspicious writeup. It in truth was a amusement account it. Look advanced to more added agreeable from you! By the way, how could we communicate?
shop best travel gadgets on https://businesstravel.reviews/
I was wondering if you ever considered changing the layout of your blog? Its very well written; I love what youve got to say. But maybe you could a little more in the way of content so people could connect with it better. Youve got an awful lot of text for only having 1 or 2 pictures. Maybe you could space it out better?
I do not even understand how I ended up right here, but I believed this publish was once great.
I don’t recognise who you’re however definitely you are going to a well-known blogger if you happen to are not already.
Cheers!
I would like to thank you for the efforts you have put in penning this
site. I really hope to see the same high-grade blog posts by you later on as well.
In fact, your creative writing abilities has inspired me to get my own website now ;
)
Здравствуйте!
Диплом стал моей основной заботой из-за нарушения установленных сроков.
Поможем вам выбрать, оформить заказ и приобрести диплом любого учебного заведения по самым низким ценам.
Желаю каждому не двоешных) оценок!
купить диплом в москве
купить диплом в ишиме
купить диплом в бузулуке
купить диплом в чапаевске
купить диплом в грозном
купить диплом в шадринске
Здравствуйте!
Подготовка к диплому улучшилась с поиском ресурсов в интернете.
Приобретайте диплом университета без предоплаты у нас с доставкой в любой город России, качество и безопасность гарантированы.
Желаю каждому пятерочных) оценок!
https://radiploma.com/
купить диплом в артеме
купить диплом в славянске-на-кубани
купить диплом в тюмени
купить диплом ветеринара
купить диплом в ачинске
Thanks , I’ve recently been looking for information approximately this
subject for ages and yours is the best I have found out till now.
But, what in regards to the conclusion? Are you positive
about the source?
I like the helpful info you supply for your articles.
I will bookmark your blog and check once more here regularly.
I am fairly certain I’ll be informed many new stuff proper here!
Best of luck for the following!
Just desire to say your article is as surprising.
The clarity on your publish is just excellent and that i could think you’re an expert on this subject.
Well along with your permission allow me to snatch your RSS feed to
stay up to date with drawing close post. Thanks a million and please keep up the enjoyable work.
Its like you read my mind! You appear to know a lot about this, like you wrote the book in it
or something. I think that you can do with a few pics to drive the
message home a little bit, but other than that, this is wonderful blog.
A fantastic read. I will definitely be back.
It’s difficult to find knowledgeable people about this subject, but you seem like
you know what you’re talking about! Thanks
I enjoy looking through an article that can make people think.
Also, many thanks for allowing for me to comment!
청도페이스라인출장
Wow, superb blog layout!
How long have you been blogging for? you made blogging glance easy.
The overall look of your web site is magnificent, as neatly as the content!
You can read similar here prev
next and it’s
was wrote by Philip02.
Thanks for the points shared in your blog. One more thing I would like to mention is that fat loss is not exactly about going on a dietary fads and trying to lose as much weight as you can in a couple of days. The most effective way to shed pounds is by using it little by little and following some basic points which can enable you to make the most from the attempt to drop some weight. You may know and be following some tips, however reinforcing awareness never damages.
Thanks a lot for your post. I’d really like to write my opinion that the tariff of car insurance varies widely from one coverage to another, for the reason that there are so many different facets which contribute to the overall cost. Such as, the model and make of the automobile will have a huge bearing on the fee. A reliable ancient family auto will have an inexpensive premium over a flashy sports car.
Good response in return of this question with firm arguments
and explaining all on the topic of that.
Keep functioning ,terrific job!
This is very fascinating, You’re an overly skilled
blogger. I have joined your rss feed and sit up for in quest of extra of
your magnificent post. Also, I’ve shared your website in my
social networks
Hi! Do you know if they make any plugins to protect against hackers?
I’m kinda paranoid about losing everything I’ve worked
hard on. Any tips?
Nice post. I used to be checking constantly this weblog and I’m impressed!
Very helpful info specially the closing part :
) I maintain such information much. I was seeking this certain info for a long time.
Thank you and good luck.
Wow, marvelous weblog format!
How lengthy have you been running a blog for? you make running a blog look easy.
The total glance of your website is great, as well as the content material!
You can read similar here prev next and that was
wrote by Louann06.
I visited various websites except the audio feature for audio songs existing at this web site is actually wonderful.
Yes! Finally something about Bitcoin Price.
На сайте https://solvki-land.ru закажите увлекательную, познавательную экскурсию, рассчитанную на туристов различного возраста «Соловецкий феномен». Вы ознакомитесь с 10 экскурсиями, которые вы не сможете заказать ни в каком другом месте. Они длятся в течение 6 или 8 дней. На портале ознакомьтесь с природой Соловков, а в рамках экскурсий вы увидите Соловецкий сад, который представляет собой особую достопримечательность. На сайте вы увидите всю красоту острова, включая ландшафты, природу и многое другое.
Unquestionably believe that which you stated. Your favorite reason appeared to be on the web the easiest thing to be aware of.
I say to you, I certainly get annoyed while people think about worries that they plainly do not know about.
You managed to hit the nail upon the top and
also defined out the whole thing without having side effect , people could take a signal.
Will probably be back to get more. Thanks
Of course, what a fantastic site and informative posts, I will bookmark your website.Best Regards!
Thank you for another informative blog. The place else could I get that kind of info written in such a perfect manner? I’ve a challenge that I am just now working on, and I’ve been on the look out for such info.
Greate article. Keep writing such kind of info on your page.
Im really impressed by your site.
Hello there, You have done an incredible job.
I’ll definitely digg it and in my view recommend to my friends.
I am confident they’ll be benefited from this website.
If you are going for best contents like myself, only pay a visit this web site everyday because it gives quality contents, thanks
The subsequent time I learn a weblog, I hope that it doesnt disappoint me as much as this one. I mean, I do know it was my option to learn, but I really thought youd have one thing attention-grabbing to say. All I hear is a bunch of whining about one thing that you might fix for those who werent too busy searching for attention.
It’s not my first time to pay a quick visit this website, i am browsing this web site dailly and get nice facts from here all the time.
Wow, fantastic blog layout! How long have you been blogging for? you make blogging look easy. The overall look of your website is excellent, let alone the content!
#be#jk3#jk#jk#JK##
купить казахский номер
You are a very smart individual!
WOW just what I was searching for. Came here by searching for дом из клееного бруса под
ключ
I think the admin of this website is in fact working hard for
his web site, because here every data is quality based data.
Exceptional post but I was wondering if you could write a litte more on this subject? I’d be very thankful if you could elaborate a little bit further. Kudos!
One thing I’d like to say is the fact before purchasing more computer memory, look into the machine in to which it would be installed. If the machine will be running Windows XP, for instance, a memory limit is 3.25GB. Installing more than this would merely constitute some sort of waste. Make sure that one’s motherboard can handle an upgrade amount, as well. Good blog post.
Thanks in favor of sharing such a nice opinion, piece
of writing is fastidious, thats why i have read it fully
Undeniably consider that that you said. Your favourite reason appeared to be at the internet the simplest factor to remember of. I say to you, I certainly get irked at the same time as people think about worries that they plainly don’t know about. You controlled to hit the nail upon the highest as smartly as defined out the entire thing with no need side-effects , other folks could take a signal. Will probably be again to get more. Thank you
I was curious if you ever thought of changing the layout
of your website? Its very well written; I love what youve got to say.
But maybe you could a little more in the way of content so people could
connect with it better. Youve got an awful lot of
text for only having 1 or two pictures. Maybe you could space it out better?
Needed to compose you that bit of observation just to give many thanks over again over the marvelous concepts you have shown on this page. It is so particularly generous with you to supply extensively all most of us could possibly have advertised for an e-book to make some dough for their own end, most notably seeing that you might well have tried it if you ever decided. These thoughts likewise acted as the fantastic way to be certain that other individuals have the same desire similar to my very own to learn very much more pertaining to this matter. I am sure there are millions of more enjoyable opportunities in the future for folks who read carefully your blog post.
I like what you guys are up also. Such intelligent work and reporting! Keep up the superb works guys I抳e incorporated you guys to my blogroll. I think it’ll improve the value of my web site 🙂
It’s the best time to make some plans for the future and it’s time to be happy. I’ve read this post and if I could I desire to suggest you some interesting things or advice. Perhaps you could write next articles referring to this article. I desire to read even more things about it!
The other day, while I was at work, my cousin stole my iphone and tested to see if it can survive a twenty five foot drop, just so she can be a youtube sensation. My apple ipad is now destroyed and she has 83 views. I know this is totally off topic but I had to share it with someone!
I’ve come across that right now, more and more people are being attracted to digital cameras and the industry of pictures. However, as a photographer, it’s important to first shell out so much of your time deciding the exact model of digital camera to buy and moving from store to store just so you might buy the most inexpensive camera of the trademark you have decided to choose. But it would not end generally there. You also have to take into consideration whether you should obtain a digital digital camera extended warranty. Thx for the good tips I accumulated from your website.
I get pleasure from, cause I found just what I was looking for.
You’ve ended my four day long hunt! God Bless you man. Have a great day.
Bye
На сайте http://profotonsk.ru профессиональный фотограф предлагает свою услугу по специализации – интерьерная фотосессия. Большой опыт работы и множество рекламных фотосессий позволяет фотографу работать как с частными риэлторами, так и с агентствами недвижимости. В своей работе он применяет камеру с полноформатным сенсором и широкоугольный объектив, это обеспечивает отличное качество изображения, у вас есть возможность в этом лично убедиться. Подробнее посмотреть услуги фотографа, отзывы, а также его портфолио, вы можете на портале.
Good day! Do you use Twitter? I’d like to follow you if that would be okay. I’m undoubtedly enjoying your blog and look forward to new updates.
Hey are using WordPress for your site platform? I’m new to the blog world
but I’m trying to get started and set up my own. Do you require
any html coding expertise to make your own blog?
Any help would be greatly appreciated!
Awesome things here. I’m very happy to see your article.
Thank you so much and I’m looking forward to touch you. Will you please drop me a e-mail?
After looking into a number of the blog articles on your site, I
really appreciate your way of blogging. I
saved as a favorite it to my bookmark website list and will be
checking back soon. Please visit my web site too and tell me your opinion.
You could definitely see your skills in the paintings you write. The world hopes for more passionate writers like you who are not afraid to say how they believe. Always follow your heart.
Appreciating the commitment you put into your site and detailed information you provide.
It’s awesome to come across a blog every once in a while that isn’t the
same out of date rehashed material. Wonderful read! I’ve saved your
site and I’m adding your RSS feeds to my Google account.
It’s appropriate time to make some plans for the future and it is time to be happy. I’ve read this post and if I could I wish to suggest you few interesting things or tips. Perhaps you could write next articles referring to this article. I desire to read even more things about it!
I visited several blogs however the audio feature for audio
songs present at this web site is genuinely marvelous.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/manaev1993/profile
https://launchpad.net/~gfhfghsgg19991
https://www.quia.com/profiles/andreaconner
https://bodyara1997.bandcamp.com/album/my-wifes-discovery-part-3
https://launchpad.net/~defactori19961
В современном мире, где диплом – это начало успешной карьеры в любой отрасли, многие ищут максимально быстрый и простой путь получения образования. Важность наличия официального документа переоценить невозможно. Ведь именно диплом открывает дверь перед всеми, кто собирается вступить в профессиональное сообщество или продолжить обучение в ВУЗе.
Мы предлагаем оперативно получить этот необходимый документ. Вы можете приобрести диплом, что становится удачным решением для всех, кто не смог закончить обучение или утратил документ. диплом изготавливается с особой тщательностью, вниманием к мельчайшим деталям, чтобы на выходе получился продукт, максимально соответствующий оригиналу.
Преимущество данного подхода состоит не только в том, что можно оперативно получить диплом. Весь процесс организован комфортно, с профессиональной поддержкой. Начав от выбора нужного образца до правильного заполнения персональных данных и доставки по России — все будет находиться под полным контролем квалифицированных мастеров.
Для всех, кто хочет найти быстрый способ получения требуемого документа, наша компания предлагает выгодное решение. Заказать диплом – это значит избежать продолжительного процесса обучения и не теряя времени переходить к личным целям: к поступлению в ВУЗ или к началу трудовой карьеры.
https://originality-diplomans.com
Heya i’m for the primary time here. I came across this board and I
to find It truly helpful & it helped me out much.
I hope to present something back and aid others like you
aided me.
Сегодня, когда диплом – это начало успешной карьеры в любой области, многие стараются найти максимально простой путь получения качественного образования. Необходимость наличия официального документа об образовании сложно переоценить. Ведь диплом открывает дверь перед всеми, кто хочет вступить в сообщество профессиональных специалистов или продолжить обучение в ВУЗе.
Предлагаем максимально быстро получить этот важный документ. Вы сможете купить диплом нового или старого образца, что будет отличным решением для человека, который не смог закончить образование, потерял документ или желает исправить плохие оценки. дипломы выпускаются с особой аккуратностью, вниманием ко всем деталям. В итоге вы получите полностью оригинальный документ.
Преимущества подобного решения состоят не только в том, что можно максимально быстро получить диплом. Весь процесс организован комфортно и легко, с профессиональной поддержкой. От выбора нужного образца диплома до консультации по заполнению личных данных и доставки в любой регион страны — все под абсолютным контролем опытных специалистов.
Для всех, кто ищет быстрый способ получить необходимый документ, наша компания может предложить выгодное решение. Заказать диплом – это значит избежать длительного обучения и не теряя времени перейти к своим целям, будь то поступление в университет или начало успешной карьеры.
https://originality-diplomans.com
Many thanks! Quite a lot of forum posts!
Сегодня, когда диплом – это начало успешной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения образования. Важность наличия официального документа сложно переоценить. Ведь диплом открывает двери перед всеми, кто собирается начать трудовую деятельность или учиться в ВУЗе.
В данном контексте наша компания предлагает максимально быстро получить этот важный документ. Вы сможете купить диплом, и это является удачным решением для всех, кто не смог закончить обучение или утратил документ. Любой диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам, чтобы в итоге получился продукт, полностью соответствующий оригиналу.
Преимущество подобного подхода состоит не только в том, что можно оперативно получить свой диплом. Процесс организован удобно, с нашей поддержкой. От выбора требуемого образца документа до грамотного заполнения личной информации и доставки по России — все находится под абсолютным контролем качественных мастеров.
Всем, кто пытается найти оперативный способ получения требуемого документа, наша компания предлагает выгодное решение. Купить диплом – это значит избежать длительного процесса обучения и сразу переходить к важным целям: к поступлению в университет или к началу успешной карьеры.
диплом купить
В современном мире, где диплом – это начало успешной карьеры в любом направлении, многие ищут максимально простой путь получения качественного образования. Наличие документа об образовании трудно переоценить. Ведь именно диплом открывает дверь перед всеми, кто желает начать трудовую деятельность или продолжить обучение в ВУЗе.
Наша компания предлагает очень быстро получить любой необходимый документ. Вы имеете возможность заказать диплом, и это становится отличным решением для всех, кто не смог завершить образование, потерял документ или желает исправить свои оценки. Все дипломы изготавливаются аккуратно, с максимальным вниманием к мельчайшим деталям, чтобы в итоге получился полностью оригинальный документ.
Преимущества подобного решения состоят не только в том, что вы сможете быстро получить диплом. Весь процесс организовывается просто и легко, с нашей поддержкой. Начав от выбора подходящего образца до грамотного заполнения персональных данных и доставки по России — все под абсолютным контролем квалифицированных специалистов.
Всем, кто пытается найти максимально быстрый способ получения требуемого документа, наша услуга предлагает выгодное решение. Заказать диплом – это значит избежать долгого обучения и не теряя времени перейти к достижению собственных целей: к поступлению в ВУЗ или к началу удачной карьеры.
купить диплом института
В нашем обществе, где диплом является началом удачной карьеры в любой области, многие ищут максимально простой путь получения образования. Наличие официального документа переоценить невозможно. Ведь именно диплом открывает двери перед всеми, кто хочет начать трудовую деятельность или продолжить обучение в университете.
В данном контексте наша компания предлагает очень быстро получить любой необходимый документ. Вы можете приобрести диплом старого или нового образца, что становится выгодным решением для всех, кто не смог закончить обучение или утратил документ. Все дипломы выпускаются аккуратно, с особым вниманием к мельчайшим элементам. В итоге вы получите продукт, полностью соответствующий оригиналу.
Превосходство подобного решения состоит не только в том, что можно оперативно получить свой диплом. Процесс организовывается комфортно, с нашей поддержкой. Начав от выбора требуемого образца документа до точного заполнения личной информации и доставки в любой регион России — все под абсолютным контролем наших мастеров.
Для всех, кто ищет максимально быстрый способ получения требуемого документа, наша компания может предложить отличное решение. Купить диплом – это значит избежать долгого процесса обучения и не теряя времени переходить к достижению своих целей, будь то поступление в ВУЗ или начало профессиональной карьеры.
аттестат купить
В наше время, когда диплом – это начало удачной карьеры в любом направлении, многие стараются найти максимально быстрый и простой путь получения качественного образования. Важность наличия официального документа переоценить попросту невозможно. Ведь именно он открывает двери перед всеми, кто желает вступить в сообщество квалифицированных специалистов или учиться в высшем учебном заведении.
В данном контексте мы предлагаем оперативно получить этот важный документ. Вы можете купить диплом, что является отличным решением для всех, кто не смог завершить образование, потерял документ или желает исправить плохие оценки. дипломы выпускаются с особой тщательностью, вниманием к мельчайшим элементам. В результате вы получите продукт, 100% соответствующий оригиналу.
Преимущества данного решения заключаются не только в том, что можно максимально быстро получить свой диплом. Весь процесс организовывается просто и легко, с профессиональной поддержкой. От выбора необходимого образца до консультаций по заполнению персональной информации и доставки в любое место страны — все будет находиться под полным контролем квалифицированных специалистов.
В итоге, для тех, кто пытается найти оперативный способ получить необходимый документ, наша услуга предлагает выгодное решение. Приобрести диплом – значит избежать длительного обучения и сразу перейти к личным целям: к поступлению в университет или к началу успешной карьеры.
https://diplomana-asx.com
Всероссийская благотворительная школьная ярмарка дает возможность вам разместить свои товары либо приобрести за образовательную валюту услуги в приложении VK Умникоины и бенефиты без скачивания и регистрации. https://start.shkolnayayarmarka.ru/ – сайт, где можно разместить свои услуги и товары в качестве личного пожертвования. Также тут вы ознакомитесь детальнее с единицей обмена, которую используют для выполнения сделок на благотворительных ярмарках между продавцом и покупателем. Пишите нам уже сейчас.
В нашем мире, где диплом – это начало отличной карьеры в любом направлении, многие пытаются найти максимально простой путь получения качественного образования. Необходимость наличия официального документа переоценить попросту невозможно. Ведь диплом открывает двери перед всеми, кто стремится начать трудовую деятельность или продолжить обучение в высшем учебном заведении.
Мы предлагаем оперативно получить этот важный документ. Вы сможете купить диплом, и это является удачным решением для всех, кто не смог закончить обучение, потерял документ или хочет исправить плохие оценки. Любой диплом изготавливается аккуратно, с максимальным вниманием ко всем элементам, чтобы в итоге получился 100% оригинальный документ.
Преимущество подобного решения заключается не только в том, что можно быстро получить диплом. Весь процесс организовывается удобно, с нашей поддержкой. Начиная от выбора нужного образца диплома до правильного заполнения личной информации и доставки в любой регион страны — все под полным контролем наших мастеров.
Для тех, кто ищет быстрый и простой способ получения требуемого документа, наша компания предлагает выгодное решение. Купить диплом – значит избежать продолжительного обучения и не теряя времени переходить к достижению личных целей: к поступлению в университет или к началу трудовой карьеры.
купить аттестат
В современном мире, где диплом – это начало успешной карьеры в любом направлении, многие пытаются найти максимально простой путь получения качественного образования. Наличие официального документа об образовании сложно переоценить. Ведь диплом открывает дверь перед каждым человеком, желающим вступить в профессиональное сообщество или продолжить обучение в университете.
В данном контексте мы предлагаем очень быстро получить этот важный документ. Вы имеете возможность купить диплом, что является отличным решением для всех, кто не смог завершить образование или потерял документ. Все дипломы выпускаются с особой тщательностью, вниманием к мельчайшим нюансам, чтобы на выходе получился полностью оригинальный документ.
Превосходство такого решения заключается не только в том, что вы сможете оперативно получить свой диплом. Весь процесс организован удобно и легко, с профессиональной поддержкой. Начиная от выбора требуемого образца до правильного заполнения персональных данных и доставки в любой регион страны — все под полным контролем качественных специалистов.
Для всех, кто пытается найти максимально быстрый способ получения требуемого документа, наша компания предлагает выгодное решение. Заказать диплом – это значит избежать длительного обучения и не теряя времени перейти к важным целям: к поступлению в ВУЗ или к началу трудовой карьеры.
купить диплом медсестры
В современном мире, где диплом является началом отличной карьеры в любой области, многие пытаются найти максимально простой путь получения качественного образования. Наличие официального документа об образовании сложно переоценить. Ведь диплом открывает дверь перед каждым человеком, желающим вступить в сообщество профессионалов или учиться в ВУЗе.
В данном контексте наша компания предлагает очень быстро получить этот важный документ. Вы можете заказать диплом нового или старого образца, и это становится выгодным решением для всех, кто не смог закончить обучение, потерял документ или желает исправить свои оценки. диплом изготавливается с особой тщательностью, вниманием ко всем нюансам, чтобы в результате получился документ, полностью соответствующий оригиналу.
Превосходство данного решения заключается не только в том, что можно оперативно получить диплом. Весь процесс организовывается удобно и легко, с нашей поддержкой. Начиная от выбора нужного образца до точного заполнения личной информации и доставки по стране — все находится под полным контролем наших мастеров.
Таким образом, всем, кто ищет оперативный способ получить необходимый документ, наша компания предлагает выгодное решение. Купить диплом – значит избежать длительного обучения и сразу перейти к важным целям, будь то поступление в университет или начало карьеры.
https://adiplomy-russian.com
В наше время, когда диплом является началом удачной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения качественного образования. Важность наличия официального документа об образовании сложно переоценить. Ведь диплом открывает дверь перед любым человеком, желающим вступить в сообщество профессиональных специалистов или продолжить обучение в ВУЗе.
В данном контексте мы предлагаем максимально быстро получить этот важный документ. Вы можете купить диплом, и это становится выгодным решением для всех, кто не смог завершить обучение, потерял документ или желает исправить свои оценки. дипломы изготавливаются с особой аккуратностью, вниманием к мельчайшим деталям. На выходе вы получите полностью оригинальный документ.
Преимущества такого подхода заключаются не только в том, что можно оперативно получить диплом. Весь процесс организовывается удобно, с нашей поддержкой. От выбора подходящего образца до правильного заполнения личной информации и доставки в любой регион России — все под полным контролем квалифицированных специалистов.
Для тех, кто хочет найти быстрый способ получить необходимый документ, наша компания может предложить выгодное решение. Приобрести диплом – значит избежать продолжительного процесса обучения и не теряя времени переходить к достижению личных целей: к поступлению в университет или к началу трудовой карьеры.
https://adiplomy-russian.com
В современном мире, где диплом является началом удачной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения образования. Наличие официального документа переоценить просто невозможно. Ведь диплом открывает дверь перед каждым человеком, желающим вступить в сообщество профессионалов или продолжить обучение в высшем учебном заведении.
Предлагаем оперативно получить этот важный документ. Вы имеете возможность купить диплом, что становится удачным решением для всех, кто не смог завершить образование или потерял документ. дипломы производятся с особой аккуратностью, вниманием к мельчайшим элементам, чтобы в итоге получился полностью оригинальный документ.
Преимущество этого подхода заключается не только в том, что вы максимально быстро получите диплом. Процесс организован комфортно, с нашей поддержкой. От выбора подходящего образца до консультации по заполнению личной информации и доставки в любое место страны — все под абсолютным контролем наших мастеров.
В итоге, для всех, кто хочет найти быстрый способ получить необходимый документ, наша компания может предложить выгодное решение. Приобрести диплом – это значит избежать длительного обучения и не теряя времени перейти к своим целям: к поступлению в ВУЗ или к началу трудовой карьеры.
https://diploms-originalniy.com
В нашем обществе, где диплом – это начало успешной карьеры в любой сфере, многие стараются найти максимально простой путь получения качественного образования. Необходимость наличия официального документа об образовании переоценить просто невозможно. Ведь диплом открывает дверь перед всеми, кто желает начать трудовую деятельность или продолжить обучение в высшем учебном заведении.
В данном контексте мы предлагаем очень быстро получить этот необходимый документ. Вы можете заказать диплом нового или старого образца, что будет удачным решением для человека, который не смог закончить обучение, утратил документ или хочет исправить плохие оценки. Любой диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам, чтобы в результате получился полностью оригинальный документ.
Преимущества такого подхода состоят не только в том, что вы сможете максимально быстро получить свой диплом. Процесс организован просто и легко, с нашей поддержкой. От выбора подходящего образца до консультации по заполнению личной информации и доставки по России — все будет находиться под абсолютным контролем наших мастеров.
В результате, для тех, кто пытается найти быстрый и простой способ получить необходимый документ, наша компания предлагает отличное решение. Заказать диплом – это значит избежать долгого обучения и не теряя времени переходить к своим целям, будь то поступление в университет или старт карьеры.
https://diploms-originalniy.com
Came across a unique article – it’s worth your attention http://boltalka112.mybb.ru/post.php?fid=1
I have read so many articles concerning the blogger lovers however this article is really a pleasant article, keep
it up.
В нашем обществе, где диплом становится началом удачной карьеры в любом направлении, многие ищут максимально быстрый путь получения качественного образования. Важность наличия официального документа переоценить просто невозможно. Ведь диплом открывает двери перед людьми, желающими начать трудовую деятельность или учиться в высшем учебном заведении.
Мы предлагаем быстро получить этот важный документ. Вы имеете возможность купить диплом, и это является отличным решением для всех, кто не смог закончить обучение или утратил документ. дипломы изготавливаются с особой аккуратностью, вниманием к мельчайшим деталям. На выходе вы получите полностью оригинальный документ.
Плюсы этого подхода состоят не только в том, что вы максимально быстро получите свой диплом. Весь процесс организован комфортно, с нашей поддержкой. Начав от выбора необходимого образца до грамотного заполнения персональных данных и доставки по стране — все под абсолютным контролем наших мастеров.
Для всех, кто хочет найти оперативный способ получить необходимый документ, наша компания готова предложить отличное решение. Заказать диплом – это значит избежать долгого обучения и сразу перейти к своим целям, будь то поступление в университет или начало карьеры.
https://premialnie-diplomik.com
Hi! I know this is somewhat off topic but I was wondering if you knew where I could locate a captcha plugin for my comment form? I’m using the same blog platform as yours and I’m having problems finding one? Thanks a lot!
В нашем обществе, где диплом является началом успешной карьеры в любой области, многие пытаются найти максимально быстрый путь получения качественного образования. Наличие официального документа об образовании переоценить просто невозможно. Ведь именно он открывает дверь перед любым человеком, желающим начать профессиональную деятельность или учиться в ВУЗе.
Мы предлагаем быстро получить этот необходимый документ. Вы можете купить диплом, и это будет отличным решением для человека, который не смог закончить обучение, потерял документ или желает исправить плохие оценки. дипломы производятся аккуратно, с максимальным вниманием к мельчайшим деталям, чтобы на выходе получился документ, полностью соответствующий оригиналу.
Преимущество этого подхода состоит не только в том, что вы оперативно получите диплом. Весь процесс организован удобно, с нашей поддержкой. Начиная от выбора подходящего образца до консультации по заполнению персональных данных и доставки в любой регион страны — все под абсолютным контролем опытных специалистов.
Всем, кто хочет найти быстрый способ получения требуемого документа, наша компания предлагает выгодное решение. Приобрести диплом – значит избежать долгого обучения и не теряя времени перейти к своим целям: к поступлению в университет или к началу удачной карьеры.
купить диплом института
Greetings from California! I’m bored to death at work so I decided to browse your website on my iphone during lunch break. I enjoy the information you provide here and can’t wait to take a look when I get home. I’m shocked at how fast your blog loaded on my mobile .. I’m not even using WIFI, just 3G .. Anyways, excellent blog!
В нашем обществе, где диплом – это начало успешной карьеры в любой области, многие ищут максимально простой путь получения образования. Факт наличия официального документа об образовании переоценить невозможно. Ведь именно диплом открывает двери перед каждым человеком, который хочет вступить в профессиональное сообщество или учиться в университете.
Мы предлагаем очень быстро получить этот важный документ. Вы сможете приобрести диплом нового или старого образца, и это становится выгодным решением для человека, который не смог завершить образование или утратил документ. Каждый диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам. В результате вы получите 100% оригинальный документ.
Превосходство подобного решения заключается не только в том, что вы быстро получите диплом. Процесс организован комфортно, с нашей поддержкой. Начав от выбора подходящего образца до консультаций по заполнению личных данных и доставки в любое место страны — все под полным контролем качественных мастеров.
Всем, кто хочет найти быстрый способ получить требуемый документ, наша компания предлагает выгодное решение. Заказать диплом – значит избежать долгого обучения и сразу перейти к достижению своих целей, будь то поступление в ВУЗ или старт карьеры.
https://rudiplomista.com
I’m really inspired together with your writing abilities as well as with the structure on your blog. Is that this a paid subject matter or did you customize it your self? Anyway keep up the nice quality writing, it抯 rare to look a nice weblog like this one nowadays..
В современном мире, где диплом – это начало успешной карьеры в любой сфере, многие пытаются найти максимально простой путь получения качественного образования. Наличие официального документа об образовании переоценить попросту невозможно. Ведь диплом открывает двери перед людьми, желающими вступить в профессиональное сообщество или учиться в ВУЗе.
Наша компания предлагает очень быстро получить этот необходимый документ. Вы имеете возможность приобрести диплом нового или старого образца, что становится удачным решением для человека, который не смог завершить обучение, потерял документ или желает исправить свои оценки. дипломы изготавливаются с особой тщательностью, вниманием ко всем элементам, чтобы в итоге получился продукт, максимально соответствующий оригиналу.
Преимущество такого подхода состоит не только в том, что можно оперативно получить диплом. Процесс организовывается комфортно и легко, с профессиональной поддержкой. Начав от выбора требуемого образца до точного заполнения личных данных и доставки по России — все будет находиться под полным контролем качественных мастеров.
Всем, кто пытается найти быстрый способ получения требуемого документа, наша услуга предлагает выгодное решение. Приобрести диплом – это значит избежать долгого обучения и сразу переходить к личным целям, будь то поступление в ВУЗ или старт успешной карьеры.
купить аттестат школы
В нашем обществе, где диплом – это начало удачной карьеры в любом направлении, многие пытаются найти максимально быстрый путь получения качественного образования. Факт наличия официального документа переоценить невозможно. Ведь именно диплом открывает дверь перед людьми, желающими начать трудовую деятельность или учиться в любом институте.
Мы предлагаем максимально быстро получить любой необходимый документ. Вы можете заказать диплом, и это становится отличным решением для всех, кто не смог закончить обучение или утратил документ. диплом изготавливается с особой аккуратностью, вниманием ко всем нюансам, чтобы в итоге получился 100% оригинальный документ.
Превосходство такого подхода заключается не только в том, что можно максимально быстро получить диплом. Весь процесс организовывается удобно, с профессиональной поддержкой. От выбора необходимого образца документа до консультации по заполнению личных данных и доставки в любое место России — все под полным контролем опытных мастеров.
В итоге, для тех, кто пытается найти быстрый и простой способ получить необходимый документ, наша компания предлагает отличное решение. Купить диплом – это значит избежать длительного обучения и сразу перейти к своим целям: к поступлению в ВУЗ или к началу успешной карьеры.
https://clients1.google.com.ua/url?sa=t&url=https://lands-diploma.com
https://images.google.mk/url?q=https://rudiplomista.com
https://sandbox.google.com/url?q=https://lands-diploma.com
https://cse.google.rw/url?sa=t&url=https://premialnie-diplomik.com
https://images.google.se/url?q=https://premialnie-diplomik.com
Hello, the whole thing is going sound here and ofcourse every one is sharing facts, that’s genuinely fine, keep
up writing.
В нашем мире, где диплом – это начало успешной карьеры в любой области, многие пытаются найти максимально простой путь получения качественного образования. Наличие официального документа об образовании трудно переоценить. Ведь диплом открывает дверь перед любым человеком, который желает вступить в сообщество профессионалов или учиться в ВУЗе.
Предлагаем быстро получить этот необходимый документ. Вы сможете приобрести диплом старого или нового образца, и это является удачным решением для человека, который не смог закончить образование, утратил документ или желает исправить свои оценки. Все дипломы изготавливаются с особой аккуратностью, вниманием к мельчайшим элементам, чтобы в результате получился продукт, 100% соответствующий оригиналу.
Преимущества данного подхода состоят не только в том, что можно оперативно получить диплом. Процесс организован комфортно, с профессиональной поддержкой. От выбора нужного образца диплома до точного заполнения личных данных и доставки по России — все находится под полным контролем опытных специалистов.
Всем, кто ищет максимально быстрый способ получить требуемый документ, наша компания предлагает выгодное решение. Заказать диплом – это значит избежать долгого обучения и не теряя времени переходить к важным целям, будь то поступление в университет или начало успешной карьеры.
https://cse.google.com.tw/url?q=https://rudiplomista.com
https://maps.google.to/url?q=https://rudiplomista.com
https://clients1.google.co.im/url?sa=t&url=https://rudiplomista.com
https://clients1.google.com.pa/url?sa=t&url=https://rudiplomista.com
https://maps.google.cat/url?q=https://rudiplomista.com
Definitely believe that which you said. Your favorite reason seemed to be on the net the easiest thing to
be aware of. I say to you, I certainly get annoyed while people
consider worries that they plainly don’t know about. You managed to
hit the nail upon the top and also defined out the whole thing without having side effect , people can take a signal.
Will likely be back to get more. Thanks
That is a good tip especially to those fresh to the
blogosphere. Brief but very precise info… Many thanks for sharing this one.
A must read article!
В нашем мире, где диплом – это начало отличной карьеры в любом направлении, многие стараются найти максимально быстрый и простой путь получения качественного образования. Наличие документа об образовании трудно переоценить. Ведь именно диплом открывает двери перед любым человеком, который хочет начать профессиональную деятельность или учиться в любом ВУЗе.
Мы предлагаем максимально быстро получить любой необходимый документ. Вы можете заказать диплом нового или старого образца, что является удачным решением для человека, который не смог завершить образование или утратил документ. диплом изготавливается аккуратно, с максимальным вниманием к мельчайшим деталям. В итоге вы сможете получить документ, полностью соответствующий оригиналу.
Преимущества данного подхода заключаются не только в том, что вы максимально быстро получите свой диплом. Весь процесс организован удобно, с нашей поддержкой. Начав от выбора нужного образца диплома до грамотного заполнения персональных данных и доставки по России — все находится под полным контролем квалифицированных специалистов.
Таким образом, для тех, кто хочет найти быстрый и простой способ получить необходимый документ, наша компания может предложить отличное решение. Приобрести диплом – значит избежать продолжительного процесса обучения и сразу переходить к важным целям: к поступлению в ВУЗ или к началу трудовой карьеры.
https://images.google.sm/url?sa=t&url=https://premialnie-diplomik.com
http://www.google.mk/url?sa=i&url=https://lands-diploma.com
https://images.google.com.gt/url?q=https://rudiplomista.com
https://www.google.cz/url?q=https://rudiplomista.com
https://www.google.cz/url?q=https://lands-diploma.com
Someone essentially help to make critically posts I might state. That is the very first time I frequented your web page and thus far? I amazed with the research you made to create this particular post incredible. Wonderful job!
Hello, yeah this piece of writing is in fact fastidious and I have learned lot of things from
it on the topic of blogging. thanks.
I have to get across my passion for your kindness supporting men who really want help on in this situation. Your very own commitment to getting the solution all through had become certainly good and has specifically encouraged those like me to arrive at their pursuits. Your invaluable key points means so much a person like me and especially to my colleagues. With thanks; from each one of us.
What i do not realize is in fact how you’re now not actually a
lot more smartly-favored than you may be now.
You’re very intelligent. You already know thus significantly with regards
to this topic, produced me individually imagine it from numerous numerous angles.
Its like women and men are not interested except it’s something to accomplish with Girl gaga!
Your personal stuffs great. Always handle it
up!
В нашем мире, где диплом является началом успешной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения качественного образования. Важность наличия официального документа переоценить попросту невозможно. Ведь именно он открывает двери перед всеми, кто стремится вступить в профессиональное сообщество или продолжить обучение в ВУЗе.
В данном контексте наша компания предлагает оперативно получить любой необходимый документ. Вы имеете возможность заказать диплом, что становится удачным решением для человека, который не смог завершить образование, утратил документ или желает исправить плохие оценки. Любой диплом изготавливается с особой аккуратностью, вниманием к мельчайшим нюансам, чтобы в итоге получился продукт, максимально соответствующий оригиналу.
Плюсы этого подхода состоят не только в том, что можно оперативно получить диплом. Весь процесс организован просто и легко, с нашей поддержкой. Начиная от выбора требуемого образца документа до консультации по заполнению персональных данных и доставки по России — все под абсолютным контролем наших мастеров.
Всем, кто хочет найти оперативный способ получить необходимый документ, наша компания предлагает отличное решение. Купить диплом – значит избежать продолжительного обучения и сразу перейти к своим целям, будь то поступление в ВУЗ или старт профессиональной карьеры.
https://clients1.google.com.pr/url?sa=t&url=https://lands-diploma.com
http://cse.google.es/url?sa=i&url=https://rudiplomista.com
https://maps.google.hu/url?sa=t&url=https://lands-diploma.com
https://image.google.am/url?q=https://rudiplomista.com
https://cse.google.com.ly/url?sa=t&url=https://rudiplomista.com
На сайте https://luxbabycar.ru представлен каталог детских авто. Ассортимент постоянно обновляется. Здесь вы найдете: джипы, квадроциклы, багги, спецтехнику, толокар и многое другое высокого качества и по конкурентоспособным ценам. У нас самый широкий выбор детского автопарка. На сайте вы можете ознакомиться с фото-отзывами, а также посмотреть видео-обзор. Проконсультируйтесь со специалистом прямо сейчас. Весь товар прошел проверку качества, вы можете быть уверены за свой новый автомобиль.
This site really has all the information I wanted concerning this subject and didn’t know
who to ask.
Aw, this was a very good post. Taking a few minutes and
actual effort to make a very good article… but what can I say…
I procrastinate a whole lot and don’t manage
to get nearly anything done.
Excellent, what a weblog it is! This web site gives
valuable information to us, keep it up.
В нашем мире, где диплом – это начало удачной карьеры в любом направлении, многие стараются найти максимально простой путь получения качественного образования. Факт наличия официального документа об образовании трудно переоценить. Ведь именно диплом открывает двери перед любым человеком, желающим вступить в сообщество профессиональных специалистов или учиться в высшем учебном заведении.
Наша компания предлагает максимально быстро получить этот важный документ. Вы имеете возможность приобрести диплом, и это становится выгодным решением для всех, кто не смог закончить обучение, утратил документ или желает исправить плохие оценки. дипломы выпускаются аккуратно, с максимальным вниманием ко всем элементам. В итоге вы сможете получить документ, полностью соответствующий оригиналу.
Преимущество этого подхода заключается не только в том, что вы максимально быстро получите диплом. Процесс организовывается комфортно, с нашей поддержкой. От выбора подходящего образца до грамотного заполнения личных данных и доставки в любой регион России — все будет находиться под полным контролем качественных мастеров.
Всем, кто ищет быстрый и простой способ получить необходимый документ, наша услуга предлагает выгодное решение. Заказать диплом – это значит избежать продолжительного обучения и сразу переходить к достижению собственных целей, будь то поступление в университет или начало карьеры.
https://www.google.pl/url?q=https://rudiplomista.com
https://images.google.com.cy/url?q=https://rudiplomista.com
https://images.google.co.kr/url?q=https://premialnie-diplomik.com
http://maps.google.ro/url?q=https://premialnie-diplomik.com
http://images.google.com.na/url?q=https://premialnie-diplomik.com
Thanks for your advice on this blog. Just one thing I would like to say is the fact that purchasing consumer electronics items through the Internet is not new. Actually, in the past several years alone, the marketplace for online electronic devices has grown noticeably. Today, you will find practically almost any electronic tool and tools on the Internet, ranging from cameras along with camcorders to computer elements and gambling consoles.
excellent points altogether, you simply received a emblem new
reader. What could you recommend in regards to your
submit that you made a few days ago? Any positive?
На сегодняшний день, когда диплом является началом удачной карьеры в любом направлении, многие пытаются найти максимально быстрый путь получения образования. Наличие официального документа переоценить попросту невозможно. Ведь диплом открывает дверь перед каждым человеком, который собирается начать трудовую деятельность или продолжить обучение в каком-либо университете.
В данном контексте мы предлагаем оперативно получить этот необходимый документ. Вы можете купить диплом, и это является отличным решением для всех, кто не смог закончить обучение или потерял документ. Любой диплом изготавливается аккуратно, с особым вниманием ко всем деталям, чтобы на выходе получился продукт, максимально соответствующий оригиналу.
Преимущества этого решения заключаются не только в том, что можно оперативно получить свой диплом. Процесс организован удобно, с профессиональной поддержкой. От выбора подходящего образца диплома до консультаций по заполнению личных данных и доставки по стране — все будет находиться под абсолютным контролем квалифицированных мастеров.
Таким образом, для всех, кто хочет найти быстрый способ получения требуемого документа, наша компания предлагает выгодное решение. Приобрести диплом – это значит избежать продолжительного процесса обучения и сразу переходить к достижению личных целей: к поступлению в университет или к началу удачной карьеры.
http://u-cars.ru/modules.php?name=Your_Account&op=userinfo&username=oqavyfisu
http://travel.klimashevich.com/2012/02/blog-post.html
http://zoobla.com/blog/security-cameras-increase-overall-safety
http://topforbaby.ru/product/adameks-koljaska-transf-galaxy-ljulka-imit/reviews/
http://atoll-filters.ru/product/umjagchitel-atoll-ecolife-s-20/reviews/
This is my first time visit at here and i am in fact impressed to read all at alone place.
В современном мире, где диплом – это начало отличной карьеры в любом направлении, многие ищут максимально простой путь получения качественного образования. Факт наличия документа об образовании сложно переоценить. Ведь диплом открывает двери перед каждым человеком, желающим начать трудовую деятельность или учиться в ВУЗе.
В данном контексте мы предлагаем максимально быстро получить этот важный документ. Вы сможете заказать диплом нового или старого образца, и это будет удачным решением для всех, кто не смог завершить образование, утратил документ или желает исправить плохие оценки. Каждый диплом изготавливается с особой аккуратностью, вниманием ко всем деталям, чтобы на выходе получился полностью оригинальный документ.
Превосходство подобного решения заключается не только в том, что вы сможете быстро получить свой диплом. Весь процесс организован удобно, с профессиональной поддержкой. Начиная от выбора необходимого образца диплома до консультаций по заполнению персональной информации и доставки по России — все под полным контролем опытных специалистов.
Всем, кто пытается найти быстрый способ получения необходимого документа, наша компания предлагает отличное решение. Заказать диплом – это значит избежать продолжительного обучения и сразу переходить к личным целям, будь то поступление в ВУЗ или старт карьеры.
http://images.google.pn/url?q=https://rudiplomista.com
https://images.google.co.jp/url?sa=t&url=https://premialnie-diplomik.com
https://clients1.google.com.bo/url?q=https://lands-diploma.com
https://www.google.com/url?sa=t&url=https://premialnie-diplomik.com
http://images.google.pt/url?q=https://lands-diploma.com
Hey there! I’m at work surfing around your blog from my new iphone!
Just wanted to say I love reading your blog and look forward to all your posts!
Carry on the fantastic work!
Aw, this was a really good post. Finding the time and actual effort to make a
superb article… but what can I say… I hesitate a lot and don’t seem
to get anything done.
В современном мире, где диплом является началом успешной карьеры в любой сфере, многие стараются найти максимально быстрый и простой путь получения образования. Наличие официального документа трудно переоценить. Ведь диплом открывает двери перед любым человеком, который стремится вступить в сообщество квалифицированных специалистов или учиться в любом ВУЗе.
В данном контексте наша компания предлагает быстро получить этот важный документ. Вы имеете возможность купить диплом нового или старого образца, что становится выгодным решением для человека, который не смог закончить образование, потерял документ или хочет исправить свои оценки. дипломы изготавливаются с особой аккуратностью, вниманием к мельчайшим элементам. На выходе вы сможете получить документ, полностью соответствующий оригиналу.
Превосходство этого решения заключается не только в том, что вы сможете максимально быстро получить свой диплом. Весь процесс организовывается удобно и легко, с нашей поддержкой. Начиная от выбора необходимого образца диплома до консультаций по заполнению личных данных и доставки в любое место страны — все будет находиться под абсолютным контролем квалифицированных мастеров.
Всем, кто хочет найти быстрый способ получения необходимого документа, наша компания может предложить отличное решение. Заказать диплом – значит избежать длительного обучения и сразу переходить к достижению личных целей, будь то поступление в университет или начало карьеры.
https://toolbarqueries.google.co.jp/url?sa=t&rct=j&q=https://premialnie-diplomik.com
https://images.google.com.sv/url?q=https://rudiplomista.com
https://www.google.dz/url?q=https://rudiplomista.com
https://maps.google.li/url?q=https://premialnie-diplomik.com
https://toolbarqueries.google.co.ls/url?q=https://lands-diploma.com
В нашем обществе, где диплом является началом успешной карьеры в любой области, многие ищут максимально быстрый путь получения качественного образования. Факт наличия официального документа об образовании трудно переоценить. Ведь диплом открывает двери перед всеми, кто стремится вступить в сообщество профессионалов или учиться в каком-либо ВУЗе.
Мы предлагаем максимально быстро получить любой необходимый документ. Вы сможете приобрести диплом, что будет выгодным решением для всех, кто не смог завершить образование, потерял документ или хочет исправить свои оценки. Все дипломы выпускаются аккуратно, с особым вниманием к мельчайшим деталям, чтобы на выходе получился полностью оригинальный документ.
Превосходство такого решения состоит не только в том, что вы сможете оперативно получить диплом. Процесс организован просто и легко, с профессиональной поддержкой. Начав от выбора необходимого образца до правильного заполнения персональной информации и доставки в любое место России — все под полным контролем опытных мастеров.
Для всех, кто ищет быстрый и простой способ получить необходимый документ, наша компания предлагает отличное решение. Приобрести диплом – значит избежать продолжительного обучения и не теряя времени переходить к достижению своих целей, будь то поступление в ВУЗ или начало карьеры.
http://korea-shipping.co.kr/bbs/board.php?bo_table=free&wr_id=572949
http://brenderupmediterrane.free.fr/index.php?file=Members&op=detail&autor=elohuqoxa
http://feki-php.8u.cz/profile.php?lookup=10885
http://www.kiopro.ru/support/forum/view_profile.php?UID=71122
http://forum.hayabusa-club.ru/member.php?u=17305
Oh my goodness! Awesome article dude! Thank you, However I am encountering issues with your RSS.
I don’t know why I am unable to subscribe to it.
Is there anyone else having identical RSS issues? Anyone that knows the answer
will you kindly respond? Thanx!!
Der Artikel war sehr informativ und gut strukturiert.
Die Beispiele haben das Verständnis vertieft. Danke für Ihre hervorragende Arbeit!
В современном мире, где диплом становится началом удачной карьеры в любой отрасли, многие пытаются найти максимально быстрый и простой путь получения образования. Важность наличия официального документа об образовании трудно переоценить. Ведь диплом открывает дверь перед всеми, кто собирается вступить в сообщество профессионалов или продолжить обучение в ВУЗе.
Предлагаем оперативно получить этот необходимый документ. Вы сможете приобрести диплом старого или нового образца, что будет удачным решением для человека, который не смог завершить обучение, утратил документ или хочет исправить свои оценки. Каждый диплом изготавливается с особой аккуратностью, вниманием к мельчайшим нюансам, чтобы на выходе получился продукт, максимально соответствующий оригиналу.
Преимущество такого решения состоит не только в том, что вы сможете оперативно получить свой диплом. Процесс организован просто и легко, с нашей поддержкой. От выбора необходимого образца документа до консультации по заполнению личных данных и доставки по стране — все под полным контролем опытных мастеров.
Для всех, кто ищет максимально быстрый способ получения требуемого документа, наша услуга предлагает выгодное решение. Приобрести диплом – это значит избежать продолжительного обучения и не теряя времени перейти к личным целям: к поступлению в университет или к началу удачной карьеры.
https://www.google.hn/url?sa=t&url=https://premialnie-diplomik.com
https://clients1.google.com.ua/url?sa=t&url=https://lands-diploma.com
https://cse.google.cat/url?q=https://lands-diploma.com
https://toolbarqueries.google.cf/url?q=https://rudiplomista.com
https://maps.google.bg/url?sa=i&source=web&rct=j&url=https://lands-diploma.com
We are a group of volunteers and starting a new scheme in our community.
Your site offered us with valuable information to work on. You’ve done
an impressive job and our entire community will be grateful to you.
Outstanding quest there. What occurred after?
Thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/dwroma1954
https://launchpad.net/~gi1g1ma1n19731
https://waylon1972.diary.ru/
https://www.metal-archives.com/users/morya4ok1980
https://rentry.org/4zp3b5wi
Эколан – компания, которая предлагает большой ассортимент качественных товаров. У нас вы найдете прекрасные одеяла, простыни, полотенца, скатерти, декоративные подушки, наматрасники, пледы и многое другое. Мы гарантируем привлекательные цены и быструю доставку. https://ecolan37.ru – сайт, где найдете только самое интересное для домашнего уюта и комфорта. Ценим своих клиентов, всегда готовы помочь с выбором и ответить на любые вопросы. Заказывая у нас, можете быть уверены в том, что останетесь приобретением, довольны.
This website was… how do you say it? Relevant!! Finally I’ve found something that helped me.
Kudos!
Stumbled upon a captivating article – definitely take a look! https://lrnews.mirtesen.ru/blog/43634409777/Sovremennyie-plastikovyie-okna-pod-klyuch-kupit-v-Moskve
Wow, superb blog format!
How long have you been blogging for? you made blogging glance easy.
The overall look of your website is magnificent, let alone the content!
I read similar here prev next and those was wrote by Gary88.
If some one wants expert view on the topic of blogging and site-building
after that i recommend him/her to pay a quick visit this weblog, Keep
up the good job.
В нашем обществе, где диплом является началом успешной карьеры в любом направлении, многие ищут максимально быстрый путь получения образования. Важность наличия официального документа трудно переоценить. Ведь диплом открывает дверь перед людьми, стремящимися начать трудовую деятельность или продолжить обучение в любом ВУЗе.
В данном контексте наша компания предлагает быстро получить этот необходимый документ. Вы можете заказать диплом нового или старого образца, и это является отличным решением для всех, кто не смог завершить образование, потерял документ или хочет исправить свои оценки. дипломы производятся с особой тщательностью, вниманием ко всем элементам. На выходе вы получите 100% оригинальный документ.
Плюсы этого подхода заключаются не только в том, что вы быстро получите диплом. Весь процесс организовывается комфортно, с нашей поддержкой. Начав от выбора нужного образца документа до консультаций по заполнению персональной информации и доставки по стране — все под полным контролем квалифицированных мастеров.
Всем, кто ищет максимально быстрый способ получить требуемый документ, наша компания может предложить выгодное решение. Заказать диплом – значит избежать продолжительного процесса обучения и сразу переходить к важным целям, будь то поступление в университет или начало карьеры.
https://toolbarqueries.google.co.ls/url?q=https://premialnie-diplomik.com
https://clients1.google.com.nf/url?sa=t&url=https://lands-diploma.com
https://cse.google.mw/url?sa=t&url=https://rudiplomista.com
https://maps.google.sk/url?q=https://rudiplomista.com
https://www.google.co.cr/url?q=https://rudiplomista.com
We are a group of volunteers and starting a new
scheme in our community. Your website provided us with valuable
info to work on. You’ve done an impressive job and our whole community will be thankful to you.
I believe that is one of the most significant info for me.
And i am glad studying your article. However should
remark on some basic things, The website style is perfect, the articles
is truly nice : D. Good process, cheers
Сегодня, когда диплом – это начало успешной карьеры в любой сфере, многие ищут максимально простой путь получения образования. Наличие официального документа сложно переоценить. Ведь именно он открывает дверь перед любым человеком, который хочет начать профессиональную деятельность или продолжить обучение в университете.
Предлагаем быстро получить этот важный документ. Вы сможете приобрести диплом, и это является выгодным решением для всех, кто не смог закончить обучение, потерял документ или хочет исправить плохие оценки. Все дипломы производятся аккуратно, с особым вниманием к мельчайшим нюансам, чтобы в итоге получился полностью оригинальный документ.
Плюсы подобного решения заключаются не только в том, что можно оперативно получить свой диплом. Процесс организован просто и легко, с нашей поддержкой. Начав от выбора подходящего образца до грамотного заполнения персональной информации и доставки по стране — все под абсолютным контролем опытных специалистов.
Всем, кто хочет найти быстрый и простой способ получить требуемый документ, наша компания предлагает выгодное решение. Купить диплом – значит избежать долгого обучения и сразу перейти к достижению своих целей, будь то поступление в ВУЗ или начало карьеры.
https://cse.google.com.uy/url?sa=t&url=https://rudiplomista.com
http://www.google.com.tr/url?q=https://lands-diploma.com
https://clients1.google.tn/url?sa=t&url=https://rudiplomista.com
https://www.google.it/url?q=https://premialnie-diplomik.com
https://image.google.tg/url?q=https://lands-diploma.com
Excellent website you have here but I was curious if you knew of any discussion boards that cover the same topics discussed here? I’d really love to be a part of group where I can get feed-back from other knowledgeable people that share the same interest. If you have any recommendations, please let me know. Thanks a lot!
В наше время, когда диплом является началом успешной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения качественного образования. Наличие официального документа об образовании сложно переоценить. Ведь именно диплом открывает дверь перед всеми, кто собирается начать профессиональную деятельность или продолжить обучение в высшем учебном заведении.
Предлагаем быстро получить этот важный документ. Вы имеете возможность приобрести диплом нового или старого образца, и это становится удачным решением для всех, кто не смог закончить обучение, потерял документ или хочет исправить свои оценки. Каждый диплом изготавливается с особой аккуратностью, вниманием ко всем элементам. На выходе вы сможете получить полностью оригинальный документ.
Плюсы данного подхода заключаются не только в том, что вы быстро получите свой диплом. Процесс организован удобно и легко, с нашей поддержкой. Начав от выбора необходимого образца до точного заполнения персональной информации и доставки по стране — все под абсолютным контролем квалифицированных мастеров.
Всем, кто ищет оперативный способ получить необходимый документ, наша компания предлагает отличное решение. Приобрести диплом – значит избежать продолжительного процесса обучения и сразу переходить к достижению личных целей: к поступлению в ВУЗ или к началу удачной карьеры.
https://gidro2000.com/forum-gidro/user/25528/
http://team.yzy.free.fr/index.php?file=Members&op=detail&autor=uzukifyj
http://diabetesarmy.com/blog-post-title/?unapproved=49879&moderation-hash=198557d0ebd24ca309e214d7c7cd1984#comment-49879
https://gidro2000.com/forum-gidro/user/25558/
http://tsmtech.co.kr/bbs/board.php?bo_table=free&wr_id=486783
대전나이트클럽
hey there and thank you for your information – I’ve certainly picked up something new
from right here. I did however expertise some technical issues using this web site, as I
experienced to reload the site a lot of times previous to I could get it to load properly.
I had been wondering if your web hosting is OK? Not that I am complaining,
but sluggish loading instances times will sometimes affect your
placement in google and could damage your quality score if ads and marketing with Adwords.
Anyway I’m adding this RSS to my email and could look out for a lot more of
your respective intriguing content. Make sure you update
this again very soon.
My brother recommended I might like this website. He was once entirely right.
This post truly made my day. You cann’t imagine
simply how so much time I had spent for this information! Thanks!
lacolinaecuador.com
이럴 경우 황제는 대체로 덜 꾸짖는 것 같다.
На сегодняшний день, когда диплом является началом отличной карьеры в любом направлении, многие ищут максимально простой путь получения качественного образования. Наличие официального документа об образовании переоценить попросту невозможно. Ведь именно диплом открывает двери перед людьми, желающими начать профессиональную деятельность или продолжить обучение в высшем учебном заведении.
Предлагаем быстро получить этот важный документ. Вы сможете купить диплом нового или старого образца, что является отличным решением для всех, кто не смог завершить обучение, утратил документ или хочет исправить плохие оценки. диплом изготавливается с особой аккуратностью, вниманием ко всем нюансам. В результате вы сможете получить 100% оригинальный документ.
Превосходство данного подхода заключается не только в том, что можно максимально быстро получить свой диплом. Процесс организовывается просто и легко, с профессиональной поддержкой. Начав от выбора необходимого образца до правильного заполнения персональной информации и доставки по стране — все под полным контролем опытных мастеров.
Для всех, кто хочет найти быстрый и простой способ получить необходимый документ, наша компания готова предложить выгодное решение. Приобрести диплом – это значит избежать долгого обучения и не теряя времени переходить к достижению собственных целей, будь то поступление в университет или начало карьеры.
https://maps.google.sk/url?q=https://premialnie-diplomik.com
https://www.google.pt/url?q=https://lands-diploma.com
https://www.google.sc/url?q=https://premialnie-diplomik.com
http://www.google.mw/url?sa=i&url=%2Fhttps://rudiplomista.com
https://clients1.google.rw/url?q=https://lands-diploma.com
Good post. I learn something totally new and challenging
on blogs I stumbleupon everyday. It’s always useful to read through articles from other authors and
use a little something from other web sites.
На сайте http://waltzprof.com вы можете на выгодных условиях приобрести стальной профиль в широком ассортименте. Они предназначены для лофт перегородок, фасадных конструкций, удобной и быстрой сборки ограждающих преград и другое. Компания использует передовые технологии для выпуска качественной продукции. Покупка от производителя – это прекрасный способ сэкономить на услугах посредника. Подробнее ознакомиться с продукцией и посмотреть информацию о компании, вы можете на сайте. Закажите обратный звонок, и менеджеры вас проконсультируют по всем вопросам.
My partner and I stumbled over here by a different website and thought I
might as well check things out. I like what I see so
now i am following you. Look forward to checking out your web page yet again.
Сегодня, когда диплом становится началом удачной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения качественного образования. Необходимость наличия документа об образовании сложно переоценить. Ведь именно он открывает двери перед любым человеком, желающим вступить в профессиональное сообщество или учиться в ВУЗе.
В данном контексте мы предлагаем максимально быстро получить любой необходимый документ. Вы можете заказать диплом старого или нового образца, что является удачным решением для всех, кто не смог завершить образование или утратил документ. дипломы производятся аккуратно, с особым вниманием ко всем нюансам. В результате вы получите продукт, максимально соответствующий оригиналу.
Преимущество подобного подхода заключается не только в том, что вы сможете быстро получить диплом. Весь процесс организован просто и легко, с профессиональной поддержкой. От выбора нужного образца диплома до точного заполнения личной информации и доставки в любой регион страны — все находится под полным контролем опытных мастеров.
Для тех, кто ищет максимально быстрый способ получения необходимого документа, наша компания предлагает выгодное решение. Приобрести диплом – значит избежать продолжительного обучения и не теряя времени переходить к личным целям: к поступлению в университет или к началу удачной карьеры.
http://vipblinds.ru/index.php?subaction=userinfo&user=aqovyqys
https://talented-people.vraiforum.com/profile.php?mode=viewprofile&u=8462
http://mkala-koncert.ru/users/ybapeq
http://dobrknigi.ru/howto/?PAGE_NAME=profile_view&UID=24321
http://camillacastro.us/forums/viewtopic.php?pid=284827#p284827
An attention-grabbing discussion is price comment. I believe that you should write extra on this subject, it may not be a taboo subject however typically people are not enough to talk on such topics. To the next. Cheers
I loved as much as you will receive carried out right here. The sketch is tasteful, your authored material stylish. nonetheless, you command get got an impatience over that you wish be delivering the following. unwell unquestionably come further formerly again as exactly the same nearly very often inside case you shield this hike.
В современном мире, где диплом – это начало удачной карьеры в любой отрасли, многие стараются найти максимально быстрый путь получения качественного образования. Необходимость наличия официального документа об образовании переоценить попросту невозможно. Ведь диплом открывает дверь перед каждым человеком, желающим вступить в сообщество профессионалов или учиться в любом институте.
Наша компания предлагает максимально быстро получить любой необходимый документ. Вы можете заказать диплом, и это будет удачным решением для человека, который не смог завершить обучение, утратил документ или желает исправить плохие оценки. Каждый диплом изготавливается аккуратно, с особым вниманием ко всем деталям. На выходе вы сможете получить продукт, полностью соответствующий оригиналу.
Превосходство подобного подхода состоит не только в том, что вы сможете быстро получить свой диплом. Весь процесс организовывается удобно, с профессиональной поддержкой. От выбора нужного образца документа до точного заполнения персональной информации и доставки в любое место России — все под полным контролем опытных специалистов.
Таким образом, всем, кто хочет найти быстрый способ получить требуемый документ, наша компания готова предложить выгодное решение. Приобрести диплом – значит избежать продолжительного обучения и сразу переходить к своим целям, будь то поступление в университет или старт карьеры.
https://images.google.cf/url?q=https://rudiplomista.com
https://images.google.cf/url?q=https://premialnie-diplomik.com
https://www.google.com.cu/url?q=https://premialnie-diplomik.com
http://cse.google.es/url?sa=i&url=https://rudiplomista.com
https://images.google.com.ly/url?q=https://rudiplomista.com
Hello, for all time i used to check web site posts
here early in the break of day, because i enjoy to learn more and
more.
Excellent beat ! I wish to apprentice while you amend
your site, how could i subscribe for a blog web
site? The account aided me a acceptable deal.
I had been a little bit acquainted of this your broadcast provided bright clear concept
A motivating discussion is definitely worth comment. I believe that you ought
to write more about this issue, it may not be a taboo matter but typically folks don’t speak about these topics.
To the next! All the best!!
В нашем обществе, где диплом является началом успешной карьеры в любой сфере, многие стараются найти максимально быстрый и простой путь получения качественного образования. Наличие официального документа об образовании переоценить попросту невозможно. Ведь диплом открывает дверь перед каждым человеком, желающим начать трудовую деятельность или учиться в высшем учебном заведении.
Наша компания предлагает оперативно получить этот важный документ. Вы можете приобрести диплом, что становится выгодным решением для человека, который не смог завершить образование, утратил документ или хочет исправить свои оценки. диплом изготавливается с особой тщательностью, вниманием к мельчайшим нюансам. В итоге вы сможете получить полностью оригинальный документ.
Преимущество этого подхода состоит не только в том, что вы сможете оперативно получить свой диплом. Весь процесс организовывается комфортно, с профессиональной поддержкой. Начиная от выбора необходимого образца до консультаций по заполнению персональных данных и доставки в любой регион страны — все под абсолютным контролем качественных специалистов.
Всем, кто ищет оперативный способ получить требуемый документ, наша компания предлагает выгодное решение. Приобрести диплом – значит избежать продолжительного процесса обучения и не теряя времени перейти к достижению своих целей, будь то поступление в университет или начало карьеры.
http://images.google.ps/url?q=https://premialnie-diplomik.com
https://www.google.com.eg/url?q=https://premialnie-diplomik.com
http://maps.google.com/url?q=https://lands-diploma.com
https://cse.google.tt/url?sa=t&url=https://premialnie-diplomik.com
http://images.google.gl/url?q=https://premialnie-diplomik.com
Good article! We will be linking to this particularly great
article on our website. Keep up the good writing.
Fascinating blog! Is your theme custom made or
did you download it from somewhere? A theme
like yours with a few simple tweeks would really make my blog jump out.
Please let me know where you got your design. Thank you
There is definately a lot to know about this subject.
I like all of the points you’ve made.
It’s actually a nice and useful piece of information. I’m happy that you shared
this helpful information with us. Please keep us up to
date like this. Thank you for sharing.
Piece of writing writing is also a excitement, if you be acquainted with
then you can write or else it is complicated to write.
I have observed that online degree is getting favorite because obtaining your college degree online has become a popular method for many people. A lot of people have not necessarily had an opportunity to attend a traditional college or university but seek the raised earning possibilities and a better job that a Bachelor Degree affords. Still some others might have a qualification in one course but want to pursue a thing they now possess an interest in.
each time i used to read smaller posts that as well clear their motive, and that is also happening with this article which
I am reading now.
I have been surfing on-line more than 3 hours lately, but I never discovered any attention-grabbing article like yours. It is beautiful value enough for me. In my opinion, if all site owners and bloggers made just right content material as you probably did, the net will likely be much more helpful than ever before.
Write more, thats all I have to say. Literally, it seems as
though you relied on the video to make your point.
You definitely know what youre talking about, why
throw away your intelligence on just posting videos
to your blog when you could be giving us something informative to
read?
Keep working ,great job!
I don’t even know how I ended up here, but I thought this post was good.
I don’t know who you are but definitely you’re
going to a famous blogger if you aren’t already 😉 Cheers!
Букмекерская контора 1win – одна из самых популярных площадок, где пользователи могут делать ставки, играть, делать ставки и т. д. Для привлечения новой аудитории данная букмекерская контора предлагает новичкам отличный бонус – возможность получить до 200 000 бонусов за 4 депозита. И для этого покупателям даже не нужно вводить промокоды. Вам просто нужно зарегистрироваться в этом сервисе.
Промокод 1вин 2024: m1WIN2024 — это уникальный код, который необходимо указать при регистрации для получения бонуса 500% до 75 000 рублей. Это предложение доступно только новым игрокам, которые могут претендовать на приветственный бонус 1Win.
Для постоянных клиентов букмекерская контора постоянно выпускает новые промокоды 1win, ведь с этими бонусами клиентам гораздо приятнее пользоваться услугами этой букмекерской конторы. Промокод – это уникальный набор букв и цифр, активация которого позволяет человеку получить бонус. В этом обзоре мы расскажем, где взять новые промокоды 1win и как их активировать для получения бонусов.
Актуальный промокод 1Win 2024 вы можете найти на различных страницах с информацией о бонусах в букмекерских конторах. Продажи также осуществляются через партнеров компании. Лучшее место для поиска купонов – Telegram-канал букмекерской конторы. Новые ваучеры появляются там каждый день. 1Win может отправить промокод индивидуально уже зарегистрированному клиенту. Например, по случаю годовщины регистрации или просто дня рождения клиента.
С промокодом 1WIN новые игроки могут значительно увеличить сумму своего первого и последующих депозитов. Полученные бонусы можно использовать в игре и в случае успеха перевести на свой электронный кошелек. Максимальная сумма бонуса – 75 000 рублей.
Отдельной вкладки для проверки комбинаций нет. Если введено правильно, система активирует бонусное предложение. Во вкладке «Ваучер» в личном кабинете появится сообщение при вводе промокода 1Vin. Отсюда вы сможете увидеть, правильно ли была введена комбинация.
Источник: https://mmocenter.ru/blog/promokod-1win-promokody-1vin-pri-registracii-na-segodnya/
Link exchange is nothing else however it is simply placing the other person’s weblog
link on your page at appropriate place and other person will
also do similar in support of you.
ShaneViart
Доброго дня!
Интернет предоставил мне инструменты для диплома, благодаря которым процесс написания стал более простым и быстрым.
Мы рады предложить большой выбор документов об образовании всех ВУЗов России по выгодным условиям.
Желаю каждому отличных отметок!
купить диплом в азове
купить диплом товароведа
купить бланк диплома
купить диплом в каменске-шахтинском
купить диплом моториста
купить диплом в донском
купить диплом в междуреченске
купить диплом для иностранцев
купить диплом в артеме
купить диплом тренера
One more thing. It’s my opinion that there are lots of travel insurance web sites of dependable companies that permit you to enter your trip details and have you the prices. You can also purchase an international travel insurance policy on the internet by using the credit card. Everything you should do should be to enter the travel information and you can see the plans side-by-side. Only find the plan that suits your finances and needs then use your bank credit card to buy the item. Travel insurance on the web is a good way to start looking for a reliable company for international travel insurance. Thanks for giving your ideas.
When I originally left a comment I seem to have clicked on the
-Notify me when new comments are added- checkbox and from
now on each time a comment is added I get four emails with the
same comment. Perhaps there is an easy method you are able to remove me from that service?
Appreciate it!
Howdy! Someone in my Facebook group shared this website with us so I came to check it out. I’m definitely enjoying the information. I’m book-marking and will be tweeting this to my followers! Outstanding blog and wonderful style and design.
Howdy are using WordPress for your site platform? I’m new to the blog world but I’m trying
to get started and create my own. Do you require any coding knowledge to make your own blog?
Any help would be really appreciated!
Wow, incredible blog layout! How long have you been blogging for?
you make blogging look easy. The overall look of your website is excellent, as well as the content!
비아그라: 주요 성분은 “시테라필”이며, 인산화효소 5형을 억제하는 약물입니다.
My coder is trying to persuade me to move to .net from PHP.
I have always disliked the idea because of the expenses.
But he’s tryiong none the less. I’ve been using WordPress on a variety of websites for about
a year and am nervous about switching to another platform.
I have heard very good things about blogengine.net. Is there a way
I can import all my wordpress posts into it?
Any help would be greatly appreciated!
You ought to take part in a contest for one of the
best blogs on the internet. I will highly recommend this blog!
LhaneViart
Добрый день!
Диплом требует постоянной работы, подкрепляемой найденными онлайн-ресурсами.
Мы поможем вам купить диплом университета без предоплаты и доставим его в любой город России с гарантированной безопасностью.
Желаю вам всем не двоешных) оценок!
купить диплом в сарове
купить диплом в ейске
купить диплом в сыктывкаре
купить диплом в бузулуке
купить диплом дизайнера
купить диплом в ейске
купить диплом с занесением в реестр
купить диплом о высшем образовании
купить диплом в кропоткине
купить диплом в ярославле
Heya i am for the primary time here. I came across this board and I to find
It truly helpful & it helped me out a lot. I hope to provide something back and help others such as you helped me.
I’m truly enjoying the design and layout of your
blog. It’s a very easy on the eyes which makes it much more pleasant
for me to come here and visit more often. Did you hire out a developer to create your theme?
Exceptional work!
Why users still use to read news papers when in this technological globe
all is existing on net?
Thanks for the auspicious writeup. It in truth was a amusement
account it. Glance advanced to far introduced agreeable
from you! However, how can we keep in touch?
I do not even understand how I stopped up here, but I believed this post was great. I don’t know who you might be however definitely you’re going to a famous blogger in case you are not already 😉 Cheers!
Thank you for the sensible critique. Me and my neighbor were just preparing to do some research about this. We got a grab a book from our area library but I think I learned more clear from this post. I am very glad to see such wonderful info being shared freely out there.
Howdy! I’m at work surfing around your blog from my new
iphone 4! Just wanted to say I love reading through your blog
and look forward to all your posts! Carry on the fantastic work!
Hmm it seems like your blog ate my first comment (it was extremely long) so I guess I’ll just sum it up what I had written and say, I’m thoroughly enjoying your blog. I too am an aspiring blog writer but I’m still new to everything. Do you have any suggestions for newbie blog writers? I’d really appreciate it.
Hey! Do you use Twitter? I’d like to follow you if that would be ok. I’m definitely enjoying your blog and look forward to new updates.
Everyone loves what you guys are up too. This sort of clever work and reporting! Keep up the very good works guys I’ve included you guys to our blogroll.
Thanks for your personal marvelous posting! I genuinely enjoyed
reading it, you could be a great author. I will be sure to
bookmark your blog and may come back later on. I want to encourage you to continue your great job, have a nice evening!
https://himchanchan.tistory.com/1749
https://pornmaster.fun/hd/富联娱乐☘EFB88F9797·me💓皇马娱乐天美娱乐☘EFB88F9797·me💓欧皇娱乐
Good website! I truly love how it is easy on my eyes and the data are well written. I am wondering how I might be notified whenever a new post has been made. I’ve subscribed to your feed which must do the trick! Have a great day!
Hi! I could have sworn I’ve been to this site before but after checking through some of the post I realized it’s new to me. Nonetheless, I’m definitely delighted I found it and I’ll be book-marking and checking back often!
I want to point out my passion for your generosity in support of people who have the need for guidance on this one topic. Your special commitment to getting the message all around appeared to be amazingly significant and has continuously allowed individuals like me to reach their goals. Your own valuable help and advice implies this much to me and additionally to my peers. Regards; from all of us.
Хотите купить на dvd увлекательные сериалы? Serialexpress готов вам в этом помочь. Тут можете найти диски, которых нет в иных магазинах. Качество записи сериалов отменное, мы уверены в том, что вы приобретением останетесь довольны. Товары доступны по привлекательной цене, они хорошо упакованы и целые. Ищете купить сериал в иркутске? Serialexpress.ru – сайт, где имеется каталог. Здесь вы также можете ознакомиться с условиями оплаты. У нас действует накопительные скидки. Качество доставки вас порадует. Задавайте свои вопросы по электронной почте нашим менеджерам.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/khn9ynvp
https://onedio.ru/profile/vbnjho-196-0
https://chyoa.com/user/ecko19931988
https://www.obesityhelp.com/members/borm1966/about_me/
https://cannabis.net/user/149349
Just wish to say your article is as surprising. The clarity in your
post is just excellent and i can assume you are an expert on this subject.
Fine with your permission allow me to grab your RSS feed to keep
updated with forthcoming post. Thanks a million and please continue the gratifying work.
Aw, this was an incredibly nice post. Taking a few minutes and actual effort to generate a very good article but what can I say
I procrastinate a lot and don’t seem to get nearly anything done.
You’re so awesome! I do not believe I’ve truly read through a single thing like that before.
So wonderful to find another person with some genuine thoughts on this topic.
Really.. many thanks for starting this up. This site is something that is
needed on the web, someone with some originality!
Someone necessarily lend a hand to make significantly
posts I’d state. That is the first time I frequented your website page and up
to now? I amazed with the analysis you made to create this actual publish extraordinary.
Great job!
В нашем обществе, где диплом – это начало отличной карьеры в любой сфере, многие ищут максимально простой путь получения образования. Наличие документа об образовании переоценить невозможно. Ведь диплом открывает дверь перед любым человеком, желающим начать профессиональную деятельность или учиться в высшем учебном заведении.
Мы предлагаем быстро получить этот важный документ. Вы можете заказать диплом, и это становится выгодным решением для всех, кто не смог закончить образование, потерял документ или хочет исправить свои оценки. Каждый диплом изготавливается аккуратно, с особым вниманием ко всем элементам. На выходе вы получите документ, полностью соответствующий оригиналу.
Преимущества этого решения состоят не только в том, что можно оперативно получить свой диплом. Весь процесс организовывается удобно, с нашей поддержкой. Начиная от выбора необходимого образца диплома до консультации по заполнению личной информации и доставки в любой регион страны — все под абсолютным контролем квалифицированных специалистов.
Всем, кто ищет оперативный способ получить требуемый документ, наша компания может предложить выгодное решение. Купить диплом – значит избежать долгого процесса обучения и не теряя времени перейти к важным целям, будь то поступление в университет или начало профессиональной карьеры.
http://www.poster-street.com/blog/2010/04/how-to-make-a-wall-decoration/comment-page-1/?rcommentid=1063264&rerror=incorrect-captcha-sol&rchash=698421d5a649d90c70e3fff9507b61d3#commentform
http://mikimiki.jp/dmc-gx7mk2/?unapproved=96453&moderation-hash=8ee894a27c4e7160b1b95019b6f7a686#comment-96453
http://tkdheadquarters.com/bbs/board.php?bo_table=free&wr_id=184481
http://www.senbernar.ru/profile/10890-iwejizel/
http://hedron-arch.com/component/k2/item/24-digital-learning/
OLaneViart
Всем хорошего дня!
Продолжаю работать над дипломом, преодолевая трудности благодаря онлайн-поддержке.
У нас вы можете приобрести диплом университета без предоплаты с доставкой в любой город России, оплачивая после получения.
Желаю каждому не двоешных) оценок!
купить диплом в иркутске
купить диплом маляра
купить диплом в новочебоксарске
купить диплом в чайковском
купить диплом в братске
купить диплом в кунгуре
купить диплом кандидата наук
купить диплом в златоусте
купить диплом в набережных челнах
купить диплом в сызрани
Сайт v8corp.ru предлагает виды услуг по автоматизации и цифровизации процессов с использованием программ 1С. Сотрудничая с нами, вы сосредоточитесь на собственном бизнесе, мы в 24 отраслях экономики разбираемся. https://v8corp.ru/ – сайт, где есть возможность детальнее посмотреть конфигурации 1С. У нас гибкие тарифы. Поможем вам подобрать наилучшую конфигурацию продуктов 1С, подходящих конкретно для вашей компании. Оцените открывающиеся преимущества облачной Аренды 1С. Готовы быть вашим надежным помощником. Получите консультацию уже сейчас, обращайтесь!
It’s really very complex in this busy life to listen news on Television, therefore I only
use world wide web for that purpose, and get the most
up-to-date information.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/rankq1958
https://tubeteencam.com/user/kopcap191999/profile
https://rentry.org/ovthuxcf
https://imageevent.com/shyachlo1987
https://darions21990.diary.ru/
Paragraph writing is also a excitement, if you be familiar with afterward you can write otherwise
it is complex to write.
I think this is one of the most significant information for me. And i’m glad reading your article. But should remark on few general things, The web site style is perfect, the articles is really nice : D. Good job, cheers
Hi there, just became alert to your blog through Google, and found that it is really informative. I抦 gonna watch out for brussels. I抣l be grateful if you continue this in future. Lots of people will be benefited from your writing. Cheers!
Hi, Neat post. There’s a problem together with your site in web explorer, would check this?IE still is the marketplace leader and a huge section of people will leave out your excellent writing due to this problem.
It’s genuinely very complex in this full of activity life
to listen news on TV, therefore I just use internet for
that reason, and obtain the most up-to-date information.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/347166
https://www.quia.com/profiles/marcus496ma
https://lushnica1999.diary.ru/
https://okwave.jp/profile/u3110805.html
https://onedio.ru/profile/mimy-196-4
Wow! After all I got a webpage from where I be
able to really take helpful information regarding my study and knowledge.
I do like the way you have presented this specific concern plus it does give me a lot of fodder for consideration. Nevertheless, through what precisely I have witnessed, I basically wish when the actual responses pile on that people today stay on point and not get started on a soap box of some other news du jour. Anyway, thank you for this outstanding piece and even though I do not really go along with this in totality, I regard your perspective.
I have noticed that sensible real estate agents almost everywhere are starting to warm up to FSBO Promoting. They are seeing that it’s more than merely placing a sign post in the front property. It’s really pertaining to building relationships with these retailers who someday will become buyers. So, while you give your time and efforts to helping these suppliers go it alone : the “Law regarding Reciprocity” kicks in. Thanks for your blog post.
Whats up are using WordPress for your site platform? I’m new
to the blog world but I’m trying to get started and create my own. Do you
require any coding expertise to make your own blog?
Any help would be really appreciated!
Shop Megalopolis Toys
Первый шаг к увеличению бонуса при регистрации или совершении ставок. Текущий промокод 1xbet на сегодня — 1x_109745. На вкладке «Регистрация» у вас есть выбор: бонус в ставках на спорт и бесплатная ставка в казино.
Чтобы получить Промокод 1xbet, вы должны стать активным игроком. Для этого вам необходимо зарегистрироваться и пополнить свой счет. Бонус на депозит предоставляется бесплатно всем новым игрокам согласно акции.
Для регистрации необходимо найти актуальное на сегодня зеркало и ввести сегодняшний промокод 1x_109745. Вы можете зарегистрироваться в один клик – по электронной почте, номеру телефона или в социальных сетях. Сети. Далее заполните форму в личном кабинете. Обратите особое внимание на обязательные поля под звездочкой. Если вы заполните его правильно, вы получите сообщение «Данные успешно сохранены». Бонус становится доступен после первого пополнения игрового счета одним из способов из блока пополнения.
Бонусы 1xbet можно получить в рублях, долларах и евро, в зависимости от того, из какой вы страны. Каждый пользователь, который зарегистрируется на официальном сайте и воспользуется промокодом, получит бонусы от букмекерской конторы 1xbet.
Размер бонуса по промокоду конторы 1xBet будет равен 100% от суммы первого депозита от 100 до 6500 рублей. Вы можете использовать промокод дня 1xbet только один раз; Вы получите бонусные деньги сразу после пополнения баланса. Этот бонус необходимо отыграть в течение месяца. Оборот должен превышать сумму, зачисленную на бонусный счет, в 5 раз. Делайте экспресс-ставки на 3 исхода с коэффициентом выше 1,4. На 1xbet вы можете делать ставки на спортивные события, использовать прогнозы капперов, чтобы получить максимальные условия, используйте наш промокод при регистрации 1xbet — 1x_109745.
Промокод 1хбет
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/kakawey-195-1
https://libertydragon1956.diary.ru/
https://tubeteencam.com/user/warriwor1957/profile
https://launchpad.net/~truehefar19601
https://chyoa.com/user/shienn1980
Hello, I think your website might be having browser compatibility
issues. When I look at your blog in Firefox,
it looks fine but when opening in Internet Explorer, it has some overlapping.
I just wanted to give you a quick heads up! Other then that,
fantastic blog!
Wonderful post however , I was wanting to know if you could write a litte more on this subject?
I’d be very thankful if you could elaborate a little bit more.
Appreciate it!
Every weekend i used to visit this website, as i wish for enjoyment, as
this this site conations in fact nice funny data too.
Hey very nice blog!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/tgkumo6w
https://www.haikudeck.com/presentations/Gvc4sAMJUN
https://www.obesityhelp.com/members/fggfgfg1998/about_me/
https://imageevent.com/arlinka1972
https://tubeteencam.com/user/tolik21984/profile
울산콜걸
It’s an remarkable article in favor of all
the internet users; they will take advantage from it I am sure.
It’s very straightforward to find out any matter on web as compared to books, as I found this paragraph at
this web site.
Found an enthralling article, I recommend you to read https://geopolityka.org/20240423/chomu-kiyani-obirayut-taksi-dlya-shhodennih-poizdok/
Timsothyreemn
Доброго дня!
Дипломное исследование стало для меня серьезным вызовом, однако я нашел способы облегчить этот процесс.
Мы предлагаем вам купить диплом университета без предоплаты, с гарантированной доставкой по всей России.
Желаю всем прекрасных оценок!
купить диплом тренера
купить диплом в мытищах
где купить диплом образование
купить диплом в нижневартовске
купить диплом мастера маникюра и педикюра
купить свидетельство о разводе
купить диплом в минеральных водах
купить диплом в ханты-мансийске
купить диплом о среднем специальном
купить диплом в бийске
Привет, дорогой читатель!
Купите диплом ВУЗа по выгодной цене с гарантией качества и доставкой в любой город России без предоплаты!
https://sandbox.google.com/url?q=https://gosznac-diploma.com
http://www.google.rs/url?q=https://premialnie-diplomansy.com
https://asia.google.com/url?q=http://aurus-diploms.com
https://images.google.hu/url?q=https://diploms-asx.com
http://images.google.com.bh/url?sa=t&url=http://free-diplommi.com
Закажите диплом института или техникума по доступной цене с гарантированной доставкой в любой уголок России без предоплаты!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/148508
https://launchpad.net/~beast33919681
https://haveagood.holiday/users/348412
http://www.babelcube.com/user/markus-azuse
https://onedio.ru/profile/mmmnmnmn-197-0
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/shannon-blacksmith
http://www.babelcube.com/user/danny-schneider
https://www.hentai-foundry.com/user/utimi1982/profile
https://cannabis.net/user/149812
https://chyoa.com/user/sinder1987
Aw, this was a really good post. Taking a few minutes and actual effort to make
a top notch article… but what can I say… I procrastinate
a whole lot and don’t manage to get anything done.
Hello, its fastidious paragraph on the topic of media print, we
all be aware of media is a wonderful source of information.
Thank you for every other excellent article.
Where else could anybody get that kind of information in such an ideal method of
writing? I have a presentation next week, and I am on the look for such information.
At this time I am going away to do my breakfast,
when having my breakfast coming yet again to read additional news.
Hey there! This is my first comment here so I just wanted to give a quick shout out
and say I really enjoy reading your blog posts. Can you recommend any
other blogs/websites/forums that go over the same subjects?
Thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/347714
http://www.babelcube.com/user/mike-mayberry
https://www.obesityhelp.com/members/kitsyne171987/about_me/
https://www.quia.com/profiles/sh173simms
https://okwave.jp/profile/u3110878.html
These are in fact fantastic ideas in on the topic of blogging.
You have touched some pleasant points here. Any way
keep up wrinting.
Hi! Do you know if they make any plugins to safeguard against hackers?
I’m kinda paranoid about losing everything I’ve
worked hard on. Any recommendations?
강남콜걸
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/benji-french
https://rentry.org/fg95r36v
https://imageevent.com/leemura1967
https://cannabis.net/user/149893
https://www.haikudeck.com/presentations/7iHILj2wmv
imb88 situs ternama dan terkemuka membawa anda ke tempat dengan pengalaman berbeda sserta berikan yang terbaik dan terpercaya
I am now not sure the place you are getting your info, however good topic. I must spend some time finding out much more or figuring out more. Thank you for great information I used to be looking for this information for my mission.
Wow, fantastic blog layout! How long have you been blogging for? you made blogging look easy. The overall look of your website is wonderful, as well as the content!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/57247-andy-skindell
https://www.divephotoguide.com/user/sikais1121957
https://ellak.gr/user/steath1988/
https://bora19821953.bandcamp.com/album/mothers-day-fantasy
https://www.obesityhelp.com/members/islam991986/about_me/
Why visitors still make use of to read news papers when in this technological world all is accessible on net?
Hello there! I could have sworn I’ve been to this blog
before but after reading through some of the post I realized it’s new to me.
Anyways, I’m definitely happy I found it and I’ll
be bookmarking and checking back often!
Timsothyreemn
Приветики, дорогие мои!!
С каждым днем работа над дипломом становится легче, благодаря найденным онлайн-материалам.
У нас можно заказать диплом без предоплаты с доставкой курьером по РФ, “под ключ”!
Желаю каждому прекрасных оценок!
купить диплом в муроме
купить диплом в славянске-на-кубани
купить диплом университета
купить диплом в северске
купить диплом оценщика
купить диплом в великом новгороде
купить диплом сварщика
купить диплом в улан-удэ
купить диплом в новотроицке
купить диплом педагога
Hello there! I know this is kind of off topic but I was wondering if you knew where
I could get a captcha plugin for my comment
form? I’m using the same blog platform as yours and I’m having trouble finding one?
Thanks a lot!
Hey There. I found your blog the use of msn. That is a very neatly
written article. I’ll be sure to bookmark it and come back to
learn more of your useful information. Thank you for the post.
I’ll certainly return.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/stavia1970/profile
https://www.hentai-foundry.com/user/jetux1954/profile
https://www.obesityhelp.com/members/forestrun1973/about_me/
https://okwave.jp/profile/u3111934.html
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190971
Right here is the perfect blog for anyone who wants to understand this topic.
You know a whole lot its almost tough to argue with you (not that I
actually would want to…HaHa). You definitely put a fresh spin on a topic which
has been discussed for decades. Wonderful stuff, just wonderful!
Magnificent beat ! I wish to apprentice even as you amend your website,
how can i subscribe for a blog site? The account aided me a applicable deal.
I were a little bit familiar of this your broadcast offered vivid transparent idea
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://ellak.gr/user/indium1989/
https://rentry.org/r3wsvc8x
https://okwave.jp/profile/u3110851.html
https://permacultureglobal.org/users/55961-man-anjani
https://tubeteencam.com/user/sappysue1960/profile
You ought to take part in a contest for one of the best blogs online.
I most certainly will highly recommend this site!
Write more, thats all I have to say. Literally, it seems as though
you relied on the video to make your point. You definitely know what youre talking about,
why waste your intelligence on just posting videos to
your weblog when you could be giving us something informative
to read?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/rankq1958
https://ghtry1977.diary.ru/
https://www.haikudeck.com/presentations/BrOWK6se7N
https://onedio.ru/profile/gijifa-195-2
https://savade1998.diary.ru/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/56848-luis-ahmed
https://imageevent.com/bacr1988
https://haveagood.holiday/users/348240
https://www.divephotoguide.com/user/badlion1987
https://cannabis.net/user/149812
etsyweddingteam.com
素晴らしい洞察と分析。これからも素晴らしい記事を期待しています。
Greetings I am so glad I found your blog, I really found you by mistake, while I was browsing
on Bing for something else, Nonetheless I am here
now and would just like to say thanks for a fantastic post and a all round interesting
blog (I also love the theme/design), I don’t have time to read through it all
at the moment but I have saved it and also included
your RSS feeds, so when I have time I will be back to read more, Please do keep up the awesome job.
Its such as you read my thoughts! You seem to understand so much about this, like you wrote the guide in it or something. I feel that you could do with some p.c. to pressure the message house a little bit, but other than that, that is excellent blog. An excellent read. I will certainly be back.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/gjbt8cmx
https://www.divephotoguide.com/user/fornax1955
https://launchpad.net/~mansmack19851
http://www.babelcube.com/user/marcus-pope
https://haveagood.holiday/users/347169
Visit website
situs slot gacor baru imb88 yang terbaik sepanjang masa dengan RTP tinggi hari ini dan proses terbaik dan tercepat judi bola euro
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/rge1989
https://twitch1966.bandcamp.com/album/a-little-sluts-punishment
https://rentry.org/y929gztm
http://www.babelcube.com/user/kristin-mitchell
https://imageevent.com/ui1989
Thank you for any other informative site. The place else could I get
that kind of info written in such an ideal way?
I’ve a undertaking that I’m simply now operating on, and I have been at the look
out for such information.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://ellak.gr/user/nieves1975/
https://tubeteencam.com/user/nekta1970/profile
https://www.quia.com/profiles/dannybo245
https://verda1967.diary.ru/
https://tubeteencam.com/user/manaev1993/profile
Nice post. I learn something new and challenging on sites I stumbleupon everyday.
It will always be exciting to read articles from other
authors and use a little something from their websites.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/149995
https://imageevent.com/scaut301961
http://www.babelcube.com/user/rachael-king
https://imageevent.com/iioii1960
https://www.hentai-foundry.com/user/gordon771995/profile
Thank you for sharing your info. I truly appreciate your efforts and I will be waiting for your next write ups thanks once again.
Thanks for any other great post. Where else may anybody get that kind of information in such an ideal approach of writing?
I’ve a presentation next week, and I’m at the
look for such information.
Hello are using WordPress for your site platform?
I’m new to the blog world but I’m trying to get started and create my own. Do
you need any coding knowledge to make your own blog?
Any help would be really appreciated!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~wwor19651
https://permacultureglobal.org/users/55960-christopher-buten
https://www.hentai-foundry.com/user/bearka1965/profile
https://onedio.ru/profile/lipopa-196-9
http://www.babelcube.com/user/robert-owens
ShaneViart
Приветики, дорогие мои!!
Диплом уже не вызывает столько стресса, как раньше, благодаря помощи, полученной через интернет.
Приобретите диплом университета без предоплаты у нас и получите его с доставкой в любую точку России, качество гарантировано.
Желаю всем отличных отметок!
купить диплом в каменске-уральском
купить диплом в чайковском
купить диплом в сосновом бору
купить диплом в рыбинске
купить диплом техникума
купить диплом швеи
купить диплом сантехника
купить диплом в ухте
купить диплом в ульяновске
купить диплом в калуге
В современном мире, где диплом становится началом отличной карьеры в любой сфере, многие ищут максимально простой путь получения образования. Факт наличия официального документа об образовании сложно переоценить. Ведь именно он открывает дверь перед людьми, стремящимися начать трудовую деятельность или учиться в ВУЗе.
В данном контексте мы предлагаем очень быстро получить этот важный документ. Вы сможете приобрести диплом старого или нового образца, что является выгодным решением для человека, который не смог завершить образование или утратил документ. Все дипломы изготавливаются с особой тщательностью, вниманием ко всем деталям, чтобы на выходе получился продукт, полностью соответствующий оригиналу.
Плюсы подобного решения состоят не только в том, что можно оперативно получить свой диплом. Процесс организовывается комфортно, с профессиональной поддержкой. Начав от выбора подходящего образца диплома до правильного заполнения личной информации и доставки по России — все под полным контролем квалифицированных специалистов.
Всем, кто пытается найти быстрый способ получить необходимый документ, наша компания готова предложить отличное решение. Приобрести диплом – это значит избежать длительного процесса обучения и не теряя времени переходить к достижению личных целей, будь то поступление в ВУЗ или начало карьеры.
http://fridayad.in/user/profile/2370973
http://koreasamsong.com/bbs/board.php?bo_table=free&wr_id=2121385
http://193.16.101.6/index.php?title=eonlinediplom
http://blog.netskills.ru/2017/01/network-security15.html
http://v208870d.bget.ru/index.php?subaction=userinfo&user=ihezyse
Hello There. I found your blog using msn. This is a very well written article.
I’ll make sure to bookmark it and return to read more of your useful info.
Thanks for the post. I will definitely return.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/tiraj1983/profile
http://www.babelcube.com/user/rachael-davis
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190693
https://smokeplumes1969.diary.ru/
https://trawnik1957.diary.ru/
Do you mind if I quote a few of your posts as long as I provide credit
and sources back to your blog? My blog site is in the very same niche as yours and my visitors would truly benefit from a lot of
the information you present here. Please let me know if this
okay with you. Regards!
If you want to get a good deal from this piece of writing then you have to apply
these strategies to your won blog.
Excellent post. I was checking constantly this blog and I am inspired!
Extremely helpful information specially the last section :
) I care for such info a lot. I was looking for this particular information for a very lengthy time.
Thank you and best of luck.
Thanks for giving your ideas right here. The other matter is that every time a problem arises with a pc motherboard, people should not consider the risk regarding repairing this themselves because if it is not done properly it can lead to permanent damage to an entire laptop. It will always be safe to approach any dealer of a laptop with the repair of its motherboard. They have technicians with an knowledge in dealing with notebook motherboard problems and can get the right analysis and accomplish repairs.
batmanapollo.ru
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~demon19691
https://verda1967.diary.ru/
http://www.nfomedia.com/profile?uid=rOiUejJ
https://imageevent.com/leemura1967
https://www.haikudeck.com/presentations/twsZcsgd4R
You made some good points there. I did a search on the issue and found most people will consent with your site.
Interesting blog post. What I would like to bring up is that laptop or computer memory ought to be purchased in case your computer can no longer cope with whatever you do along with it. One can add two RAM memory boards with 1GB each, by way of example, but not certainly one of 1GB and one of 2GB. One should make sure the manufacturer’s documentation for own PC to be sure what type of memory space it can take.
Hi to every single one, it’s in fact a pleasant for me to go to see this site, it consists of important Information.
I was suggested this blog by my cousin. I’m not sure whether this post is written by him as nobody else know such detailed about my problem. You are wonderful! Thanks!
В нашем обществе, где диплом становится началом успешной карьеры в любом направлении, многие пытаются найти максимально быстрый и простой путь получения образования. Наличие документа об образовании переоценить попросту невозможно. Ведь диплом открывает двери перед людьми, стремящимися начать трудовую деятельность или продолжить обучение в университете.
В данном контексте наша компания предлагает максимально быстро получить этот необходимый документ. Вы имеете возможность купить диплом, и это является выгодным решением для всех, кто не смог завершить образование, потерял документ или желает исправить свои оценки. дипломы выпускаются с особой тщательностью, вниманием к мельчайшим элементам, чтобы на выходе получился документ, 100% соответствующий оригиналу.
Преимущества данного подхода состоят не только в том, что вы максимально быстро получите свой диплом. Весь процесс организовывается комфортно, с нашей поддержкой. Начиная от выбора подходящего образца до консультации по заполнению персональной информации и доставки в любое место страны — все находится под полным контролем опытных мастеров.
В результате, для всех, кто пытается найти максимально быстрый способ получить требуемый документ, наша услуга предлагает отличное решение. Заказать диплом – значит избежать продолжительного обучения и сразу перейти к личным целям: к поступлению в ВУЗ или к началу трудовой карьеры.
https://clicktr.com/blog/yeni-baslayanlar-icin-affiliate-marketing/?unapproved=58669&moderation-hash=181966edce70704f846eeb40a77456ed#comment-58669
http://emseyi.com/index.php?qa=user&qa_1=ujaqi
http://protprom.ru/index.php?subaction=userinfo&user=alyxapo
http://avestalaw.ru/index.php?subaction=userinfo&user=oluvade
http://e-audit.gefest.me/index.php?subaction=userinfo&user=okoje
If you would like to improve your knowledge simply keep
visiting this website and be updated with the hottest information posted here.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/gecnj1959/profile
https://launchpad.net/~ghostxxx19691
https://rentry.org/9xx5z8x7
https://manferty1956.bandcamp.com/album/aimees-first-massage
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190367
Hello there, There’s no doubt that your web site
may be having web browser compatibility problems.
Whenever I look at your web site in Safari, it looks
fine but when opening in I.E., it has some overlapping issues.
I just wanted to give you a quick heads up!
Aside from that, wonderful blog!
fantastic submit, very informative. I ponder why the other
specialists of this sector don’t realize this.
You should continue your writing. I’m sure, you have a great readers’ base already!
Hi, i read your blog occasionally and i own a similar one
and i was just wondering if you get a lot of spam remarks?
If so how do you prevent it, any plugin or anything you can suggest?
I get so much lately it’s driving me mad so any help is very much appreciated.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/infan-196-3
https://permacultureglobal.org/users/56839-amber-butler
https://haveagood.holiday/users/347110
https://www.metal-archives.com/users/sanek22661998
http://www.babelcube.com/user/brandi-hollis-1
My brother suggested I would possibly like this website.
He used to be entirely right. This publish actually made my day.
You can not consider just how a lot time I had spent for this info!
Thank you!
В современном мире, где диплом является началом отличной карьеры в любом направлении, многие стараются найти максимально быстрый путь получения качественного образования. Наличие официального документа переоценить невозможно. Ведь именно он открывает дверь перед каждым человеком, желающим начать трудовую деятельность или учиться в университете.
В данном контексте наша компания предлагает очень быстро получить любой необходимый документ. Вы можете купить диплом нового или старого образца, и это становится выгодным решением для человека, который не смог завершить образование или потерял документ. диплом изготавливается аккуратно, с особым вниманием ко всем деталям, чтобы на выходе получился полностью оригинальный документ.
Преимущество данного решения состоит не только в том, что можно максимально быстро получить диплом. Процесс организован просто и легко, с нашей поддержкой. Начав от выбора необходимого образца до консультаций по заполнению персональных данных и доставки по стране — все будет находиться под полным контролем качественных специалистов.
Всем, кто ищет максимально быстрый способ получения требуемого документа, наша компания предлагает выгодное решение. Купить диплом – значит избежать длительного процесса обучения и не теряя времени переходить к своим целям: к поступлению в ВУЗ или к началу удачной карьеры.
http://www.ntsr.info/forum/user/91072/
http://book-club.rggu.ru/2013/10/blog-post.html
http://forum.farmer.pl/profile/150015-akotud/
http://islamdini.kg/index.php?subaction=userinfo&user=ahaze
http://u-cars.ru/modules.php?name=Your_Account&op=userinfo&username=icasujag
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/344063
https://tubeteencam.com/user/jloseg1956/profile
https://melamory1983.diary.ru/
https://www.hentai-foundry.com/user/lisaxx1981/profile
http://www.babelcube.com/user/rachel-jones-1
I used to be recommended this blog by means of my cousin. I am now not sure whether or
not this post is written by way of him as no one else know such particular about my difficulty.
You’re incredible! Thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~isadron219751
https://imageevent.com/redlof1953
http://www.nfomedia.com/profile?uid=rOiUdiG
https://rentry.org/2bgoxfyo
https://onedio.ru/profile/dreadlight-197-3
Howdy just wanted to give you a quick heads up.
The words in your content seem to be running off the screen in Chrome.
I’m not sure if this is a formatting issue or something to do with browser compatibility but I thought I’d post to let you know.
The layout look great though! Hope you get the issue solved soon. Many thanks
В наше время, когда диплом является началом отличной карьеры в любом направлении, многие ищут максимально быстрый путь получения качественного образования. Важность наличия документа об образовании переоценить просто невозможно. Ведь диплом открывает двери перед каждым человеком, который стремится начать профессиональную деятельность или учиться в любом институте.
Мы предлагаем очень быстро получить этот важный документ. Вы имеете возможность купить диплом старого или нового образца, что становится выгодным решением для человека, который не смог завершить образование или потерял документ. дипломы изготавливаются с особой аккуратностью, вниманием к мельчайшим деталям, чтобы на выходе получился продукт, 100% соответствующий оригиналу.
Превосходство этого подхода заключается не только в том, что вы быстро получите диплом. Процесс организовывается просто и легко, с нашей поддержкой. От выбора подходящего образца документа до консультации по заполнению личной информации и доставки по России — все будет находиться под полным контролем квалифицированных специалистов.
Для всех, кто ищет быстрый и простой способ получения требуемого документа, наша компания предлагает отличное решение. Купить диплом – это значит избежать продолжительного обучения и сразу перейти к своим целям, будь то поступление в ВУЗ или старт трудовой карьеры.
http://jinsanbag.com/bbs/board.php?bo_table=free&wr_id=160792
http://www.diywiki.org/index.php?title=landsdiploms
http://ilovecovers.com/product/blow-your-house-down/?unapproved=296594&moderation-hash=9a94b0f89f261b2259020ee657d6bbf6#comment-296594
http://autoschool-progress.kz/index.php?subaction=userinfo&user=omylusi
http://mos-it.ru/index.php?subaction=userinfo&user=ugawod
Hi there are using WordPress for your site platform? I’m new to the blog world but
I’m trying to get started and create my own. Do you require any coding
knowledge to make your own blog? Any help would be really appreciated!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/featjem19961985/profile
https://www.hentai-foundry.com/user/robingod21976/profile
http://www.babelcube.com/user/rachel-lopez
https://rentry.org/wxp8yows
https://permacultureglobal.org/users/56246-christina-montesini
Fantastic blog! Do you have any helpful hints for aspiring writers?
I’m planning to start my own blog soon but I’m a little lost on everything.
Would you advise starting with a free platform like
Wordpress or go for a paid option? There are so many options
out there that I’m totally confused .. Any tips? Cheers!
Wonderful blog! Do you have any suggestions for aspiring writers?
I’m hoping to start my own site soon but I’m a little lost on everything.
Would you propose starting with a free platform like WordPress or go for
a paid option? There are so many choices out there that
I’m totally overwhelmed .. Any suggestions? Thanks a lot!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190663
https://www.hentai-foundry.com/user/eviji251979/profile
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3192283
http://www.babelcube.com/user/monica-bailey
https://www.divephotoguide.com/user/jasmen1991
Hi there outstanding website! Does running a blog similar to this take a large amount of work?
I have virtually no knowledge of coding but I was hoping to start my
own blog in the near future. Anyway, should you have any ideas or tips
for new blog owners please share. I understand this is off subject nevertheless I simply wanted to ask.
Appreciate it!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/hghgfhf1981/profile
https://haveagood.holiday/users/348001
https://www.obesityhelp.com/members/ter1t1995/about_me/
https://haveagood.holiday/users/347497
https://www.haikudeck.com/presentations/ELmIVG1dlv
На сегодняшний день, когда диплом – это начало успешной карьеры в любой отрасли, многие ищут максимально быстрый и простой путь получения образования. Наличие официального документа трудно переоценить. Ведь диплом открывает двери перед всеми, кто стремится вступить в профессиональное сообщество или учиться в университете.
Наша компания предлагает оперативно получить этот необходимый документ. Вы сможете приобрести диплом старого или нового образца, что становится отличным решением для человека, который не смог закончить обучение, утратил документ или хочет исправить плохие оценки. диплом изготавливается с особой тщательностью, вниманием к мельчайшим деталям. В результате вы получите документ, полностью соответствующий оригиналу.
Преимущества данного решения заключаются не только в том, что можно оперативно получить свой диплом. Весь процесс организован комфортно, с нашей поддержкой. От выбора подходящего образца до грамотного заполнения персональной информации и доставки в любое место страны — все будет находиться под абсолютным контролем качественных специалистов.
Для всех, кто ищет быстрый способ получить необходимый документ, наша услуга предлагает отличное решение. Купить диплом – это значит избежать долгого обучения и не теряя времени перейти к достижению личных целей: к поступлению в ВУЗ или к началу трудовой карьеры.
http://images.google.ac/url?q=https://rudiplomista.com
https://clients1.google.mn/url?q=https://lands-diploma.com
http://maps.google.vg/url?q=https://premialnie-diplomik.com
https://www.google.hn/url?sa=t&url=https://lands-diploma.com
https://europe.google.com/url?q=https://lands-diploma.com
Nice replies in return of this difficulty with
real arguments and explaining everything about that.
OLaneViart
Добрый день!
Но благодаря ресурсам в интернете, моя работа над дипломом стала намного эффективнее.
У нас вы можете приобрести диплом университета без предоплаты с доставкой в любой город России, оплачивая после получения.
Желаю всем честных оценок!
купить диплом программиста
купить диплом в петропавловске-камчатском
купить диплом в нижним тагиле
купить диплом нового образца
купить диплом в москве
купить диплом в находке
купить диплом в кунгуре
купить диплом в бердске
куплю диплом с занесением
купить диплом в новошахтинске
I blog quite often and I seriously appreciate your information. The article has truly peaked
my interest. I am going to book mark your site and keep checking
for new details about once a week. I opted
in for your Feed too.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/cohuwko1989
http://www.babelcube.com/user/johnny-barton
https://haveagood.holiday/users/348380
https://tubeteencam.com/user/manaev1993/profile
https://www.divephotoguide.com/user/avelardos1952
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://holdey1988.diary.ru/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191439
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191629
https://onedio.ru/profile/aramo-196-4
https://launchpad.net/~lasdafgn19731
Excellent post. I used to be checking continuously
this weblog and I’m inspired! Very helpful info specially the
closing part 🙂 I care for such information much. I used to be looking for this particular info for a
long time. Thanks and good luck.
В современном мире, где диплом является началом успешной карьеры в любом направлении, многие ищут максимально простой путь получения образования. Важность наличия документа об образовании сложно переоценить. Ведь именно он открывает двери перед каждым человеком, который стремится начать трудовую деятельность или учиться в университете.
В данном контексте мы предлагаем оперативно получить любой необходимый документ. Вы имеете возможность приобрести диплом старого или нового образца, что становится удачным решением для всех, кто не смог закончить обучение или утратил документ. диплом изготавливается с особой аккуратностью, вниманием к мельчайшим элементам. В результате вы получите продукт, максимально соответствующий оригиналу.
Плюсы этого подхода заключаются не только в том, что можно быстро получить диплом. Весь процесс организован комфортно, с нашей поддержкой. От выбора требуемого образца документа до консультации по заполнению личных данных и доставки в любое место России — все находится под абсолютным контролем качественных специалистов.
Всем, кто пытается найти оперативный способ получения необходимого документа, наша компания предлагает выгодное решение. Приобрести диплом – это значит избежать длительного процесса обучения и сразу переходить к достижению собственных целей: к поступлению в университет или к началу удачной карьеры.
https://directolog.com/member.php?14665-ymuweji
http://www.idsys.kr/bbs/board.php?bo_table=free&wr_id=473979
http://www.hagi.co.kr/bbs/board.php?bo_table=free&wr_id=248448
http://www.koelnmedia2.de/fastelovend/member.php?action=showprofile&user_id=14409
http://kazrti.kz/index.php?subaction=userinfo&user=upocubo
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/57279-david-bragg
http://www.babelcube.com/user/michelle-harvey
https://www.metal-archives.com/users/prios1967
https://www.quia.com/profiles/j363stagg
http://www.babelcube.com/user/robert-johnson-2
I always emailed this website post page to all my friends, as if like to
read it after that my contacts will too.
What i don’t realize is in fact how you’re now not actually a lot more smartly-favored than you may
be now. You’re so intelligent. You realize thus considerably on the subject of this matter, made me in my
opinion imagine it from numerous various angles. Its like women and men aren’t involved except it is something to
do with Girl gaga! Your personal stuffs great. All the time
handle it up!
It’s genuinely very difficult in this busy life to
listen news on Television, thus I just use world wide web for that reason, and get
the hottest news.
I am genuinely thankful to the holder of this web page
who has shared this enormous post at at this place.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://entalphy1972.diary.ru/
https://chyoa.com/user/reach9971980
https://okwave.jp/profile/u3109554.html
https://haveagood.holiday/users/344913
http://www.babelcube.com/user/sapna-tapia
Hey! I could have sworn I’ve been to this blog
before but after checking through some of the post I realized it’s new
to me. Nonetheless, I’m definitely happy I found
it and I’ll be bookmarking and checking back frequently!
OLaneViart
Приветики, дорогие мои!!
Диплом продолжает быть моим вызовом, но теперь у меня есть все необходимые ресурсы для его успешного завершения.
Закажите диплом ВУЗа России недорого, без предоплаты и с гарантией возврата средств.
Желаю вам всем не двоешных) отметок!
купить диплом о среднем
купить диплом учителя
купить диплом в глазове
купить диплом в ессентуках
купить диплом в нижнем тагиле
https://www.google.co.ug/url?q=https://diploms-vuza.com/
https://www.google.com.ni/url?sa=t&url=https://kazdiplomas.com/
https://maps.google.com.ar/url?q=https://diploms-vuza.com/
https://images.google.jo/url?q=https://diplom-bez-problem.com/
http://images.google.com.gh/url?sa=t&url=https://1magistr.ru/
На сегодняшний день, когда диплом становится началом успешной карьеры в любом направлении, многие стараются найти максимально быстрый и простой путь получения качественного образования. Факт наличия официального документа переоценить невозможно. Ведь диплом открывает двери перед людьми, желающими вступить в профессиональное сообщество или учиться в университете.
Наша компания предлагает максимально быстро получить этот важный документ. Вы имеете возможность приобрести диплом нового или старого образца, что является выгодным решением для человека, который не смог закончить обучение или утратил документ. дипломы выпускаются с особой аккуратностью, вниманием ко всем деталям, чтобы на выходе получился продукт, 100% соответствующий оригиналу.
Превосходство такого решения заключается не только в том, что можно максимально быстро получить диплом. Весь процесс организован комфортно, с профессиональной поддержкой. От выбора нужного образца диплома до правильного заполнения персональной информации и доставки по России — все под полным контролем качественных мастеров.
Всем, кто ищет оперативный способ получения требуемого документа, наша компания может предложить отличное решение. Заказать диплом – значит избежать длительного процесса обучения и сразу переходить к достижению личных целей, будь то поступление в ВУЗ или старт карьеры.
http://proffpereezd.ru/index.php?subaction=userinfo&user=ylirede
http://loket.kr/free/32874
http://advanced-retail.a2hosted.com/doku.php?id=russianydiplomans
http://s-s-o.ru/forum.php?PAGE_NAME=profile_view&UID=50809
https://yozhstudio.ru/2019/09/08/%d0%bf%d1%80%d0%b5%d0%b8%d0%bc%d1%83%d1%89%d0%b5%d1%81%d1%82%d0%b2%d0%b0-%d0%b8%d0%bd%d0%b4%d0%b8%d0%b2%d0%b8%d0%b4%d1%83%d0%b0%d0%bb%d1%8c%d0%bd%d0%be%d0%b3%d0%be-%d0%bf%d0%be%d1%88%d0%b8%d0%b2%d0%b0/?unapproved=184912&moderation-hash=ae1a9c36dfa4d1e49a247032c897fa63#comment-184912
I do trust all of the concepts you’ve presented for your post.
They are very convincing and can definitely work.
Nonetheless, the posts are too quick for starters.
Could you please lengthen them a little from subsequent time?
Thank you for the post.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://kirkam1968.diary.ru/
https://anotepad.com/notes/c9nj6xb9
http://www.babelcube.com/user/freddie-mcanelly
https://chyoa.com/user/fogpro31973
https://okwave.jp/profile/u3113004.html
After I originally commented I seem to have clicked on the -Notify me when new comments are added- checkbox and now whenever a comment is added I receive four emails with the exact same comment.
There has to be a means you are able to remove me from that
service? Thank you!
В нашем мире, где диплом – это начало успешной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения качественного образования. Наличие официального документа об образовании сложно переоценить. Ведь диплом открывает дверь перед людьми, желающими начать трудовую деятельность или продолжить обучение в любом университете.
Мы предлагаем оперативно получить этот важный документ. Вы можете приобрести диплом, что является выгодным решением для человека, который не смог закончить обучение или потерял документ. Каждый диплом изготавливается аккуратно, с особым вниманием ко всем нюансам. На выходе вы сможете получить полностью оригинальный документ.
Преимущества данного подхода заключаются не только в том, что вы сможете оперативно получить диплом. Процесс организован удобно и легко, с нашей поддержкой. От выбора требуемого образца документа до точного заполнения персональных данных и доставки в любое место России — все под абсолютным контролем квалифицированных специалистов.
Таким образом, для тех, кто пытается найти быстрый способ получения требуемого документа, наша услуга предлагает выгодное решение. Приобрести диплом – значит избежать продолжительного обучения и сразу перейти к достижению своих целей: к поступлению в ВУЗ или к началу удачной карьеры.
https://maps.google.la/url?q=https://lands-diploma.com
https://images.google.es/url?q=https://premialnie-diplomik.com
https://maps.google.com.mm/url?sa=t&url=https://rudiplomista.com
https://images.google.es/url?q=https://lands-diploma.com
https://www.google.cm/url?sa=i&rct=j&q=https://premialnie-diplomik.com
I will immediately clutch your rss as I can not to find your email
subscription link or newsletter service. Do you have any?
Kindly allow me know so that I may subscribe. Thanks.
Useful info. Lucky me I found your web site accidentally, and
I am surprised why this accident didn’t took place earlier!
I bookmarked it.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/150525
https://www.divephotoguide.com/user/arguz1984
https://tubeteencam.com/user/sasukemix1996/profile
https://www.hentai-foundry.com/user/bandit851956/profile
https://launchpad.net/~powergirl19601
I really love your site.. Excellent colors & theme. Did
you develop this web site yourself? Please reply
back as I’m attempting to create my own website and would love to find out where you got this from or exactly what the theme is called.
Kudos!
mulong4d I am very interested in the article, I feel this article is very useful and useful for many other people.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/nikpoman1977
https://www.quia.com/profiles/pmarie106
https://www.hentai-foundry.com/user/geliot1994/profile
https://imageevent.com/b00sinka1999
https://www.haikudeck.com/presentations/FgPK2Qcll5
Good way of telling, and fastidious article to take
facts on the topic of my presentation topic, which i am going to present in school.
Ремонт электродвигателей
: основные этапы процедуры
Электродвигатель — это основной рабочий узел во многих бытовых и промышленных устройствах. Чтобы он исправно служил долгое время, необходимо регулярно проводить квалифицированное техническое обслуживание. Однако даже самое надёжное оборудование со временем изнашивается, и тогда требуется ремонт электродвигателя.
Основные этапы процедуры ремонта электродвигателя включают:
Очистка от загрязнений: электродвигатель очищают от пыли и грязи с помощью сжатого воздуха и тряпки, смоченной в чистом бензине.
Выявление внешних повреждений: на этом этапе обнаруживаются возможные причины поломки оборудования.
Снятие защитных кожухов и корпуса: после очистки можно приступать к разборке электродвигателя.
Проверка состояния механических узлов: проверяют состояние подшипников, вала и других механических элементов.
Демонтаж вышедших из строя подшипников и запрессовка новых: если обнаружены дефекты, подшипники заменяют на новые.
Перемотка электродвигателя: если проблема связана с повреждением статора или якоря, производят перемотку двигателя.
Для проведения качественного ремонта электродвигателя рекомендуется обращаться в специализированные мастерские или сервисные центры.
Перемотка электродвигателей: основные этапы и особенности процесса
Перемотка электродвигателей — это процесс замены старой или повреждённой обмотки на новую. Она может потребоваться в различных ситуациях, например, при износе рабочих обмоток, межвитковом пробое изоляции, коротком замыкании витков или изменении рабочего напряжения.
Этапы перемотки электродвигателя:
Дефектация: визуальный осмотр двигателя, определение наличия вмятин, царапин и оценка состояния существующих обмоток.
Удаление старых обмоток: срезание бандажных креплений и фиксация схемы соединения обмоток.
Очистка пазов статора: освобождение пазов от старой обмотки и очистка от наплывов лака, остатков изолирующих материалов.
Монтаж новых изолирующих прокладок: установка прокладок в пазы статора.
Намотка новых катушечных групп: на специальном оборудовании наматываются новые катушки, которые затем размещаются в пустых пазах статора и фиксируются.
Укладка обмоток: установка межкатушечных изолирующих элементов и обвязки (бандажа).
Подключение катушек согласно схеме: проверка электрических параметров и замыкание на корпус.
Пропитка лаком: статор пропитывается лаком для улучшения изоляционных свойств.
Полное отверждение лака и финишный контроль параметров: контроль напряжения пробоя и механических характеристик.
Механическая сборка двигателя и подключение главных выводов обмоток к клеммам: завершение процесса перемотки.
После завершения всех этапов перемотки проводится тестовый прогон электродвигателя для проверки его работоспособности.
Балансировка — это процесс устранения дисбаланса ротора, который возникает из-за неравномерного распределения массы или дефектов конструкции. Это важная процедура, которая помогает предотвратить повышенный износ подшипников, повреждение ротора и снижение эффективности работы двигателя.
Причины потери балансировки могут быть разными: ремонт ротора, заводской брак или работа двигателя в тяжёлых условиях с превышением паспортных значений нагрузки.
Существует два основных метода балансировки электродвигателей: статический и динамический. Статический метод используется для устранения основного дисбаланса, а динамический метод обеспечивает максимальную точность и применяется для балансировки двигателей, работающих на высоких оборотах.
В процессе динамической балансировки специальное оборудование раскручивает ротор электродвигателя и указывает точки дисбаланса с помощью датчиков. Добавление или уменьшение массы ротора в этих точках позволяет достичь максимальной балансировки.
Для некоторых моделей больших и мощных электродвигателей применяется только статический метод балансировки из-за невозможности выполнения динамической балансировки.
Качественная балансировка электродвигателя позволяет избежать проблем с вибрацией, повышенным износом и повреждением оборудования, связанного с работой неисправного двигателя.
Attractive section of content. I just stumbled upon your website
and in accession capital to assert that I acquire in fact enjoyed account your
blog posts. Anyway I will be subscribing to your feeds and even I achievement you access
consistently quickly.
It’s a pity you don’t have a donate button! I’d certainly donate
to this outstanding blog! I guess for now i’ll
settle for book-marking and adding your RSS feed to my Google account.
I look forward to new updates and will talk about this
website with my Facebook group. Talk soon!
Hmm is anyone else having problems with the pictures on this blog loading?
I’m trying to figure out if its a problem on my end or if it’s the blog.
Any suggestions would be greatly appreciated.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/427ze5mn
https://chyoa.com/user/kease1951
http://www.nfomedia.com/profile?uid=rOiVhiD
https://www.divephotoguide.com/user/battosai1989
https://ellak.gr/user/dainty1959/
wonderful points altogether, you just won a brand new
reader. What might you suggest in regards to your publish
that you just made some days ago? Any sure?
Hey I am so thrilled I found your web site, I really found you by
error, while I was looking on Digg for something else, Anyhow I am here now and would just like to say many thanks for a incredible post and a all round exciting blog (I also love the theme/design), I don’t have time to
go through it all at the moment but I have book-marked it
and also added in your RSS feeds, so when I have time I will be back to
read much more, Please do keep up the excellent work.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/holeymole1978
http://www.babelcube.com/user/serena-hitzmann-1
https://rentry.org/2m3umyxr
https://cannabis.net/user/148816
http://www.babelcube.com/user/whitney-hunter
Dichaelautob
Всем хорошего дня!
Диплом потребовал от меня работы ночью, что плохо сказалось на здоровье.
Заказать и купить диплом в России без предоплаты и с гарантией можно только у нас, гарантия “под ключ”.
Желаю каждому прекрасных оценок!
купить диплом в оренбурге
купить диплом повара
купить диплом в смоленске
купить диплом в уфе
купить диплом в кемерово
http://alt1.toolbarqueries.google.co.ve/url?q=https://diplom-insti.ru/
http://maps.google.com.jm/url?q=https://diplomoz-197.com/
https://cse.google.bj/url?q=https://diploms-vuza.com/
https://cse.google.dz/url?sa=t&url=https://ry-diplom.com/
https://www.google.co.ls/url?q=https://diplom-profi.ru/
etsyweddingteam.com
素晴らしい記事!いつも新鮮な視点をありがとう。
It is perfect time to make some plans for the future and it
is time to be happy. I have read this post and if I could I desire to suggest you few interesting things or suggestions.
Maybe you could write next articles referring to this article.
I wish to read more things about it!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/ray-castillo
https://chyoa.com/user/evolve1990
https://www.metal-archives.com/users/saw1962
https://okwave.jp/profile/u3111123.html
https://www.metal-archives.com/users/kosy1981
Woah! I’m really digging the template/theme of this
website. It’s simple, yet effective. A lot of times
it’s hard to get that “perfect balance” between superb
usability and visual appeal. I must say you have done a excellent
job with this. Also, the blog loads very quick for me on Internet explorer.
Superb Blog!
Helpful information. Lucky me I found your site accidentally, and I
am surprised why this twist of fate didn’t happened in advance!
I bookmarked it.
Букмекерская контора 1win – одна из самых популярных площадок, где пользователи могут делать ставки, играть, делать ставки и т. д. Для привлечения новой аудитории данная букмекерская контора предлагает новичкам отличный бонус – возможность получить до 200 000 бонусов за 4 депозита. И для этого покупателям даже не нужно вводить промокоды. Вам просто нужно зарегистрироваться в этом сервисе.
Промокод 1Win 2024: m1WIN2024 — это уникальный код, который необходимо указать при регистрации для получения бонуса 500% до 75 000 рублей. Это предложение доступно только новым игрокам, которые могут претендовать на приветственный бонус 1Win.
Для постоянных клиентов букмекерская контора постоянно выпускает новые промокоды 1win, ведь с этими бонусами клиентам гораздо приятнее пользоваться услугами этой букмекерской конторы. Промокод – это уникальный набор букв и цифр, активация которого позволяет человеку получить бонус. В этом обзоре мы расскажем, где взять новые промокоды 1win и как их активировать для получения бонусов.
Актуальный промокод 1Win 2024 вы можете найти на различных страницах с информацией о бонусах в букмекерских конторах. Продажи также осуществляются через партнеров компании. Лучшее место для поиска купонов – Telegram-канал букмекерской конторы. Новые ваучеры появляются там каждый день. 1Win может отправить промокод индивидуально уже зарегистрированному клиенту. Например, по случаю годовщины регистрации или просто дня рождения клиента.
С промокодом 1WIN новые игроки могут значительно увеличить сумму своего первого и последующих депозитов. Полученные бонусы можно использовать в игре и в случае успеха перевести на свой электронный кошелек. Максимальная сумма бонуса – 75 000 рублей.
Отдельной вкладки для проверки комбинаций нет. Если введено правильно, система активирует бонусное предложение. Во вкладке «Ваучер» в личном кабинете появится сообщение при вводе промокода 1Vin. Отсюда вы сможете увидеть, правильно ли была введена комбинация.
Источник: https://mmocenter.ru/blog/promokod-1win-promokody-1vin-pri-registracii-na-segodnya/
Appreciate this post. Will try it out.
I have been exploring for a bit for any high quality articles
or blog posts on this kind of area . Exploring in Yahoo
I eventually stumbled upon this web site. Reading this info So i’m happy to express that I have an incredibly just right
uncanny feeling I discovered just what I needed.
I so much undoubtedly will make sure to don?t forget this web
site and give it a glance on a continuing basis.
Thanks in favor of sharing such a nice idea, piece of
writing is nice, thats why i have read it fully
Сегодня, когда диплом является началом отличной карьеры в любой области, многие ищут максимально быстрый и простой путь получения качественного образования. Факт наличия официального документа сложно переоценить. Ведь диплом открывает дверь перед любым человеком, желающим начать профессиональную деятельность или продолжить обучение в ВУЗе.
Наша компания предлагает очень быстро получить этот важный документ. Вы сможете купить диплом старого или нового образца, что будет отличным решением для человека, который не смог завершить образование или потерял документ. диплом изготавливается аккуратно, с максимальным вниманием к мельчайшим деталям. В результате вы сможете получить документ, максимально соответствующий оригиналу.
Плюсы подобного подхода состоят не только в том, что можно оперативно получить свой диплом. Весь процесс организовывается удобно, с профессиональной поддержкой. Начиная от выбора подходящего образца диплома до консультации по заполнению личных данных и доставки в любой регион страны — все будет находиться под полным контролем качественных мастеров.
Всем, кто ищет быстрый и простой способ получить требуемый документ, наша компания готова предложить отличное решение. Купить диплом – это значит избежать долгого процесса обучения и сразу перейти к достижению своих целей: к поступлению в университет или к началу удачной карьеры.
https://cse.google.ne/url?sa=i&url=https://lands-diploma.com
https://cse.google.com.lb/url?sa=t&url=https://premialnie-diplomik.com
https://clients1.google.sn/url?sa=t&url=https://rudiplomista.com
https://images.google.tl/url?sa=t&url=https://premialnie-diplomik.com
https://image.google.co.ck/url?sa=j&source=web&rct=j&url=https://lands-diploma.com
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/losttim1984
https://www.credly.com/users/sam-henry.32f0ea85/badges
https://www.hentai-foundry.com/user/q3koshmar1956/profile
http://www.babelcube.com/user/agunk-simmons
https://www.quia.com/profiles/sarahkaur
Hello there! I know this is kinda off topic but I was wondering which blog platform are you using for this website?
I’m getting tired of WordPress because I’ve had problems
with hackers and I’m looking at alternatives for another platform.
I would be fantastic if you could point me in the direction of
a good platform.
My relatives every time say that I am wasting my time here at web, but
I know I am getting familiarity all the time by reading such
pleasant content.
It’s really a cool and useful piece of information. I’m happy
that you shared this useful information with us. Please keep us up to
date like this. Thank you for sharing.
With havin so much content and articles do you ever run into any problems of plagorism or copyright violation? My site has a lot of completely unique content
I’ve either authored myself or outsourced but it looks like a lot of it is popping it up
all over the internet without my permission. Do you
know any solutions to help protect against content from being stolen? I’d genuinely appreciate it.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://akapi1957.diary.ru/
https://www.metal-archives.com/users/denis19291965
https://chyoa.com/user/fogpro31973
https://tubeteencam.com/user/defain1956/profile
https://haveagood.holiday/users/343944
Why users still make use of to read news papers when in this technological world the whole thing is existing on web?
I’m not sure exactly why but this web site is loading extremely slow for me.
Is anyone else having this problem or is it a issue on my end?
I’ll check back later on and see if the problem still exists.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/HMFj9H2JjH
https://sentinell1950.diary.ru/
https://tubeteencam.com/user/gecnj1959/profile
https://www.haikudeck.com/presentations/RuOtRHPi5F
http://www.babelcube.com/user/wendy-taylor
Excellent article. Keep posting such kind of info on your
page. Im really impressed by it.
Hi there, You have performed a fantastic job.
I will certainly digg it and for my part suggest to my friends.
I am sure they will be benefited from this site.
I do agree with all the ideas you’ve introduced in your post.
They’re really convincing and will certainly work. Nonetheless, the posts are very quick for newbies.
Could you please extend them a bit from next time?
Thanks for the post.
Having read this I thought it was very informative.
I appreciate you taking the time and energy to put this short article together.
I once again find myself spending a lot of time both reading and posting comments.
But so what, it was still worthwhile!
I always emailed this weblog post page to all my associates, since if like to read
it afterward my links will too.
I got this web site from my buddy who told me regarding this web page and at the moment this time I am visiting this web page and reading very informative content here.
http://www.google.gr/url?q=https://ry-diplom.com/
https://maps.google.com.pg/url?rct=j&sa=t&url=https://kazdiplomas.com/
https://www.google.com.gh/url?q=https://diplomoz-197.com/
http://images.google.co.ck/url?q=https://diplom-bez-problem.com/
http://www.google.is/url?sa=t&url=https://diplom-profi.ru/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/comablue1973/profile
https://www.haikudeck.com/presentations/2BfokujAX9
https://ellak.gr/user/sylker1968/
https://haveagood.holiday/users/346204
https://tubeteencam.com/user/mm1081999/profile
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/vjgu001979
https://anotepad.com/notes/xwi8gfi5
https://www.quia.com/profiles/mlively205
https://www.metal-archives.com/users/coppa1950
https://www.haikudeck.com/presentations/ELmIVG1dlv
На сегодняшний день, когда диплом – это начало успешной карьеры в любой отрасли, многие ищут максимально быстрый и простой путь получения качественного образования. Наличие документа об образовании сложно переоценить. Ведь именно он открывает дверь перед любым человеком, желающим вступить в сообщество профессионалов или продолжить обучение в каком-либо институте.
В данном контексте наша компания предлагает очень быстро получить любой необходимый документ. Вы сможете купить диплом нового или старого образца, и это является удачным решением для всех, кто не смог завершить обучение, утратил документ или желает исправить плохие оценки. дипломы выпускаются аккуратно, с особым вниманием к мельчайшим нюансам. На выходе вы получите документ, максимально соответствующий оригиналу.
Преимущество такого подхода заключается не только в том, что можно оперативно получить свой диплом. Процесс организовывается удобно, с нашей поддержкой. От выбора подходящего образца до правильного заполнения персональной информации и доставки по России — все под абсолютным контролем качественных специалистов.
Для тех, кто ищет оперативный способ получить требуемый документ, наша компания предлагает отличное решение. Купить диплом – значит избежать длительного процесса обучения и сразу переходить к своим целям, будь то поступление в ВУЗ или начало успешной карьеры.
http://images.google.com.pg/url?q=https://lands-diploma.com
https://www.google.me/url?q=https://lands-diploma.com
http://cse.google.co.bw/url?sa=t&url=https://rudiplomista.com
https://cse.google.ci/url?q=https://lands-diploma.com
https://images.google.com.kh/url?q=https://lands-diploma.com
Hello to all, it’s actually a good for me to visit this web page, it
contains priceless Information.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/ramzesl1958/profile
https://www.hentai-foundry.com/user/psh1041992/profile
https://cannabis.net/mycannabis/p-basic-info/submit
https://tubeteencam.com/user/manifiko1986/profile
https://www.haikudeck.com/presentations/EralrRLAyx
Hi i am kavin, its my first occasion to commenting anywhere,
when i read this paragraph i thought i could also create
comment due to this sensible article.
game1kb.com
동시에 흥분한 사업가들은 다양한 사업 기회를 찾기 시작했습니다.
The other day, while I was at work, my sister stole my apple ipad
and tested to see if it can survive a forty foot drop,
just so she can be a youtube sensation. My apple ipad is now destroyed and she has 83 views.
I know this is entirely off topic but I had to share it with someone!
Having read this I thought it was really informative.
I appreciate you spending some time and effort to put this short article together.
I once again find myself spending a lot of time both reading and
commenting. But so what, it was still worthwhile!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/57226-sandy-petersen
https://www.obesityhelp.com/members/stysemm1982/about_me/
https://rentry.org/5hyrsygq
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3189198
https://www.divephotoguide.com/user/cyran1955
Why users still use to read news papers when in this technological globe all is presented on web?
If some one needs expert view concerning blogging and site-building afterward i recommend him/her
to pay a quick visit this website, Keep up the nice
job.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/valance1967
https://www.divephotoguide.com/user/feppapu1955
https://anotepad.com/notes/bh5wm86b
https://akapi1957.diary.ru/
https://www.hentai-foundry.com/user/roan971955/profile
I do believe all the ideas you have introduced in your post.
They are really convincing and will definitely work. Nonetheless, the posts are too short
for novices. May just you please extend them a little from next time?
Thank you for the post.
I am really inspired together with your writing talents as neatly as
with the structure to your blog. Is this
a paid subject or did you modify it yourself?
Anyway stay up the nice high quality writing, it’s rare to look a nice
weblog like this one nowadays..
I love what you guys are up too. Such clever work and exposure!
Keep up the awesome works guys I’ve you guys to my own blogroll.
I have fun with, result in I discovered exactly what I used to be having a look for.
You have ended my four day long hunt! God Bless you man. Have a
nice day. Bye
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/meexan1997
https://launchpad.net/~ilecfg19951
https://rentry.org/vb7fg4am
https://cannabis.net/user/149632
https://rockymars1985.diary.ru/
My brother suggested I would possibly like this blog. He was once
entirely right. This post actually made my day. You can not believe just how a lot time I had
spent for this info! Thanks!
Yes! Finally someone writes about fortune tiger.
Hello there! This is my 1st comment here so I just wanted to
give a quick shout out and say I genuinely enjoy reading your articles.
Can you suggest any other blogs/websites/forums that go over the
same subjects? Thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/56787-ana-williams
https://rentry.org/t5oony6e
https://ellak.gr/user/godmag1996/
https://www.quia.com/profiles/leslie460
https://tubeteencam.com/user/iiiuba1996/profile
IsmaelBog
Привет всем!
Мои усилия по диплому получили поддержку через полезные ресурсы, найденные в сети.
Закажите диплом ВУЗа России недорого, без предоплаты и с гарантией возврата средств.
Желаю всем прекрасных отметок!
купить диплом медбрата
купить диплом в воронеже
куплю диплом высшего образования
купить диплом в северске
купить диплом провизора
https://images.google.ml/url?sa=t&url=https://okdiplom.com/
https://cse.google.li/url?q=https://diplom45.ru/
https://cse.google.dz/url?sa=t&url=https://diplom-zakaz.ru/
https://cse.google.sm/url?q=https://bisness-diplom.ru/
https://www.google.com.np/url?sa=t&url=https://diplom5.com/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/dharma1081994/profile
http://www.nfomedia.com/profile?uid=rOiWcbK
https://anotepad.com/notes/p4gnyfeg
https://imageevent.com/livass1975
https://launchpad.net/~gi1g1ma1n19731
Thank you for sharing your thoughts. I really appreciate your efforts and I
will be waiting for your further write ups thank you once again.
Currently it sounds like WordPress is the top blogging platform available right now.
(from what I’ve read) Is that what you’re using on your blog?
I have read so many posts concerning the blogger lovers
however this paragraph is genuinely a pleasant piece of writing, keep
it up.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/lildream1964
https://launchpad.net/~wnyhtuk19741
https://ar2r1965.diary.ru/
https://www.haikudeck.com/presentations/EralrRLAyx
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191598
Link exchange is nothing else however it is only placing the other person’s weblog link on your page at
suitable place and other person will also do similar in favor of you.
В наше время, когда диплом становится началом отличной карьеры в любом направлении, многие пытаются найти максимально простой путь получения качественного образования. Важность наличия официального документа трудно переоценить. Ведь именно он открывает двери перед всеми, кто собирается начать профессиональную деятельность или учиться в высшем учебном заведении.
Мы предлагаем очень быстро получить этот необходимый документ. Вы имеете возможность заказать диплом, и это становится отличным решением для всех, кто не смог завершить обучение, потерял документ или желает исправить свои оценки. диплом изготавливается аккуратно, с особым вниманием ко всем деталям. В результате вы получите документ, 100% соответствующий оригиналу.
Плюсы данного решения состоят не только в том, что можно быстро получить свой диплом. Процесс организовывается просто и легко, с профессиональной поддержкой. Начиная от выбора нужного образца до грамотного заполнения личной информации и доставки по России — все будет находиться под абсолютным контролем квалифицированных специалистов.
Всем, кто ищет быстрый и простой способ получения требуемого документа, наша компания предлагает выгодное решение. Купить диплом – значит избежать длительного процесса обучения и сразу переходить к своим целям, будь то поступление в ВУЗ или начало карьеры.
https://toolbarqueries.google.co.tz/url?q=https://premialnie-diplomik.com
http://maps.google.com.om/url?q=https://premialnie-diplomik.com
https://images.google.dk/url?sa=t&url=https://premialnie-diplomik.com
https://cse.google.rw/url?sa=t&url=https://rudiplomista.com
https://clients1.google.bs/url?sa=t&url=https://lands-diploma.com
Great blog right here! Additionally your website a lot up fast!
What web host are you the usage of? Can I get your affiliate link for your host?
I desire my site loaded up as fast as yours lol
Hey there! Quick question that’s entirely off topic. Do you know how to make your site
mobile friendly? My website looks weird when viewing from my iphone4.
I’m trying to find a theme or plugin that might be able to correct this problem.
If you have any suggestions, please share. Many thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/waric1964
https://www.divephotoguide.com/user/druidmzf1971
https://rentry.org/7nymofhb
https://www.hentai-foundry.com/user/demwit1990/profile
https://launchpad.net/~dm12319781
Здравствуйте!
Закажите диплом ВУЗа с доставкой по всей России без предоплаты и с гарантией качества!
https://cse.google.tm/url?q=http://diplomrussian.com
https://clients1.google.rw/url?q=http://aurus-diploms.com
http://toolbarqueries.google.bf/url?sa=t&url=http://diplomsagroups.com
https://cse.google.tm/url?q=https://russian-diplom.com
http://maps.google.im/url?q=http://free-diplommi.com
Мы предлагаем приобрести диплом института или колледжа с доставкой по России без предоплаты.
Awesome blog! Do you have any suggestions for aspiring writers?
I’m hoping to start my own blog soon but I’m a little lost
on everything. Would you propose starting with a free platform like WordPress or go
for a paid option? There are so many options out there that I’m totally
overwhelmed .. Any tips? Thanks a lot!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/sakamot1950
https://rentry.org/wz39qqes
https://www.haikudeck.com/presentations/VmeuDYLxSI
http://www.nfomedia.com/profile?uid=rOiUhhH
https://www.hentai-foundry.com/user/linkfis1967/profile
Nice blog! Is your theme custom made or did you download
it from somewhere? A design like yours with a few simple tweeks would really make my blog jump out.
Please let me know where you got your design. Thank you
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/antimothe1992/about_me/
https://www.divephotoguide.com/user/schokk3421969
https://haveagood.holiday/users/348304
https://www.divephotoguide.com/user/muginec051996
https://imageevent.com/korvolol1990
LewisAluri
Привет всем!
Диплом стал моей основной заботой из-за нарушения установленных сроков.
Мы предлагаем всем желающим приобрести диплом университета России по доступной цене с доставкой “под ключ”.
Желаю для каждого положительных оценок!
купить аттестат за 11 класс
купить диплом в новоуральске
купить диплом в находке
купить диплом в барнауле
купить диплом в нижнем новгороде
http://alt1.toolbarqueries.google.so/url?q=https://okdiplom.com/
http://images.google.vg/url?q=https://diplom-zakaz.ru/
https://maps.google.hu/url?sa=t&url=https://prodiplome.com/
https://maps.google.bg/url?sa=i&source=web&rct=j&url=https://peoplediplom.ru/
https://maps.google.com.br/url?q=https://diplom5.com/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/narutta1989
https://www.obesityhelp.com/members/shane541956/about_me/
https://rentry.org/qyz5swt7
https://permacultureglobal.org/users/56654-jaclyn-yoon
https://chyoa.com/user/peredozz1999
What’s up to every one, since I am in fact eager of reading this website’s post
to be updated daily. It includes fastidious material.
After going over a number of the articles on your web page, I really appreciate your
technique of blogging. I book-marked it to my bookmark webpage list and
will be checking back soon. Please check out my web site too and tell me
how you feel.
I constantly spent my half an hour to read this webpage’s
content daily along with a mug of coffee.
Spot on with this write-up, I really feel this website needs much more attention. I’ll probably be back again to read more, thanks for
the information!
Приветики!
Закажите диплом института или техникума по доступной цене с гарантированной доставкой в любой уголок России без предоплаты!
https://cse.google.co.il/url?sa=t&url=https://russian-diplom.com
https://maps.google.cl/url?q=http://originality-diplomy.com
https://image.google.bs/url?q=http://diplomsagroups.com
https://images.google.co.za/url?q=http://diplomrussian.com
https://cse.google.cat/url?q=http://russdiplomiki.com
Приобретите российский диплом с гарантией качества и доставкой в любую точку страны без предварительной оплаты!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://friezer1980.diary.ru/
http://www.babelcube.com/user/markus-azuse
https://www.divephotoguide.com/user/basho1995
https://rentry.org/3dafrbnu
https://imageevent.com/nikpoman1977
Hello there! Do you know if they make any plugins to protect against hackers?
I’m kinda paranoid about losing everything I’ve worked hard on. Any
recommendations?
I could not resist commenting. Very well written!
Appreciate the recommendation. Will try it out.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/bdieker
https://www.hentai-foundry.com/user/narccop1955/profile
https://www.metal-archives.com/users/lenora1986
https://www.divephotoguide.com/user/uiipo1972
https://ellak.gr/user/tarytu1971/
Приветики!
Приобретите диплом института или колледжа недорого с доставкой по РФ и гарантией качества без предварительной оплаты!
http://www.google.gp/url?q=https://anny-diploms.com
https://toolbarqueries.google.td/url?sa=j&source=web&rct=j&url=http://rdiplomik24.com
http://images.google.co.nz/url?sa=t&url=https://eonline-diploms.com
http://maps.google.cg/url?q=https://diploms-asx.com
http://toolbarqueries.google.co.in/url?sa=t&url=http://diplomsagroups.com
Закажите диплом ВУЗа недорого с доставкой курьером по России без предоплаты – это быстро и надежно!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://spirit681985.diary.ru/
https://okwave.jp/profile/u3110910.html
https://haveagood.holiday/users/347616
https://imageevent.com/killer451955
https://imageevent.com/luckymike1953
Gichardadvox
Всем хорошего дня!
Диплом теперь развивается благодаря разнообразным ресурсам из сети.
На нашем сайте вы можете купить диплом ВУЗа с постоплатой и помощью 24/7.
Желаю для каждого честных оценок!
купить диплом в рубцовске
купить диплом в новоуральске
купить диплом парикмахера
купить диплом в когалыме
купить диплом журналиста
https://maps.google.ie/url?q=https://diploms-vuza.com/
https://www.google.com.uy/url?q=https://10000diplomov.ru/
https://maps.google.fr/url?q=https://kdiplom.ru/
https://clients1.google.com.bh/url?sa=t&url=https://kdiplom.ru/
http://images.google.rw/url?q=https://1magistr.ru/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3192511
https://anotepad.com/notes/28qdg2jf
https://www.obesityhelp.com/members/pelfox1963/about_me/
https://tubeteencam.com/user/tiraj1983/profile
https://launchpad.net/~bearddemon19621
Exceptional post however I was wondering if you could write a litte more on this subject?
I’d be very thankful if you could elaborate a little bit further.
Thank you!
Hi there! Do you know if they make any plugins to safeguard against hackers?
I’m kinda paranoid about losing everything I’ve worked hard on. Any recommendations?
Magnificent items from you, man. I have take into accout
your stuff prior to and you’re simply too excellent.
I actually like what you have got right here, certainly like what you
are stating and the way during which you are saying it. You’re making it entertaining and you
still take care of to keep it wise. I cant wait to read much more from you.
That is really a terrific website.
Today, I went to the beach with my children. I found a sea shell and gave it to
my 4 year old daughter and said “You can hear the ocean if you put this to your ear.” She put the shell to her ear and screamed.
There was a hermit crab inside and it pinched her ear. She never wants
to go back! LoL I know this is totally off topic but I had to tell someone!
After checking out a handful of the blog posts on your website, I
really like your way of blogging. I saved it to my bookmark website list and will be checking back soon. Take a look at
my web site as well and tell me how you feel.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/shamb6np
https://haveagood.holiday/users/347946
https://www.quia.com/profiles/andreawalls
https://tubeteencam.com/user/kahu1965/profile
https://chyoa.com/user/treecher1972
Добрый день всем!
Купите диплом института или колледжа без предоплаты с гарантированной доставкой в любой уголок России.
http://images.google.com.na/url?q=https://anny-diploms.com
https://toolbarqueries.google.mn/url?q=http://aurus-diploms.com
http://maps.google.tt/url?q=http://diploms-service.com
http://www.google.bt/url?q=http://originality-diplomy.com
https://www.google.mv/url?sa=t&url=http://rudiplomisty24.com
Получите документы об образовании ВУЗов России с доставкой по РФ и возможностью оплаты после получения – просто и надежно!
Hello all, here every one is sharing such knowledge, therefore it’s pleasant to read this website, and I used to pay a visit this weblog everyday.
I was extremely pleased to discover this great site.
I wanted to thank you for ones time for this particularly wonderful read!!
I definitely appreciated every bit of it and i also have you bookmarked to check out new information on your
web site.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/shawn-keller
https://ellak.gr/user/mere1990/
https://imageevent.com/urricus1990
https://permacultureglobal.org/users/56498-michael-staebler
https://www.hentai-foundry.com/user/ivan7121985/profile
Elevate Your Beauty Routine with Clean Glam: Natural, Sustainable, and Cruelty-Free
Hey! I’m at work browsing your blog from my new iphone 3gs!
Just wanted to say I love reading your blog and look
forward to all your posts! Carry on the great work!
Thanks to my father who informed me on the topic of this web site, this blog is genuinely amazing.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/348154
https://chyoa.com/user/assaultive1963
http://www.nfomedia.com/profile?uid=rOiWbkI
https://anotepad.com/notes/qf2mcnge
https://www.metal-archives.com/users/viking591966
Привет всем!
Приобретите документ об образовании по выгодной цене с доставкой по РФ и оплатой после получения ваших руками!
https://maps.google.ng/url?q=http://aurus-diploms.com
https://www.google.com.ag/url?sa=t&url=https://anny-diploms.com
https://cse.google.is/url?sa=t&url=https://anny-diploms.com
https://www.google.sr/url?q=http://free-diplommi.com
https://cse.google.co.in/url?q=https://diploms-asx.com
Получите диплом ВУЗа без предварительной оплаты с доставкой по всей России – просто, надежно, выгодно!
hello there and thank you for your info – I’ve definitely picked
up anything new from right here. I did however expertise
some technical points using this website,
as I experienced to reload the website many times previous to I could get it to load
properly. I had been wondering if your web hosting is OK?
Not that I’m complaining, but slow loading instances times will very frequently affect
your placement in google and could damage your high quality score if advertising and marketing with Adwords.
Anyway I’m adding this RSS to my email and could look out for much more of your respective fascinating content.
Make sure you update this again soon.
Very good post. I am facing a few of these issues as well..
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://syrini1961.bandcamp.com/album/wifes-revenge
https://rentry.org/t5oony6e
https://www.metal-archives.com/users/xssdwsa1967
https://www.divephotoguide.com/user/missgr1011960
https://haveagood.holiday/users/348324
Very good blog! Do you have any tips for aspiring writers?
I’m hoping to start my own website soon but I’m a little lost on everything.
Would you propose starting with a free platform like WordPress or go
for a paid option? There are so many choices out there that I’m
completely overwhelmed .. Any suggestions? Appreciate it!
Наша компания предлагает высококачественные услуги Демонтаж / Снос зданий и домов Алматы Мы обеспечиваем надежное и профессиональное оборудование для выполнения различных земляных работ на строительных площадках и других объектах. Наш опытный персонал и гибкие условия аренды делают нас надежным партнером для вашего проекта.
Greetings! I’ve been reading your blog for
a while now and finally got the courage to go ahead and
give you a shout out from Lubbock Texas! Just wanted to tell you
keep up the good job!
Great article. I’m facing many of these issues as well..
Wow, this paragraph is fastidious, my younger sister is analyzing such things, thus
I am going to let know her.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/janice-rubin
https://anotepad.com/notes/j8rhgfkb
https://www.divephotoguide.com/user/ildus1958
https://launchpad.net/~jacksolo19541
https://www.obesityhelp.com/members/mortician1973/about_me/
Привет всем!
Закажите диплом ВУЗа с доставкой по всей России без предоплаты и с гарантией качества!
https://image.google.tk/url?sa=t&source=web&rct=j&url=http://diplomsagroups.com
https://maps.google.ki/url?sa=t&url=https://russian-diplom.com
https://clients1.google.com.qa/url?sa=t&url=http://originality-diplomy.com
https://maps.google.com.bd/url?sa=t&url=https://premialnie-diplomansy.com
http://partnerpage.google.com/url?q=http://diploms-service.com
Приобретите диплом колледжа недорого с доставкой по РФ и гарантией качества без предварительной оплаты – просто и надежно!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/kimwilliams519
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3192585
https://brachyo1952.diary.ru/
https://cannabis.net/user/149752
https://tubeteencam.com/user/keb4ik1954/profile
It’s an remarkable article designed for all the web viewers; they
will get advantage from it I am sure.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/mountbatten1954
https://www.haikudeck.com/presentations/EG2Yh0lf9t
https://onedio.ru/profile/ko-4-ca-197-0
http://www.nfomedia.com/profile?uid=rOiUfgJ
https://www.obesityhelp.com/members/roginald1993/about_me/
Way cool! Some very valid points! I appreciate you penning this
write-up plus the rest of the website is also very good.
Timsothyreemn
Привет всем!
Диплом стал для меня огромной проблемой, ведь я сильно задержался со сроками его сдачи.
Закажите диплом у нас и получите его быстро и надежно с оплатой после получения, доставка в любой регион РФ.
Желаю каждому не двоешных) оценок!
купить диплом в челябинске
купить диплом в сыктывкаре
купить диплом бухгалтера
купить диплом в брянске
купить диплом в клинцах
https://maps.google.com.ag/url?q=https://bisness-diplom.ru/
https://clients1.google.com.fj/url?sa=t&url=https://diploms-vuza.com/
https://www.google.co.vi/url?q=https://ry-diplom.com/
https://clients1.google.com.fj/url?sa=t&url=https://peoplediplom.ru/
https://images.google.ml/url?sa=t&url=https://diploms-vuza.com/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/sometimes1975
https://permacultureglobal.org/users/56712-valerie-fernandez
https://www.quia.com/profiles/dannybo245
https://www.haikudeck.com/presentations/eYshqNhzhl
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190360
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/xt8g36oe
https://tubeteencam.com/user/crhr291968/profile
https://www.obesityhelp.com/members/offerad1952/about_me/
https://launchpad.net/~noyen19931
https://tubeteencam.com/user/oboru1954/profile
This design is incredible! You obviously know how to keep a reader entertained.
Between your wit and your videos, I was almost moved to start
my own blog (well, almost…HaHa!) Fantastic job.
I really loved what you had to say, and more than that,
how you presented it. Too cool!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/nejniy1992
https://www.metal-archives.com/users/romkka1978
http://www.babelcube.com/user/serena-hitzmann-1
https://ellak.gr/user/baltamel1967/
https://rentry.org/ap4a4wnc
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiUdcI
https://rentry.org/63xhid6m
https://rentry.org/nkt6qod2
https://rentry.org/d6k6swmn
https://anotepad.com/notes/ykbgg4xf
Heya i am for the primary time here. I found this board and
I to find It really helpful & it helped me out a lot. I’m hoping to give something back and help others such as you
helped me.
I have been browsing on-line greater than 3 hours nowadays, yet I never
found any attention-grabbing article like yours.
It is beautiful price enough for me. In my view, if all webmasters and bloggers made excellent content material as you did, the web
can be much more useful than ever before.
You actually make it seem so easy with your presentation but I find this matter
to be really something which I think I would never understand.
It seems too complex and extremely broad for me.
I am looking forward for your next post, I will try to get the hang of it!
Приветики!
Купите диплом института или техникума с доставкой по РФ по выгодной цене без предоплаты!
Купите диплом техникума с доставкой по РФ по выгодной цене без предоплаты – просто и надежно!
https://www.google.com.mm/url?q=https://russiany-diplomans.com
http://cse.google.co.zw/url?sa=t&url=https://frees-diplom.com
https://maps.google.no/url?rct=t&sa=t&url=http://diploman-spb24.com
https://maps.google.com.gt/url?q=https://russiany-diplomans.com
https://www.google.md/url?sa=t&url=https://russdiplomag.com
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://ellak.gr/user/firepoint1986/
https://anotepad.com/notes/wa968byn
https://www.haikudeck.com/presentations/rgDSBwWAmM
https://rentry.org/8gu9bssv
https://permacultureglobal.org/users/58244-leah-leonard
What’s up it’s me, I am also visiting this web site on a regular basis, this site is actually good and
the visitors are really sharing nice thoughts.
I’m really enjoying the design and layout of your blog.
It’s a very easy on the eyes which makes it much more pleasant for
me to come here and visit more often. Did you hire out a developer to create
your theme? Excellent work!
Having read this I thought it was very enlightening.
I appreciate you finding the time and effort to put this article together.
I once again find myself spending a lot of time both
reading and leaving comments. But so what, it was still worth it!
What you wrote made a lot of sense. However, consider this,
what if you wrote a catchier title? I am not suggesting your content is
not solid., but what if you added a title that grabbed a person’s attention? I mean Fornix (crus)/ Stria
Terminalis – fxst | Whole Brain Protocol for
Tractography with Empirical MRI is kinda plain. You ought to peek at Yahoo’s home
page and see how they write news headlines to grab people to open the links.
You might add a video or a pic or two to get people
interested about everything’ve written. In my opinion, it could bring your website a little
bit more interesting.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/p4gnyfeg
http://www.babelcube.com/user/gary-carter-1
https://imageevent.com/bassa1991
https://www.quia.com/profiles/roger582b
https://ellak.gr/user/mystikos1981/
This is my first time visit at here and i am actually impressed to read everthing at one place.
Hmm is anyone else experiencing problems with the images on this blog loading?
I’m trying to figure out if its a problem on my end or if it’s the blog.
Any responses would be greatly appreciated.
Hey! This is my first visit to your blog! We are a collection of volunteers and starting a new
initiative in a community in the same niche. Your blog provided us beneficial information to
work on. You have done a extraordinary job!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/56048-brandy-green
https://rentry.org/e4dtewv2
https://sofast1978.diary.ru/
http://www.babelcube.com/user/cristal-rose
https://rentry.org/8sxgqhcg
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~meou19661
http://www.babelcube.com/user/keith-langley
https://www.metal-archives.com/users/lenokhel1982
https://haveagood.holiday/users/348304
https://bhujibk1959.diary.ru/
hello!,I love your writing very a lot! percentage we be
in contact extra approximately your post on AOL?
I require an expert in this area to solve my problem.
May be that is you! Having a look ahead to see you.
You’re so cool! I don’t believe I have read a single thing like that before.
So great to discover another person with a few unique thoughts on this subject matter.
Really.. thanks for starting this up. This web site is something
that’s needed on the internet, someone with a little originality!
Great blog you’ve got here.. It’s hard to find quality writing like yours these days.
I honestly appreciate people like you! Take care!!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3112113.html
https://joca1968.bandcamp.com/album/i-fucked-my-roommates-boyfriend
https://haveagood.holiday/users/348119
https://chyoa.com/user/lineta1966
https://slipak1992.diary.ru/
Hi everyone, it’s my first go to see at this site, and article is really fruitful in favor of
me, keep up posting these types of posts.
No matter if some one searches for his vital thing, therefore he/she desires
to be available that in detail, therefore that thing is maintained over here.
I savor, cause I discovered just what I used to be having a look for.
You’ve ended my four day long hunt! God Bless you man. Have a great day.
Bye
Its like you read my mind! You appear to know a lot about this, such as you wrote the e-book in it or
something. I think that you just could do with
some p.c. to force the message house a bit, but other than that, that is fantastic blog.
A fantastic read. I’ll certainly be back.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/explo1963/about_me/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190607
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190543
https://www.obesityhelp.com/members/kran1959/about_me/
https://iomi1967.bandcamp.com/album/a-welcome-guest-12
After I initially left a comment I seem to have clicked the -Notify me when new comments are added- checkbox
and from now on each time a comment is added I recieve 4 emails with the exact same comment.
Perhaps there is a way you can remove me from that service?
Thanks a lot!
With havin so much content do you ever run into any problems
of plagorism or copyright infringement? My site has a lot of completely unique
content I’ve either created myself or outsourced but it appears a lot of it is popping it up
all over the internet without my permission.
Do you know any techniques to help reduce content from being stolen? I’d really appreciate it.
Сегодня, когда диплом становится началом удачной карьеры в любом направлении, многие пытаются найти максимально простой путь получения образования. Наличие официального документа об образовании сложно переоценить. Ведь именно он открывает дверь перед любым человеком, который стремится начать трудовую деятельность или учиться в любом университете.
Наша компания предлагает оперативно получить этот необходимый документ. Вы имеете возможность приобрести диплом нового или старого образца, что становится выгодным решением для человека, который не смог завершить обучение, утратил документ или хочет исправить плохие оценки. Любой диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам. В итоге вы сможете получить документ, максимально соответствующий оригиналу.
Преимущества подобного подхода заключаются не только в том, что можно оперативно получить диплом. Процесс организовывается комфортно, с нашей поддержкой. Начиная от выбора подходящего образца документа до консультаций по заполнению личной информации и доставки по стране — все находится под абсолютным контролем качественных мастеров.
Для тех, кто ищет максимально быстрый способ получения требуемого документа, наша компания предлагает отличное решение. Заказать диплом – это значит избежать продолжительного процесса обучения и сразу переходить к личным целям, будь то поступление в университет или начало удачной карьеры.
http://alt1.toolbarqueries.google.com.iq/url?q=https://rudiplomista.com
https://image.google.tg/url?q=https://lands-diploma.com
https://image.google.co.ck/url?sa=j&source=web&rct=j&url=https://premialnie-diplomik.com
https://clients1.google.td/url?sa=t&url=https://premialnie-diplomik.com
http://toolbarqueries.google.se/url?sa=t&url=https://lands-diploma.com
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/assadidas1953
https://www.metal-archives.com/users/saviks1987
https://ellak.gr/user/retytyt1957/
https://www.divephotoguide.com/user/feppapu1955
https://cannabis.net/user/149648
It’s actually a nice and useful piece of information. I am satisfied that you simply shared this useful info with us.
Please keep us up to date like this. Thanks for sharing.
Thank you for sharing your info. I truly appreciate your
efforts and I am waiting for your further write ups thank
you once again.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/rycjian-196-6
https://www.obesityhelp.com/members/eagle1953/about_me/
https://tubeteencam.com/user/nekta1970/profile
https://onedio.ru/profile/olfin-199-4
https://www.hentai-foundry.com/user/tanyki1965/profile
When I originally left a comment I seem to have clicked on the -Notify me when new comments
are added- checkbox and now every time a comment is added
I get four emails with the same comment. Perhaps there is a way you are able to remove me from that service?
Appreciate it!
Every weekend i used to visit this web page, because i
want enjoyment, for the reason that this this web site conations in fact nice funny
information too.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/kosy1981
https://www.divephotoguide.com/user/bushca1995
https://imageevent.com/witos1987
https://chyoa.com/user/vbncvc1992
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3192591
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/xalkfak1958
https://chyoa.com/user/drakeua1985
https://anotepad.com/notes/bksf39k7
https://haveagood.holiday/users/346709
https://tubeteencam.com/user/magonia1977/profile
Normally I don’t learn article on blogs, but I would like to say that this write-up very pressured me to take a look at and do it!
Your writing taste has been amazed me. Thanks, very great article.
What’s Taking place i’m new to this, I stumbled upon this I
have found It absolutely useful and it has helped me out loads.
I am hoping to contribute & help different customers
like its aided me. Good job.
Hurrah! At last I got a website from where I be
capable of truly obtain helpful information concerning my study and knowledge.
I do not know whether it’s just me or if perhaps everyone else encountering problems
with your website. It looks like some of the text in your posts are running
off the screen. Can somebody else please comment and let me know if this is happening to them
too? This could be a problem with my browser because I’ve had this
happen before. Many thanks
Today, I went to the beach front with my kids.
I found a sea shell and gave it to my 4 year old daughter and said “You can hear the ocean if you put this to your ear.” She placed the shell to her ear
and screamed. There was a hermit crab inside and it pinched
her ear. She never wants to go back! LoL I know this is entirely off topic but I had to
tell someone!
Hi my friend! I wish to say that this article is amazing, great written and come with almost
all significant infos. I would like to look more posts like this .
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3189089
https://www.haikudeck.com/presentations/NeEU2zFK5j
http://www.babelcube.com/user/diana-davis-1
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3189101
https://www.quia.com/profiles/croom208
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/56645-miguelittle-ton
https://ellak.gr/user/africa1990/
https://allianc1990.diary.ru/
https://www.quia.com/profiles/andreahogan
https://www.metal-archives.com/users/jkeeeeee1987
Everyone loves what you guys tend to be up too. This sort of clever work and exposure!
Keep up the superb works guys I’ve included you guys to blogroll.
my web blog :: 비아그라
It is actually a great and helpful piece of information. I’m happy that you simply shared this helpful information with us.
Please keep us informed like this. Thank you for sharing.
Excellent write-up. I definitely love this site.
Keep it up!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/mimata
https://www.quia.com/profiles/ta558davenport
https://rentry.org/yoo98u87
https://www.hentai-foundry.com/user/hkyytu1973/profile
https://www.haikudeck.com/presentations/GHcUtPCPj4
Have you ever thought about writing an e-book or guest authoring on other sites?
I have a blog centered on the same topics you discuss and would really like to have you share some
stories/information. I know my viewers would value your work.
If you’re even remotely interested, feel free to send me an e mail.
Your style is so unique in comparison to other people I’ve read stuff from.
Many thanks for posting when you’ve got the opportunity, Guess I’ll just book mark this page.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/gtnjcfnq
https://anotepad.com/notes/tqis8ybm
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191598
https://rentry.org/d6k6swmn
https://permacultureglobal.org/users/56497-christopher-groves
This post provides clear idea designed for the new users of blogging, that truly how to do blogging.
I’m not that much of a internet reader to be honest but your sites really
nice, keep it up! I’ll go ahead and bookmark your site to come back down the road.
Cheers
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3111691.html
https://onedio.ru/profile/caries-197-2
https://www.metal-archives.com/users/nigiliya1979
https://haveagood.holiday/users/348319
https://cannabis.net/user/149282
This is really interesting, You’re a very skilled blogger. I have joined your feed and look forward to seeking more of your wonderful post. Also, I have shared your web site in my social networks!
https://www.google.sc/url?q=https://notes.io/wgEfb
Pretty nice post. I just stumbled upon your weblog and wished to say that I have truly enjoyed surfing around your blog
posts. After all I will be subscribing to your rss
feed and I hope you write again soon!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://gruio1985.diary.ru/
https://www.metal-archives.com/users/llcoolj1992
https://imageevent.com/shopeg1977
https://www.obesityhelp.com/members/forestrun1973/about_me/
https://haveagood.holiday/users/347263
My programmer is trying to persuade me to move to
.net from PHP. I have always disliked the idea because of
the costs. But he’s tryiong none the less. I’ve been using WordPress on a
number of websites for about a year and am worried about switching to
another platform. I have heard excellent things about blogengine.net.
Is there a way I can transfer all my wordpress content into it?
Any kind of help would be really appreciated!
Keep on writing, great job!
https://privatebin.net/?31ecb7b14d78a784#Bb4hdBycJtomWuLmdJLyLFeBFUQh2ADzW3kAwfCujPZU
Ahaa, its nice discussion concerning this article here at this web site, I
have read all that, so at this time me also commenting here.
I’m not that much of a internet reader to be honest but your blogs really nice, keep it up!
I’ll go ahead and bookmark your site to come back in the future.
Many thanks
I used to be suggested this web site through my cousin. I’m no longer sure whether this submit is written by
him as no one else recognise such certain about my trouble.
You are incredible! Thank you!
Greetings! This is my first visit to your blog!
We are a group of volunteers and starting a new initiative in a community in the same niche.
Your blog provided us beneficial information to work on. You have done a wonderful job!
I visited several web pages however the audio quality for audio songs current
at this web page is truly wonderful.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/mike490pa
https://www.haikudeck.com/presentations/oQsvmGH39A
https://www.obesityhelp.com/members/styleoleg1965/about_me/
https://www.hentai-foundry.com/user/pe3ak1984/profile
https://chyoa.com/user/xavi1968
Superb, what a web site it is! This weblog provides helpful information to us, keep it up.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/veter621973
https://rentry.org/28nv2yhm
https://www.quia.com/profiles/leahmitchell
https://tubeteencam.com/user/awafmklam1994/profile
https://www.metal-archives.com/users/taupykle1986
I’ve been exploring for a bit for any high-quality articles or blog
posts on this kind of space . Exploring in Yahoo I finally stumbled upon this web
site. Studying this information So i’m satisfied to convey that I’ve an incredibly excellent
uncanny feeling I found out exactly what I needed.
I so much for sure will make sure to don?t disregard this site and give it a look regularly.
Лендинг-пейдж — это одностраничный сайт, предназначенный для рекламы и продажи товаров или услуг, а также для сбора контактных данных потенциальных клиентов. Вот несколько причин, почему лендинг-пейдж важен для бизнеса:
Увеличение узнаваемости компании. Лендинг-пейдж позволяет представить компанию и её продукты или услуги в выгодном свете, что способствует росту узнаваемости бренда.
Повышение продаж. Заказать лендинг можно здесь – 1landingpage.ru Одностраничные сайты позволяют сосредоточиться на конкретных предложениях и акциях, что повышает вероятность совершения покупки.
Оптимизация SEO-показателей. Лендинг-пейдж создаются с учётом ключевых слов и фраз, что улучшает позиции сайта в результатах поиска и привлекает больше целевых посетителей.
Привлечение новой аудитории. Одностраничные сайты могут использоваться для продвижения новых продуктов или услуг, а также для привлечения внимания к определённым кампаниям или акциям.
Расширение клиентской базы. Лендинг-пейдж собирают контактные данные потенциальных клиентов, что позволяет компании поддерживать связь с ними и предлагать дополнительные услуги или товары.
Простота генерации лидов. Лендинг-пейдж предоставляют краткую и понятную информацию о продуктах или услугах, что облегчает процесс принятия решения для потенциальных клиентов.
Сбор персональных данных. Лендинг-пейдж позволяют собирать информацию о потенциальных клиентах, такую как email-адрес, имя и контактные данные, что помогает компании лучше понимать свою аудиторию и предоставлять более персонализированные услуги.
Улучшение поискового трафика. Лендинг-пейдж создаются с учётом определённых поисковых запросов, что позволяет привлекать больше целевых посетителей на сайт.
Эффективное продвижение новой продукции. Лендинг-пейдж можно использовать для продвижения новых товаров или услуг, что позволяет привлечь внимание потенциальных клиентов и стимулировать их к покупке.
Лёгкий процесс принятия решений. Лендинг-пейдж содержат только самую необходимую информацию, что упрощает процесс принятия решения для потенциальных клиентов.
В целом, лендинг-пейдж являются мощным инструментом для продвижения бизнеса, увеличения продаж и привлечения новых клиентов.
Заказать лендинг
I think this is one of the most vital information for me.
And i am glad reading your article. But should remark on few general things, The website style is perfect, the articles is really great : D.
Good job, cheers
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/L8DsAs1eCI
https://www.quia.com/profiles/keritu230
https://rentry.org/crhiz5ou
https://imageevent.com/mire1985
https://r1997.diary.ru/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/vjgu001979
https://manferty1956.bandcamp.com/album/aimees-first-massage
https://www.divephotoguide.com/user/ifablei1994
https://www.hentai-foundry.com/user/speedmen1987/profile
https://bagira221996.diary.ru/
Hi! I’m at work surfing around your blog from my new iphone 3gs!
Just wanted to say I love reading through your blog and look forward
to all your posts! Keep up the excellent work!
I enjoy what you guys are up too. This type of clever work and coverage!
Keep up the amazing works guys I’ve included you guys to my blogroll.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~ronio19511
https://www.hentai-foundry.com/user/kakyshka1956/profile
https://www.divephotoguide.com/user/dimook1982
https://www.metal-archives.com/users/heliotopia1972
https://imageevent.com/redlof1953
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/salikov1957/profile
https://www.hentai-foundry.com/user/mnhgg1981/profile
https://imageevent.com/akimyra1951
https://sensei2001988.diary.ru/
https://www.haikudeck.com/presentations/rj79VEWJ0R
I’ve been surfing on-line more than 3 hours these days, yet I never discovered any interesting article like yours.
It’s pretty worth enough for me. In my opinion, if all website owners and bloggers
made just right content material as you did, the internet will probably be a lot more helpful than ever before.
Here is my web blog; https://youtu.be/mqUGzJOgICM?si=4wZgdrD_L3KsEayV
Sweet blog! I found it while browsing on Yahoo
News. Do you have any suggestions on how to get listed in Yahoo News?
I’ve been trying for a while but I never seem to get
there! Thank you
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://piksa1953.diary.ru/
https://www.quia.com/profiles/amscobey
https://onedio.ru/profile/elizabbet-195-0
https://ellak.gr/user/sielsi1962/
https://tubeteencam.com/user/cska091998/profile
Привет всем!
Приобретите документы об образовании всех ВУЗов России с гарантированной подлинностью и доставкой по РФ без предварительной оплаты – просто, надежно, выгодно!
https://diplomany-asx.ru/
Доброго всем дня!
Наши услуги помогут вам приобрести диплом ВУЗа с доставкой по всей России без предварительной оплаты – быстро и надежно!
http://www.diplomany-asx.ru
Genuinely no matter if someone doesn’t understand then its up to other viewers that they will help, so here it happens.
Right here is the perfect webpage for everyone who would like to find out about this topic. You understand so much its almost hard to argue with you (not that I actually will need to…HaHa). You certainly put a fresh spin on a topic that has been written about for years. Great stuff, just great!
http://www.diploman-russiyans.com/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/drivers1990
https://chyoa.com/user/predator21977
https://onedio.ru/profile/iceburn-195-1
https://launchpad.net/~sabotage19821
https://tubeteencam.com/user/dadangel1969/profile
This is really interesting, You are a very skilled blogger.
I have joined your feed and look forward to seeking more of your wonderful post.
Also, I’ve shared your web site in my social networks!
I have read so many articles on the topic of the blogger lovers but this piece of writing is actually a good article, keep it up.
Hello, its fastidious article concerning media print, we all
be familiar with media is a impressive source of data.
Hi would you mind letting me know which webhost you’re using?
I’ve loaded your blog in 3 completely different web browsers
and I must say this blog loads a lot quicker
then most. Can you recommend a good web hosting provider at a reasonable price?
Thank you, I appreciate it!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://ellak.gr/user/eldor12341966/
https://www.divephotoguide.com/user/pattonlee1953
https://www.hentai-foundry.com/user/mutednewt1962/profile
https://www.hentai-foundry.com/user/georina1976/profile
https://rentry.org/vfpzsryz
Как купить диплом без усилий
купить диплом Гознак http://www.diplom-msk.ru/ .
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/tgt4qnqs
https://launchpad.net/~lasdafgn19731
https://www.hentai-foundry.com/user/locn1981/profile
https://launchpad.net/~gylg19571
https://www.obesityhelp.com/members/drooss1961/about_me/
Как купить диплом легально
купить диплом техникума http://www.diplom-msk.ru .
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/149248
https://permacultureglobal.org/users/57288-kari-ford
https://www.divephotoguide.com/user/loscxns1978
https://tubeteencam.com/user/tirg1995/profile
https://loppi1988.diary.ru/
Woah! I’m really loving the template/theme of this website.
It’s simple, yet effective. A lot of times it’s challenging to get that
“perfect balance” between superb usability
and appearance. I must say you have done a fantastic job with this.
Additionally, the blog loads extremely quick for me on Opera.
Excellent Blog!
Купить диплом с рефератом в подарок
купить диплом http://diplom-msk.ru .
Good day very nice site!! Guy .. Beautiful .. Amazing
.. I’ll bookmark your site and take the feeds additionally?
I’m satisfied to find a lot of useful info right here within the publish, we
need develop more strategies on this regard, thanks for sharing.
. . . . .
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/danterul11999
https://anotepad.com/notes/eidp5w7x
https://launchpad.net/~imbasin19601
http://www.nfomedia.com/profile?uid=rOiWbkI
https://www.hentai-foundry.com/user/vashsa1984/profile
This website was… how do I say it? Relevant!! Finally I’ve found
something that helped me. Kudos!
Quality articles or reviews is the secret to attract the viewers to pay a quick visit the
site, that’s what this website is providing.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/57077-penny-robertson
https://launchpad.net/~mansmack19851
https://launchpad.net/~kannik20019861
https://www.haikudeck.com/presentations/7AFYIq6znD
https://okwave.jp/profile/u3112591.html
each time i used to read smaller articles or reviews which also clear
their motive, and that is also happening with this post which
I am reading at this time.
great put up, very informative. I ponder why the opposite experts of this sector don’t
realize this. You must continue your writing.
I am sure, you’ve a great readers’ base already!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/C2rONCgnJe
https://www.hentai-foundry.com/user/kisto1980/profile
https://www.hentai-foundry.com/user/stsya1980/profile
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3192038
https://chyoa.com/user/quaresma1976
We are a group of volunteers and opening a new scheme in our
community. Your web site provided us with valuable info to work on. You have
done a formidable job and our whole community will be
grateful to you.
I do consider all the ideas you have introduced for your
post. They’re very convincing and can definitely work.
Still, the posts are too quick for newbies. May you please prolong them a bit from subsequent time?
Thanks for the post.
Букмекерская контора 1Вин – новинка в игровом мире, которая стремительно набирает популярность с 2016 года. Игрока привлекает действующий промокод 1winbonusbk. Увлекательные игры от 1win с впечатляющими суммами ставок и невероятными размерами ставок не оставят равнодушным даже самого требовательного игрока. Это произошло благодаря выгодным тарифам и высококачественным финансовым услугам. Например, приветственный бонус от 1winbonusbk составляет до 200%. Ни один букмекер этого не предлагает.
Чтобы получить доступ к бонусам и акциям 1win, вам необходимо пройти верификацию.
Промокод 1вин тогда примените промокод на сайте или в приложении и мгновенно выигрывайте в полевых или в игровых автоматах 1win и получайте до 50 000 рублей. Если вам понадобится помощь, вы всегда можете обратиться в онлайн-поддержку букмекерской конторы 1win.
Зарегистрироваться в 1win можно через акционную страницу или главную страницу сайта. И здесь, и там вы можете использовать промокод 1winbonusbk.
После регистрации код автоматически активируется, вы можете получить до 50 000 на первый депозит, пополнив игровой счет.
Сегодня букмекерская контора предлагает несколько вариантов вознаграждения. Большинство из них активируются после ввода специальных кодов, каждый из которых уникален. Клиент может использовать комбинацию только 1 раз. Продолжительность ограничена, как и количество участников.
Клиенты 1Win могут получать актуальные промокоды в группах букмекерской конторы в социальных сетях, в мессенджерах и у партнеров букмекерской конторы. Поощрения рассчитываются автоматически. Код необходимо активировать перед следующим пополнением счета. Если комбинация не была использована при регистрации, получить подарок от букмекера будет невозможно.
Промокоды 1вин
https://bonus-promokod-bk.ru/promokody-bk/promokod-1win/
Superb, what a website it is! This website provides useful information to us, keep it up.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/57505-ebony-robinson
https://www.obesityhelp.com/members/gewrton1956/about_me/
http://www.babelcube.com/user/johnny-barton
https://onedio.ru/profile/g-13-b1-96-5
https://onedio.ru/profile/screama-198-8
Wonderful, what a blog it is! This webpage presents useful data to us,
keep it up.
Wonderful blog! I found it while surfing around on Yahoo News.
Do you have any suggestions on how to get listed in Yahoo
News? I’ve been trying for a while but I never seem to get there!
Many thanks
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3111999.html
https://www.divephotoguide.com/user/wkbjzwuaa1972
https://www.obesityhelp.com/members/eljoker1974/about_me/
https://www.hentai-foundry.com/user/savelij1983/profile
https://onedio.ru/profile/elizabbet-195-0
Hi colleagues, its fantastic article on the topic of educationand fully explained, keep it up all the time.
I couldn’t refrain from commenting. Perfectly written!
This is a topic that’s near to my heart… Cheers!
Exactly where are your contact details though?
Привет, дорогой читатель!
Купите диплом института или колледжа с гарантированной подлинностью и доставкой по России без предоплаты – удобно, безопасно, выгодно!
Приобретите документ об образовании по выгодной цене с доставкой по РФ и оплатой после получения ваших руками!
https://clients1.google.com.cy/url?sa=t&url=https://gosznac-diplomy.com
https://cse.google.lu/url?q=http://volga-diplomy.com
https://maps.google.com.pg/url?rct=j&sa=t&url=https://rudiplomisty.com
http://www.google.cl/url?q=http://diploman-spb24.com
https://images.google.dk/url?sa=t&url=https://premialnie-diplomansy.com
I believe this is one of the such a lot significant
information for me. And i am glad reading your article.
But want to remark on some normal issues, The web site taste is perfect, the articles is in point of fact excellent : D.
Just right process, cheers
I was more than happy to find this great site. I need to to
thank you for your time for this wonderful read!! I definitely savored
every part of it and i also have you book marked to
check out new things on your website.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/hustla1986
https://www.haikudeck.com/presentations/jIC911HzXy
https://www.hentai-foundry.com/user/lisaxx1981/profile
https://www.metal-archives.com/users/mozgbi41967
https://mrtini1955.bandcamp.com/album/lonely-night-1
Hi there! I know this is somewhat off topic but I was wondering if you knew where
I could locate a captcha plugin for my comment form?
I’m using the same blog platform as yours and I’m having
problems finding one? Thanks a lot!
Oh my goodness! Amazing article dude! Thanks, However I am going through problems
with your RSS. I don’t understand the reason why I can’t join it.
Is there anyone else getting the same RSS
issues? Anybody who knows the answer can you kindly respond?
Thanks!!
Greate pieces. Keep writing such kind of information on your page.
Im really impressed by your blog.
Hi there, You’ve performed a fantastic job. I
will definitely digg it and individually recommend to my friends.
I’m sure they’ll be benefited from this website.
I visited many web pages but the audio feature for audio songs current at this
site is actually fabulous.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/149392
https://chyoa.com/user/cofee1987
https://imageevent.com/riork1988
https://www.divephotoguide.com/user/spammm1994
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190371
k8 かじ の
素敵な視点と洞察に富んだ内容でした。感謝しています。
I like the helpful information you provide in your
articles. I will bookmark your weblog and check again here frequently.
I am quite sure I’ll learn many new stuff right here! Good luck for the next!
It’s very effortless to find out any topic on web as compared to textbooks,
as I found this article at this site.
I all the time used to study paragraph in news papers but now as I am a user
of web therefore from now I am using net for content, thanks to web.
Greetings! Very helpful advice in this particular article! It’s the little
changes which will make the largest changes.
Many thanks for sharing!
I like the valuable information you provide in your articles.
I’ll bookmark your blog and check again here frequently. I am quite sure I will learn plenty
of new stuff right here! Best of luck for the next!
Hello, i think that i saw you visited my website so i came to “return the favor”.I’m attempting to find things to improve my
site!I suppose its ok to use some of your ideas!!
Привет всем!
У нас вы можете заказать документы об образовании всех ВУЗов России с доставкой по РФ и оплатой после получения – просто и удобно!
Закажите диплом ВУЗа по выгодной цене с доставкой в любой регион России без предоплаты и с полной уверенностью в его законности!
https://www.google.cd/url?sa=i&url=http://diploman-spb24.com
https://www.google.com.eg/url?q=https://rudiplomisty.com
https://www.google.com.pa/url?q=https://originality-diplomiki.com
https://cse.google.li/url?q=https://originality-diplomiki.com
https://www.google.hu/url?q=https://lands-diploms.com
I really like what you guys are usually up too. This sort
of clever work and exposure! Keep up the amazing works guys I’ve
included you guys to blogroll.
This piece of writing is in fact a good one it assists new the web visitors, who are wishing in favor of
blogging.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://hjss1964.diary.ru/
https://permacultureglobal.org/users/56718-greg-king
https://www.metal-archives.com/users/daaaaaaaa1979
https://okwave.jp/profile/u3111534.html
https://cannabis.net/user/149812
You really make it appear so easy along with your presentation but I in finding
this matter to be really something which I feel I’d never understand.
It sort of feels too complex and extremely extensive
for me. I’m having a look forward to your subsequent submit, I
will attempt to get the hang of it!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/robert-owens
https://okwave.jp/profile/u3110261.html
https://anotepad.com/notes/kw4b7442
https://www.hentai-foundry.com/user/robingod21976/profile
http://www.babelcube.com/user/campy-rodriguez
Howdy I am so grateful I found your site, I
really found you by mistake, while I was searching on Google for something else, Regardless I am here now and would
just like to say cheers for a incredible post and
a all round interesting blog (I also love the theme/design), I don’t have time to browse it all at the moment but I have bookmarked it and also added your RSS feeds, so when I have time I will be back to read a lot more,
Please do keep up the excellent jo.
I’ve been exploring for a bit for any high quality articles or weblog posts on this sort of house .
Exploring in Yahoo I eventually stumbled upon this website.
Reading this info So i’m glad to exhibit that I have a very just right uncanny feeling I discovered just what I needed.
I so much indubitably will make certain to do not disregard
this web site and give it a glance regularly.
Здравствуйте!
На нашем сайте вы можете заказать диплом ВУЗа недорого с возможностью оплаты после получения и круглосуточной поддержкой!
Получите российский диплом с гарантированной подлинностью и доставкой по всей стране без предварительной оплаты – просто и удобно!
https://images.google.com.np/url?sa=t&url=http://rudiplomisty24.com
https://www.google.com.eg/url?q=https://lands-diploms.com
http://images.google.cg/url?q=http://rudiplomisty24.com
https://cse.google.ki/url?q=https://rudiplomisty.com
https://www.google.iq/url?sa=t&url=https://gosznac-diplomy.com
If some one wishes expert view concerning blogging afterward i suggest him/her to pay a
quick visit this web site, Keep up the good job.
Attractive section of content. I simply stumbled upon your web site and in accession capital to claim that I acquire in fact loved account your weblog posts. Any way I’ll be subscribing in your augment and even I achievement you get admission to persistently quickly.
https://www.diplomanc-russia24.com/
Right away I am going to do my breakfast, after having my breakfast coming yet again to read more
news.
Pretty component of content. I just stumbled upon your site and in accession capital to
say that I get in fact enjoyed account your weblog posts.
Any way I’ll be subscribing for your augment or even I fulfillment you get entry to constantly fast.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3192431
https://www.quia.com/profiles/m524hudson
https://www.hentai-foundry.com/user/nomer111985/profile
https://haveagood.holiday/users/346760
https://imageevent.com/peretta1960
В нашем мире, где диплом – это начало удачной карьеры в любом направлении, многие стараются найти максимально простой путь получения качественного образования. Наличие официального документа об образовании сложно переоценить. Ведь диплом открывает двери перед людьми, желающими начать профессиональную деятельность или продолжить обучение в университете.
В данном контексте мы предлагаем максимально быстро получить этот важный документ. Вы сможете приобрести диплом, что будет выгодным решением для человека, который не смог завершить обучение, утратил документ или хочет исправить плохие оценки. Каждый диплом изготавливается аккуратно, с особым вниманием к мельчайшим деталям, чтобы в итоге получился продукт, полностью соответствующий оригиналу.
Превосходство такого решения состоит не только в том, что вы сможете оперативно получить диплом. Процесс организован комфортно, с профессиональной поддержкой. От выбора подходящего образца документа до консультаций по заполнению персональных данных и доставки по России — все под полным контролем опытных мастеров.
В результате, для тех, кто ищет быстрый и простой способ получения необходимого документа, наша услуга предлагает выгодное решение. Заказать диплом – значит избежать длительного процесса обучения и сразу перейти к достижению личных целей, будь то поступление в ВУЗ или начало карьеры.
http://diplom-net.ru
В нашем обществе, где диплом является началом успешной карьеры в любом направлении, многие пытаются найти максимально быстрый путь получения качественного образования. Необходимость наличия официального документа переоценить невозможно. Ведь диплом открывает двери перед людьми, желающими вступить в сообщество профессионалов или продолжить обучение в высшем учебном заведении.
Предлагаем быстро получить этот необходимый документ. Вы имеете возможность купить диплом, что будет удачным решением для человека, который не смог завершить образование, утратил документ или хочет исправить плохие оценки. диплом изготавливается с особой аккуратностью, вниманием к мельчайшим элементам, чтобы на выходе получился 100% оригинальный документ.
Преимущество данного подхода заключается не только в том, что вы оперативно получите диплом. Весь процесс организовывается удобно, с нашей поддержкой. От выбора необходимого образца до грамотного заполнения личных данных и доставки по стране — все находится под полным контролем качественных мастеров.
Всем, кто ищет максимально быстрый способ получить необходимый документ, наша компания предлагает отличное решение. Купить диплом – значит избежать продолжительного обучения и не теряя времени перейти к личным целям, будь то поступление в ВУЗ или старт карьеры.
https://anaparegion.ru/news/237-dyussh_viktoriya_vstretila_uchastnikov_vsekubansko/
http://alter-energo.ru/topic2673.html?view=previous
http://www.writemob.com/index.php?qa=user&qa_1=enitoku
http://www.artcalendar.ru/783.html
http://www.livecenter.ru/index.php?showtopic=737&pid=1623&mode=threaded&show=&st=0
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3112358.html
https://www.haikudeck.com/presentations/tuvxDhFfSG
https://www.divephotoguide.com/user/xfreimx1968
http://www.babelcube.com/user/sapna-tapia
https://launchpad.net/~alicom19591
I simply couldn’t depart your web site before suggesting that
I extremely loved the usual info a person supply for your guests?
Is going to be back continuously in order to inspect new posts
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiWabC
https://rentry.org/4avcz56n
https://imageevent.com/gaspea1954
https://www.hentai-foundry.com/user/kellie1995/profile
https://rentry.org/qfk2xz5a
You should be a part of a contest for one of the most useful websites on the web.
I’m going to recommend this blog!
Have you ever considered writing an e-book or guest authoring on other sites?
I have a blog based upon on the same subjects you discuss and would love to have you share some stories/information. I know my audience would enjoy your work.
If you are even remotely interested, feel free to send
me an email.
Greate post. Keep posting such kind of information on your blog.
Im really impressed by it.
Hello there, You have done a great job. I will certainly digg it and in my opinion recommend to
my friends. I’m sure they will be benefited from this website.
Simply wish to say your article is as surprising. The clarity on your put up is just spectacular and that i could assume you’re knowledgeable in this subject. Well along with your permission allow me to clutch your RSS feed to stay updated with drawing close post. Thanks 1,000,000 and please carry on the enjoyable work.
https://animedia.xyz/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/rassavara1986/about_me/
https://fjjssad1985.diary.ru/
https://haveagood.holiday/users/343698
https://rentry.org/vvze95ga
https://haveagood.holiday/users/347861
I think that everything posted made a bunch of sense.
But, what about this? what if you wrote a catchier title? I mean, I don’t want to tell
you how to run your blog, however what if you added something that makes people want
more? I mean Fornix (crus)/ Stria Terminalis – fxst | Whole Brain Protocol for Tractography with Empirical MRI
is a little plain. You could look at Yahoo’s front page and see how they create news titles to grab
viewers interested. You might add a related video or a related picture or two to get readers excited about everything’ve got
to say. Just my opinion, it could make your blog a little bit more interesting.
В нашем мире, где диплом – это начало удачной карьеры в любой сфере, многие стараются найти максимально быстрый и простой путь получения образования. Наличие документа об образовании переоценить невозможно. Ведь именно диплом открывает дверь перед всеми, кто желает начать трудовую деятельность или продолжить обучение в высшем учебном заведении.
Мы предлагаем оперативно получить этот важный документ. Вы имеете возможность приобрести диплом, и это будет выгодным решением для всех, кто не смог закончить обучение или утратил документ. диплом изготавливается аккуратно, с максимальным вниманием к мельчайшим элементам, чтобы в итоге получился 100% оригинальный документ.
Преимущество данного подхода состоит не только в том, что можно максимально быстро получить свой диплом. Весь процесс организовывается комфортно, с профессиональной поддержкой. Начав от выбора требуемого образца до правильного заполнения персональных данных и доставки в любой регион страны — все находится под полным контролем наших специалистов.
Всем, кто пытается найти максимально быстрый способ получить необходимый документ, наша услуга предлагает выгодное решение. Купить диплом – значит избежать длительного процесса обучения и сразу перейти к достижению своих целей: к поступлению в университет или к началу удачной карьеры.
http://diplomany.ru
Great website. Lots of useful info here. I am sending it
to a few buddies ans also sharing in delicious. And
of course, thank you to your effort!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/stacyr600
http://www.babelcube.com/user/jolett-juarez
https://kristalls1967.diary.ru/
http://www.nfomedia.com/profile?uid=rOiWZeD
https://rentry.org/rns5mkr8
В нашем мире, где диплом – это начало отличной карьеры в любом направлении, многие ищут максимально простой путь получения образования. Важность наличия официального документа об образовании трудно переоценить. Ведь диплом открывает двери перед каждым человеком, желающим вступить в сообщество квалифицированных специалистов или продолжить обучение в любом университете.
В данном контексте наша компания предлагает быстро получить этот важный документ. Вы сможете заказать диплом нового или старого образца, и это становится удачным решением для всех, кто не смог завершить образование или потерял документ. дипломы производятся аккуратно, с особым вниманием ко всем нюансам. В итоге вы получите документ, 100% соответствующий оригиналу.
Плюсы такого решения состоят не только в том, что можно оперативно получить свой диплом. Весь процесс организовывается комфортно, с профессиональной поддержкой. Начав от выбора подходящего образца до точного заполнения личной информации и доставки в любой регион России — все находится под абсолютным контролем опытных специалистов.
Всем, кто хочет найти оперативный способ получить необходимый документ, наша компания готова предложить отличное решение. Заказать диплом – это значит избежать долгого обучения и сразу переходить к достижению своих целей, будь то поступление в ВУЗ или старт карьеры.
http://www.diplomany.ru
If you desire to grow your experience just keep visiting
this website and be updated with the newest news posted here.
В нашем мире, где диплом – это начало отличной карьеры в любой отрасли, многие пытаются найти максимально быстрый и простой путь получения качественного образования. Факт наличия документа об образовании сложно переоценить. Ведь именно диплом открывает дверь перед людьми, желающими начать профессиональную деятельность или продолжить обучение в высшем учебном заведении.
В данном контексте наша компания предлагает максимально быстро получить этот необходимый документ. Вы можете купить диплом, и это является выгодным решением для человека, который не смог завершить образование, потерял документ или желает исправить свои оценки. Любой диплом изготавливается с особой аккуратностью, вниманием к мельчайшим элементам, чтобы на выходе получился документ, полностью соответствующий оригиналу.
Преимущества данного решения заключаются не только в том, что вы сможете быстро получить диплом. Весь процесс организовывается удобно, с профессиональной поддержкой. От выбора нужного образца до консультаций по заполнению персональной информации и доставки в любое место страны — все находится под абсолютным контролем качественных мастеров.
Таким образом, для тех, кто пытается найти максимально быстрый способ получения требуемого документа, наша компания предлагает выгодное решение. Заказать диплом – это значит избежать продолжительного процесса обучения и не теряя времени перейти к важным целям: к поступлению в ВУЗ или к началу трудовой карьеры.
http://opendelreyoaks.net/doku.php?id=originalitydiplomiki
http://xn--80adt5btj4c.xn--80asehdb/index.php?subaction=userinfo&user=enapawe
http://www.dachamania.ru/photo_gallrey/profile.php?uid=55214
http://munservis.mirniy.ru/index.php?subaction=userinfo&user=ohobeco
https://maxiotzyv.ru/catalog/landsdiploms
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/348227
http://www.babelcube.com/user/chanel-blackwell
https://imageevent.com/flex13911982
https://cannabis.net/user/149727
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3189390
I’m really enjoying the design and layout of your blog. It’s a very easy on the eyes which makes
it much more enjoyable for me to come here and visit more often. Did you hire out
a developer to create your theme? Superb work!
Very rapidly this website will be famous amid all
blogging and site-building visitors, due to it’s nice articles or reviews
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/valentiin1970
https://tubeteencam.com/user/gecnj1959/profile
https://cannabis.net/user/149769
https://okwave.jp/profile/u3112601.html
https://okwave.jp/profile/u3112072.html
Wow, fantastic blog layout! How long have you been blogging for?
you make blogging look easy. The overall look of your site is magnificent, as
well as the content!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/defsoul1973
https://ellak.gr/user/edifir1958/
https://onedio.ru/profile/stolen-86195-4
https://www.quia.com/profiles/jessicalove249
https://tubeteencam.com/user/jeanegray1983/profile
В наше время, когда диплом становится началом успешной карьеры в любой области, многие пытаются найти максимально быстрый и простой путь получения образования. Наличие официального документа переоценить просто невозможно. Ведь диплом открывает дверь перед людьми, желающими вступить в профессиональное сообщество или продолжить обучение в каком-либо университете.
Мы предлагаем оперативно получить этот необходимый документ. Вы имеете возможность заказать диплом, и это будет удачным решением для всех, кто не смог закончить обучение, утратил документ или желает исправить плохие оценки. диплом изготавливается аккуратно, с максимальным вниманием к мельчайшим нюансам. В результате вы получите 100% оригинальный документ.
Превосходство такого подхода состоит не только в том, что можно оперативно получить диплом. Процесс организован комфортно, с нашей поддержкой. От выбора нужного образца документа до консультации по заполнению личных данных и доставки в любое место России — все под полным контролем опытных специалистов.
Для тех, кто хочет найти оперативный способ получить необходимый документ, наша компания может предложить отличное решение. Купить диплом – значит избежать длительного обучения и сразу перейти к важным целям: к поступлению в университет или к началу успешной карьеры.
http://www.pavelgorbenko.ru/2018/02/blog-post.html
https://narodovmnogo-omsk.ru/forum/user/2834/
http://zsaek.tsu.ru/user/2295
https://imperiasport.ru/product/utjazheliteli-2h45-kg-reguliruemye-bstaw20-/reviews/
http://frichinvestigation.org/node/3924
Magnificent beat ! I wish to apprentice while you amend your site, how can i
subscribe for a blog website? The account helped me a acceptable deal.
I had been a little bit acquainted of this your broadcast offered bright clear idea
Hi mates, pleasant article and pleasant arguments commented at this place, I am
actually enjoying by these.
I’m gone to inform my little brother, that he should also pay a visit this blog on regular basis to obtain updated from newest reports.
diplom07.ru
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://baton1974.bandcamp.com/album/my-grandmother-teaches-me-about-life-ch-01
https://www.haikudeck.com/presentations/r5j1YHKYXB
https://www.hentai-foundry.com/user/kapaka1959/profile
https://launchpad.net/~eugenes19671
https://ellak.gr/user/octagonalo1972/
I used to be able to find good advice from your articles.
Way cool! Some extremely valid points! I appreciate you writing
this write-up and also the rest of the website is really good.
Hey! Would you mind if I share your blog with my facebook group?
There’s a lot of folks that I think would really enjoy
your content. Please let me know. Cheers
Just wish to say your article is as amazing. The clarity for your publish is simply spectacular and i could think you’re a professional on this subject.
Well along with your permission let me to grab your RSS feed to keep updated with imminent post.
Thank you a million and please carry on the gratifying
work.
Good day! This post could not be written any better!
Reading this post reminds me of my good old room mate!
He always kept chatting about this. I will forward this page to him.
Pretty sure he will have a good read. Thank you for sharing!
Wow, this post is good, my younger sister is
analyzing these things, therefore I am going to convey her.
Hello, I would like to subscribe for this web site to obtain newest updates, therefore where can i do it
please help out.
Hello, this weekend is fastidious designed for me, as this time i am reading this fantastic educational piece of writing here at
my residence.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/149031
https://www.haikudeck.com/presentations/GHcUtPCPj4
https://www.hentai-foundry.com/user/inslight1998/profile
https://launchpad.net/~svetlui19721
https://haveagood.holiday/users/348454
You can definitely see your expertise within the work you write.
The sector hopes for even more passionate writers such as you who
are not afraid to say how they believe. At all times follow your heart.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/GHcUtPCPj4
https://okwave.jp/profile/u3110762.html
https://www.divephotoguide.com/user/b1osaint1964
https://permacultureglobal.org/users/56768-alexis-riley
https://chyoa.com/user/flaki1959
В нашем обществе, где диплом становится началом отличной карьеры в любом направлении, многие стараются найти максимально простой путь получения образования. Важность наличия официального документа об образовании переоценить невозможно. Ведь именно он открывает дверь перед каждым человеком, желающим начать трудовую деятельность или учиться в университете.
В данном контексте мы предлагаем оперативно получить этот важный документ. Вы сможете заказать диплом старого или нового образца, что становится удачным решением для человека, который не смог закончить обучение или утратил документ. дипломы производятся с особой тщательностью, вниманием к мельчайшим элементам. На выходе вы сможете получить продукт, 100% соответствующий оригиналу.
Преимущества этого подхода состоят не только в том, что можно максимально быстро получить диплом. Весь процесс организовывается удобно, с нашей поддержкой. От выбора нужного образца до консультации по заполнению личных данных и доставки по России — все находится под полным контролем наших мастеров.
В итоге, для всех, кто ищет быстрый и простой способ получения необходимого документа, наша компания предлагает отличное решение. Заказать диплом – это значит избежать длительного обучения и сразу перейти к своим целям, будь то поступление в ВУЗ или начало карьеры.
http://www.aissa.ru/2020/05/blog-post_98.html
http://creatoys.ru/product/pokernyj-nabor-royal-flush-200-/reviews/
http://getamped.yxhi.com/doku.php?id=diplomaasx
http://38.104.231.181/doku.php?id=russianydiplomans
https://gidro2000.com/forum-gidro/user/25528/
It’s appropriate time to make some plans for the long run and
it is time to be happy. I have learn this publish and
if I may just I desire to recommend you some interesting things or advice.
Perhaps you could write next articles referring to this article.
I desire to learn even more issues about it!
Great blog! Is your theme custom made or did you download it from somewhere?
A design like yours with a few simple tweeks would really make my blog shine.
Please let me know where you got your design. Thank you
Thank you, I have recently been searching for info approximately this topic for a while and yours is the best I’ve came upon so
far. However, what in regards to the conclusion? Are you certain concerning the source?
В наше время, когда диплом – это начало успешной карьеры в любом направлении, многие стараются найти максимально быстрый путь получения качественного образования. Наличие документа об образовании переоценить просто невозможно. Ведь диплом открывает дверь перед каждым человеком, который собирается начать трудовую деятельность или продолжить обучение в высшем учебном заведении.
В данном контексте наша компания предлагает быстро получить этот важный документ. Вы имеете возможность купить диплом старого или нового образца, и это становится отличным решением для всех, кто не смог завершить образование или потерял документ. Любой диплом изготавливается с особой аккуратностью, вниманием к мельчайшим нюансам, чтобы в итоге получился 100% оригинальный документ.
Преимущества подобного подхода состоят не только в том, что вы оперативно получите диплом. Процесс организовывается комфортно, с профессиональной поддержкой. Начиная от выбора необходимого образца до консультации по заполнению личной информации и доставки по стране — все находится под абсолютным контролем опытных специалистов.
В результате, для всех, кто ищет быстрый и простой способ получения необходимого документа, наша компания готова предложить выгодное решение. Приобрести диплом – значит избежать продолжительного процесса обучения и не теряя времени перейти к важным целям: к поступлению в ВУЗ или к началу успешной карьеры.
ab-diplom.ru
Thanks in favor of sharing such a nice thought, post is nice, thats why i have
read it completely
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/solii1988
https://rentry.org/hctsq4nn
https://cannabis.net/user/148844
https://imageevent.com/dasha6661974
http://www.babelcube.com/user/michelle-steen
Someone necessarily lend a hand to make severely posts I would state.
This is the very first time I frequented your web page and to
this point? I amazed with the analysis you made to make this particular
post amazing. Magnificent task!
В наше время, когда диплом является началом успешной карьеры в любом направлении, многие ищут максимально простой путь получения качественного образования. Наличие документа об образовании переоценить невозможно. Ведь именно диплом открывает дверь перед всеми, кто собирается вступить в сообщество профессиональных специалистов или продолжить обучение в любом университете.
Предлагаем очень быстро получить этот важный документ. Вы сможете приобрести диплом старого или нового образца, и это становится удачным решением для всех, кто не смог завершить обучение, потерял документ или желает исправить плохие оценки. дипломы изготавливаются аккуратно, с максимальным вниманием к мельчайшим элементам, чтобы в результате получился документ, максимально соответствующий оригиналу.
Преимущество такого решения заключается не только в том, что вы быстро получите диплом. Процесс организовывается комфортно и легко, с профессиональной поддержкой. От выбора необходимого образца до консультаций по заполнению персональной информации и доставки в любой регион страны — все будет находиться под полным контролем наших специалистов.
Таким образом, для тех, кто ищет максимально быстрый способ получить требуемый документ, наша компания готова предложить отличное решение. Приобрести диплом – это значит избежать продолжительного процесса обучения и не теряя времени перейти к своим целям: к поступлению в университет или к началу удачной карьеры.
http://ab-diplom.ru
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/ichiban1964
https://rentry.org/daazsoeh
https://cannabis.net/user/149043
https://spirit681985.diary.ru/
https://www.divephotoguide.com/user/jasmen1991
Good day I am so delighted I found your blog, I really found you by accident, while I was searching on Yahoo for something else, Regardless I am here now and would just like to say cheers for a marvelous post and a all round interesting blog (I also love the theme/design), I don’t have time to go through it all at the moment but I have bookmarked it and also added in your RSS feeds, so when I have time I will be back to read a great deal more, Please do keep up the superb work.
http://www.server-attestats.com
Keep on writing, great job!
http://www.server-attestats.com
Greetings! Very useful advice in this particular article!
It is the little changes which will make the largest changes.
Thanks a lot for sharing!
Superb blog you have here but I was wondering
if you knew of any discussion boards that cover the same topics talked about
in this article? I’d really like to be a part of group where I can get advice from other
knowledgeable individuals that share the same interest. If you have any recommendations, please let me
know. Thanks a lot!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/pelfox1956
https://onedio.ru/profile/shevchouk-196-0
https://imageevent.com/chinaplate1964
https://www.divephotoguide.com/user/curse21960
https://rentry.org/b6syouy5
You actually make it seem so easy with your presentation but I find this topic to be
actually something that I think I would never understand.
It seems too complex and very broad for me. I’m looking forward
for your next post, I’ll try to get the hang of it!
В наше время, когда диплом – это начало отличной карьеры в любой отрасли, многие стараются найти максимально быстрый путь получения образования. Наличие документа об образовании переоценить просто невозможно. Ведь именно он открывает двери перед любым человеком, который стремится вступить в профессиональное сообщество или продолжить обучение в любом университете.
Мы предлагаем очень быстро получить любой необходимый документ. Вы имеете возможность купить диплом, и это будет удачным решением для всех, кто не смог закончить обучение или утратил документ. диплом изготавливается с особой аккуратностью, вниманием к мельчайшим деталям, чтобы на выходе получился продукт, 100% соответствующий оригиналу.
Плюсы подобного подхода заключаются не только в том, что вы сможете оперативно получить свой диплом. Весь процесс организован удобно, с нашей поддержкой. От выбора требуемого образца документа до консультации по заполнению личной информации и доставки по России — все под полным контролем квалифицированных специалистов.
Всем, кто пытается найти быстрый способ получить требуемый документ, наша компания предлагает отличное решение. Заказать диплом – это значит избежать долгого процесса обучения и не теряя времени переходить к достижению своих целей, будь то поступление в университет или начало успешной карьеры.
http://www.premier-climat.ru/product/dimplex-obsidian-serija-hi-tech/reviews/
http://www.postelkr.com/bbs/board.php?bo_table=free&wr_id=409966
http://www.schornfelsen.de/index.php?option=com_easybookreloaded
http://www.lada-xray.net/showthread.php?p=32602#post32602
http://www.livecenter.ru/index.php?showtopic=737&mode=linearplus
Cool blog! Is your theme custom made or did you download it from somewhere?
A design like yours with a few simple tweeks would really make my blog shine.
Please let me know where you got your design. Appreciate it
Добрый день всем!
Получите документ ВУЗа без предоплаты и доставку в руки по России – просто, удобно, безопасно!
Рады предложить Большой выбор документов об образовании всех Вузов России, недорого, с постоплатой, помошь 24/7
https://clients1.google.com/url?sa=t&url=http://diploman-spb24.com
https://cse.google.fi/url?sa=t&url=https://russdiplomag.com
https://clients1.google.cv/url?sa=t&url=https://gosznac-diplomy.com
https://clients1.google.com.bo/url?q=https://diploms-originalniy.com
https://cse.google.com.tr/url?q=https://lands-diploms.com
Keep on writing, great job!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~gastonec19871
https://rentry.org/9fxg4e95
https://cannabis.net/user/148587
https://www.obesityhelp.com/members/shadowpc1986/about_me/
https://rentry.org/uxo5wro5
I enjoy what you guys tend to be up too. Such clever work and
reporting! Keep up the awesome works guys I’ve you guys to blogroll.
I’m impressed, I must say. Seldom do I encounter a blog that’s both educative and interesting, and without a doubt, you’ve hit the nail on the
head. The problem is something which not enough men and
women are speaking intelligently about. I’m very happy I found this in my search for something regarding this.
Howdy very nice blog!! Man .. Excellent .. Amazing ..
I will bookmark your site and take the feeds also? I am glad to search out numerous helpful info right here within the submit, we want develop extra
strategies in this regard, thanks for sharing.
. . . . .
Attractive component to content. I simply stumbled upon your weblog and in accession capital to say that I get actually enjoyed account your blog posts.
Anyway I’ll be subscribing in your augment or even I fulfillment you get right of entry to persistently fast.
I was able to find good info from your articles.
With havin so much written content do you ever run into any issues of plagorism
or copyright violation? My blog has a lot of unique content I’ve either authored myself or outsourced but it appears
a lot of it is popping it up all over the internet without
my permission. Do you know any solutions to help reduce
content from being stolen? I’d definitely appreciate it.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/locn1981/profile
https://cannabis.net/user/149464
https://rentry.org/8sxgqhcg
https://cannabis.net/user/149043
http://www.nfomedia.com/profile?uid=rOiVhkJ
Write more, thats all I have to say. Literally,
it seems as though you relied on the video to make your point.
You obviously know what youre talking about, why throw away your intelligence on just posting videos to your blog when you could be giving us something informative to read?
I do not know if it’s just me or if everybody else encountering issues with your website.
It appears as if some of the written text within your posts are running
off the screen. Can somebody else please provide feedback and
let me know if this is happening to them as well?
This could be a issue with my web browser because I’ve had this happen previously.
Appreciate it
I need to to thank you for this excellent read!!
I certainly loved every bit of it. I have you book-marked to
check out new stuff you post…
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://piksa1953.diary.ru/
https://rentry.org/capghhu4
https://www.quia.com/profiles/juliewe339
https://cannabis.net/user/149392
https://www.metal-archives.com/users/dra1996
Hello there! I know this is somewhat off topic but I
was wondering which blog platform are you using for this
site? I’m getting sick and tired of WordPress because I’ve had issues with hackers and I’m looking at options for another platform.
I would be great if you could point me in the direction of a good platform.
Hey There. I discovered your weblog using msn. This is a very smartly written article.
I will make sure to bookmark it and come back to learn more of your helpful info.
Thank you for the post. I’ll certainly comeback.
I was curious if you ever considered changing the layout of your blog?
Its very well written; I love what youve got to say.
But maybe you could a little more in the way of content so people
could connect with it better. Youve got an awful lot of text for
only having one or two pictures. Maybe you could space it out better?
I’ve been exploring for a little for any high-quality articles or weblog posts in this kind of house .
Exploring in Yahoo I finally stumbled upon this website.
Studying this info So i’m satisfied to show that I have an incredibly
good uncanny feeling I found out exactly what I needed.
I so much indisputably will make sure to don?t overlook
this website and give it a glance on a relentless basis.
An impressive share! I have just forwarded this onto a coworker who has been doing
a little research on this. And he actually bought me dinner because
I discovered it for him… lol. So allow me to reword this….
Thanks for the meal!! But yeah, thanx for spending some time
to talk about this matter here on your web page.
certainly like your website but you need to test the spelling on quite a
few of your posts. Many of them are rife with spelling issues and I
to find it very troublesome to inform the reality nevertheless I will definitely come back again.
В нашем обществе, где диплом является началом отличной карьеры в любом направлении, многие ищут максимально простой путь получения качественного образования. Наличие официального документа переоценить просто невозможно. Ведь именно диплом открывает двери перед каждым человеком, который хочет вступить в сообщество профессиональных специалистов или учиться в высшем учебном заведении.
Мы предлагаем быстро получить любой необходимый документ. Вы сможете приобрести диплом старого или нового образца, что становится выгодным решением для всех, кто не смог закончить обучение или утратил документ. дипломы изготавливаются аккуратно, с максимальным вниманием к мельчайшим элементам. В итоге вы получите продукт, полностью соответствующий оригиналу.
Плюсы данного решения состоят не только в том, что вы оперативно получите свой диплом. Процесс организован комфортно, с профессиональной поддержкой. От выбора необходимого образца до правильного заполнения личной информации и доставки в любой регион России — все находится под полным контролем наших специалистов.
Всем, кто хочет найти быстрый и простой способ получить требуемый документ, наша компания предлагает выгодное решение. Заказать диплом – это значит избежать длительного обучения и не теряя времени переходить к личным целям: к поступлению в университет или к началу успешной карьеры.
http://www.lada-4×4.net/member.php?u=22192
http://realgeek.free.fr/index.php?file=Members&op=detail&autor=esybyqeg
http://buhtarma.ru/users/aheka
http://www.inetpartners.ru/forum/member.php?u=195534
http://blog.dyos.ru/2010/05/on-rocks-bad-romance.html
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/h8f8zwfp
https://haveagood.holiday/users/347921
https://onedio.ru/profile/dambach-195-2
https://tubeteencam.com/user/crhr291968/profile
https://tubeteencam.com/user/smershi1954/profile
I simply could not leave your website prior to suggesting that I really loved the standard info a person supply
to your visitors? Is gonna be back incessantly in order to inspect new posts
Great site. A lot of useful information here.
I am sending it to several buddies ans additionally sharing in delicious.
And naturally, thanks for your effort!
I think the admin of this web page is genuinely
working hard for his web page, since here every data is quality based stuff.
Yesterday, while I was at work, my sister stole my apple ipad and tested to see if it can survive a twenty five foot drop, just so she can be a youtube sensation. My apple ipad is now destroyed and she has 83 views. I know this is entirely off topic but I had to share it with someone!
https://arusak-attestats.ru/
Excellent site you have here.. It’s hard to find quality writing like yours
nowadays. I truly appreciate people like you!
Take care!!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/148305
https://imageevent.com/nickol1966
https://www.divephotoguide.com/user/nerka1977
https://www.haikudeck.com/presentations/6bvzkJafk9
https://www.haikudeck.com/presentations/SVnAD6WRr2
Remarkable! Its genuinely amazing post, I have got much clear idea about from this post.
https://arusak-attestats.ru/
Привет, дорогой читатель!
У нас вы можете заказать диплом Гознака по специальной цене с доставкой в любой регион России без предоплаты – надежно и выгодно!
Наша компания поможет купить диплом Вуза со скидкой и доставкой в любую точку России
https://maps.google.com.au/url?q=https://russdiplomik.com
https://maps.google.com.tr/url?q=https://lands-diploms.com
https://cse.google.com.ly/url?sa=t&url=https://originality-diploman.com
https://cse.google.cat/url?q=https://russdiplomag.com
https://www.google.pn/url?q=https://lands-diploms.com
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://ellak.gr/user/gttyyyy1991/
https://www.hentai-foundry.com/user/hkyytu1973/profile
https://www.divephotoguide.com/user/daemonity1993
https://onedio.ru/profile/strongs-195-7
https://launchpad.net/~polemic19721
Unquestionably believe that which you stated. Your favorite
reason appeared to be on the internet the simplest thing to be aware
of. I say to you, I certainly get annoyed while people think about worries that they just do not know about.
You managed to hit the nail upon the top and also
defined out the whole thing without having side effect , people can take a signal.
Will probably be back to get more. Thanks
Excellent pieces. Keep writing such kind of info on your blog.
Im really impressed by it.
Hello there, You have performed a great job.
I will certainly digg it and for my part suggest to my friends.
I’m confident they’ll be benefited from this website.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/romkka1978
https://imageevent.com/holydemon1998
https://www.haikudeck.com/presentations/7iHILj2wmv
https://www.metal-archives.com/users/ridick1963
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191365
I truly love your website.. Great colors & theme.
Did you create this site yourself? Please reply back
as I’m hoping to create my own personal website and would like to find out where you got this from
or exactly what the theme is called. Cheers!
I am no longer sure where you’re getting your information,
but good topic. I must spend some time studying more or figuring out more.
Thanks for excellent information I used to be on the lookout for this info for
my mission.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/56535-veronica-carter
https://rentry.org/iyfphb7x
http://www.babelcube.com/user/jennifer-smith-3
https://www.obesityhelp.com/members/bernkastl1950/about_me/
https://www.haikudeck.com/presentations/Pe5uWEvJEm
Great article.
Heya just wanted to give you a brief heads up and let you know a few of the images aren’t loading correctly.
I’m not sure why but I think its a linking issue.
I’ve tried it in two different browsers and both show the
same outcome.
This is really interesting, You’re an overly professional blogger.
I have joined your rss feed and sit up for in quest of more of your wonderful post.
Also, I’ve shared your site in my social networks
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://ghost01231990.bandcamp.com/album/oops-i-must-be-dreaming
https://tubeteencam.com/user/drago20161969/profile
https://www.divephotoguide.com/user/b1osaint1964
https://onedio.ru/profile/poluno-4-ka-195-3
https://www.haikudeck.com/presentations/VpQW13SBWK
Nice post. I learn something new and challenging
on sites I stumbleupon everyday. It will always be
useful to read articles from other writers and practice something from other
websites.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/347913
https://launchpad.net/~marzakin19781
https://www.metal-archives.com/users/jkeeeeee1987
https://www.obesityhelp.com/members/lordvolk1963/about_me/
https://imageevent.com/vladssfm1994
I couldn’t refrain from commenting. Very well written!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/Q9V590PAxY
https://anotepad.com/notes/yrfsna5w
https://haveagood.holiday/users/346233
https://anotepad.com/notes/e2py7nbd
https://anotepad.com/notes/tqis8ybm
Very nice post. I just stumbled upon your weblog and wanted to say that I have truly enjoyed surfing
around your blog posts. In any case I’ll be subscribing to your rss feed
and I hope you write again very soon!
Hi! I’m at work surfing around your blog from my new iphone 4!
Just wanted to say I love reading through your blog and look forward to all your posts!
Carry on the outstanding work!
Pretty component to content. I just stumbled upon your web site and in accession capital
to claim that I acquire in fact loved account your weblog posts.
Anyway I’ll be subscribing in your feeds or even I fulfillment you get
right of entry to persistently quickly.
Excellent blog here! Also your site loads up fast!
What web host are you using? Can I get your affiliate link to your host?
I wish my website loaded up as quickly as yours lol
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/gaur5zr9
https://imperka1958.bandcamp.com/album/the-truth-about-nikki-pt6
https://lekci1981.diary.ru/
https://permacultureglobal.org/users/57030-todd-taylor
https://rentry.org/nepgwiuu
I am not sure where you are getting your information, but good topic.
I needs to spend some time learning more or understanding more.
Thanks for great info I was looking for this info for my mission.
I read this piece of writing fully regarding the difference of latest
and preceding technologies, it’s remarkable article.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/bmyy9xea
https://rentry.org/dovdcemx
https://chyoa.com/user/sorcermag1956
https://imageevent.com/leemura1967
https://rentry.org/c76ya2d5
Hi, i feel that i saw you visited my web site so i came to return the prefer?.I’m trying to to find things to enhance my site!I guess
its ok to make use of some of your ideas!!
I know this if off topic but I’m looking into starting my own blog
and was wondering what all is required to get set up?
I’m assuming having a blog like yours would cost a pretty penny?
I’m not very internet smart so I’m not 100% certain. Any tips or
advice would be greatly appreciated. Many thanks
Hello there, There’s no doubt that your blog could be having internet browser compatibility issues.
Whenever I look at your web site in Safari, it looks
fine however, when opening in IE, it’s got some overlapping issues.
I merely wanted to give you a quick heads up!
Apart from that, fantastic blog!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/alinira1951/profile
https://launchpad.net/~lomax7919751
https://imageevent.com/xcvbgf1955
https://okwave.jp/profile/u3112184.html
https://www.obesityhelp.com/members/kopud1966/about_me/
Woah! I’m really loving the template/theme of this site.
It’s simple, yet effective. A lot of times
it’s very hard to get that “perfect balance”
between usability and visual appearance. I must say you’ve done a superb job with this.
In addition, the blog loads very quick for me on Opera. Superb Blog!
This is a topic that is close to my heart… Many thanks!
Exactly where are your contact details though?
Very good info. Lucky me I found your site by chance (stumbleupon).
I’ve saved as a favorite for later!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/atackk1978
https://cannabis.net/user/149113
http://www.nfomedia.com/profile?uid=rOiVcdB
https://rentry.org/hv7e4sqx
https://tubeteencam.com/user/manifiko1986/profile
Hello, i think that i saw you visited my blog thus i came to “return the favor”.I’m trying
to find things to improve my site!I suppose its ok to use a few of
your ideas!!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190484
https://imageevent.com/hfff1982
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3189917
https://rentry.org/7nsgi5xu
https://onedio.ru/profile/vilemmira-198-5
It’s going to be finish of mine day, however before end I am reading this enormous paragraph to improve my experience.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/dGFKWt5GFU
https://www.obesityhelp.com/members/tha1951/about_me/
https://cannabis.net/user/149304
https://www.divephotoguide.com/user/mildewed1969
https://anotepad.com/notes/tmggmfib
I have been surfing on-line more than three
hours lately, yet I by no means discovered any fascinating article like yours.
It’s lovely worth enough for me. Personally, if all web owners and bloggers made good
content material as you did, the web might be
much more useful than ever before.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/drakeua1985
https://okwave.jp/profile/u3110642.html
https://www.quia.com/profiles/amanda513sa
https://grafhell1958.diary.ru/
https://rentry.org/ndysrxtf
Hey there! I just want to offer you a big thumbs up for your excellent information you have got right here on this post.
I’ll be returning to your blog for more soon.
It’s an awesome paragraph in support of all the web people; they will get benefit from
it I am sure.
Hello There. I discovered your weblog the use of msn. That is an extremely
neatly written article. I’ll be sure to bookmark it
and come back to learn extra of your helpful info.
Thank you for the post. I will definitely comeback.
A person necessarily help to make seriously articles I would state.
This is the first time I frequented your website page and thus far?
I surprised with the research you made to make this particular
post amazing. Great process!
Great info. Lucky me I ran across your blog by
accident (stumbleupon). I’ve saved it for later!
Thanks for your personal marvelous posting! I quite enjoyed reading it, you’re a great author.
I will ensure that I bookmark your blog and may come back down the road.
I want to encourage one to continue your great posts, have a nice weekend!
I am curious to find out what blog system you are using?
I’m having some small security issues with
my latest blog and I’d like to find something more
risk-free. Do you have any solutions?
It’s remarkable to visit this website and reading the views of all colleagues regarding this article, while I am also
eager of getting experience.
Whoa! This blog looks exactly like my old one! It’s on a totally different subject
but it has pretty much the same page layout and design. Great choice
of colors!
프라그마틱 플레이의 다채로운 슬롯으로 특별한 순간을 즐겨보세요.
프라그마틱 무료
프라그마틱의 라이브 카지노는 정말 현장감 넘치게 즐길 수 있는데, 여기서 더 많은 정보를 얻을 수 있어 좋아요!
https://www.jaswanthch.com
https://www.12315nw.cn/
https://bidclipz.com/link/
Amazing issues here. I’m very happy to see your post.
Thank you so much and I’m having a look forward to touch you.
Will you please drop me a e-mail?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/149126
https://manferty1956.bandcamp.com/album/aimees-first-massage
https://tubeteencam.com/user/maxim12221979/profile
https://www.metal-archives.com/users/lenora1986
https://www.divephotoguide.com/user/minds1952
With havin so much written content do you ever run into any issues of plagorism or copyright violation? My site has a lot of exclusive content I’ve either written myself or outsourced but
it seems a lot of it is popping it up all over the web without my authorization. Do you know any
methods to help stop content from being ripped off? I’d really
appreciate it.
Wow! After all I got a web site from where I be able to really obtain useful
information concerning my study and knowledge.
Hello, for all time i used to check website posts here early in the morning, as i enjoy to find out more and more.
Magnificent goods from you, man. I’ve understand your stuff previous
to and you are just extremely wonderful. I really like what
you have acquired here, certainly like what you’re stating and the way
in which you say it. You make it entertaining and you still
take care of to keep it wise. I can not wait to read
far more from you. This is actually a tremendous
website.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/dashing1968/profile
https://www.divephotoguide.com/user/oleg2301990
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191184
https://haveagood.holiday/users/348134
https://ellak.gr/user/dev4onk1983/
Right away I am going away to do my breakfast, after having my breakfast coming yet again to read other news.
Good post. I learn something new and challenging on blogs I stumbleupon every day. It will always be interesting to read articles from other authors and practice something from other websites.
http://diplomy-originaly.com
Доброго всем дня!
Наши услуги позволят вам купить диплом ВУЗа с доставкой по России без предоплаты и с полной уверенностью в его подлинности – просто и удобно!
Как можно приобрести диплом Вуза в России без предоплаты на сайте? Мы готовы доставить его в любую точку страны.
diploms-asx.com
What’s up i am kavin, its my first time to commenting anyplace, when i read this post i thought i could also create comment due to this brilliant article.
Hi there, You have done a fantastic job. I’ll definitely digg it and personally recommend to my friends. I’m sure they will be benefited from this website.
http://diplomy-originaly.com
I visited many blogs except the audio feature for audio songs present at this site is really wonderful.
Thank you a lot for sharing this with all folks you actually realize what you’re speaking about! Bookmarked. Kindly additionally consult with my web site =). We could have a hyperlink alternate arrangement between us
http://www.diplomy-originaly.com
Hello, I think your blog might be having browser compatibility issues. When I look at your blog in Opera, it looks fine but when opening in Internet Explorer, it has some overlapping. I just wanted to give you a quick heads up! Other then that, excellent blog!
https://diplomy-originaly.com/
Hey! This is kind of off topic but I need some advice from an established blog. Is it tough to set up your own blog? I’m not very techincal but I can figure things out pretty quick. I’m thinking about creating my own but I’m not sure where to start. Do you have any ideas or suggestions? Thank you
Кнопка чорно-червона, 2 положення, KCD2 (250V, 15A, 6 контактів)
Доброго всем дня!
Компания предлагает купить диплом Вуза России недорого, без предоплаты и с гарантией возврата средств.
https://eonline-diploms.com/
Wow that was odd. I just wrote an incredibly long comment but
after I clicked submit my comment didn’t appear. Grrrr…
well I’m not writing all that over again. Regardless, just wanted to
say great blog!
I do not even know how I ended up here, but I thought this post
was good. I do not know who you are but definitely you’re going to a famous blogger if you aren’t already
😉 Cheers!
Nice weblog here! Also your site loads up very fast!
What web host are you the use of? Can I get
your associate hyperlink in your host? I
desire my website loaded up as fast as yours lol
Hello! This is kind of off topic but I need some guidance from an established blog.
Is it hard to set up your own blog? I’m not very techincal but
I can figure things out pretty fast. I’m thinking about making my own but I’m not
sure where to begin. Do you have any ideas
or suggestions? Thank you
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/her1975/about_me/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3192126
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190299
https://haveagood.holiday/users/347389
https://www.hentai-foundry.com/user/mysato1961/profile
Wonderful beat ! I would like to apprentice while you amend your website, how
can i subscribe for a blog site? The account helped me a acceptable deal.
I had been a little bit acquainted of this your broadcast provided bright clear concept
I’m not sure exactly why but this weblog is loading incredibly slow for me.
Is anyone else having this issue or is it a issue on my end?
I’ll check back later on and see if the problem still
exists.
I blog quite often and I seriously appreciate your content. The article has really peaked my interest. I am going to take a note of your site and keep checking for new information about once per week. I subscribed to your RSS feed as well.
https://chart-studio.plotly.com/~Sklofar.ua
Write more, thats all I have to say. Literally, it seems as though you relied on the video to make your point.
You obviously know what youre talking about, why throw away
your intelligence on just posting videos to your weblog
when you could be giving us something informative to read?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/347961
https://okwave.jp/profile/u3110626.html
https://rentry.org/vfwb4psh
https://rentry.org/a8dkox72
https://www.haikudeck.com/presentations/Fo9xqL0H0A
Fantastic blog you have here but I was curious if you knew of any forums that cover the same topics talked about here? I’d really like to be a part of community where I can get suggestions from other experienced individuals that share the same interest. If you have any suggestions, please let me know. Appreciate it!
http://kvitka-ua.com/linzy-u-fary-dlya-vashogo-avtomobilya
Hello Dear, are you in fact visiting this website on a regular basis, if so afterward you will absolutely obtain pleasant experience.
http://kvitka-ua.com/linzy-u-fary-dlya-vashogo-avtomobilya
Hey there great blog! Does running a blog such as this require a lot of work? I’ve no expertise in computer programming but I had been hoping to start my own blog soon. Anyway, if you have any ideas or techniques for new blog owners please share. I understand this is off topic but I just needed to ask. Many thanks!
http://kvitka-ua.com/linzy-u-fary-dlya-vashogo-avtomobilya
Hi, I think your site might be having browser compatibility issues.
When I look at your website in Ie, it looks
fine but when opening in Internet Explorer,
it has some overlapping. I just wanted to give you a quick
heads up! Other then that, fantastic blog!
It’s very simple to find out any matter on net as compared to books, as I found this article at this web page.
Greetings from Colorado! I’m bored at work so I decided to check out your blog on my iphone during lunch
break. I enjoy the info you present here and can’t
wait to take a look when I get home. I’m shocked at how quick your blog loaded on my phone ..
I’m not even using WIFI, just 3G .. Anyhow, fantastic site!
Good day! I could have sworn I’ve been to this website before but after reading through some of the post I
realized it’s new to me. Anyways, I’m definitely happy I
found it and I’ll be book-marking and checking back often!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190430
https://rentry.org/k67ax72m
https://tubeteencam.com/user/rus181998/profile
https://okwave.jp/profile/u3111047.html
https://www.metal-archives.com/users/saw1962
Hi there, You have done an incredible job. I will definitely digg it and personally recommend to
my friends. I’m sure they will be benefited from this web site.
My coder is trying to convince me to move to .net from PHP.
I have always disliked the idea because of the costs. But he’s tryiong none the less.
I’ve been using Movable-type on a variety of websites for about a year and am
nervous about switching to another platform. I have heard fantastic things about blogengine.net.
Is there a way I can transfer all my wordpress content into it?
Any kind of help would be really appreciated!
I think this is one of the most significant information for me.
And i’m glad reading your article. But want to remark
on few general things, The site style is wonderful, the articles
is really excellent : D. Good job, cheers
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3110161.html
https://www.hentai-foundry.com/user/alandro1960/profile
https://onedio.ru/profile/demonoid-196-9
https://haveagood.holiday/users/348301
http://www.nfomedia.com/profile?uid=rOiWgeH
Добрый день всем!
Наша компания предлагает широкий выбор дипломов Вузов России с доставкой по всей стране и гарантией качества.
Купите диплом ВУЗа с гарантированной доставкой в любой город России без предварительной оплаты и уверенностью в его легальности!
http://www.diploms-asx.com
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/uik3csu9
https://b0brb0mba1978.diary.ru/
https://tubeteencam.com/user/magyar1995/profile
https://www.divephotoguide.com/user/ardiasoz1956
https://www.obesityhelp.com/members/mishelsl1991/about_me/
Thank you, I’ve recently been looking for info about this subject for ages and yours is the greatest I have found out so far.
But, what concerning the conclusion? Are you certain in regards
to the source?
Good article. I absolutely appreciate this website. Thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/MJQEPsnSCL
https://www.metal-archives.com/users/chestizer1950
https://imageevent.com/chuksa1975
https://launchpad.net/~isadron219751
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190777
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/linevv-197-3
https://launchpad.net/~gaper19901
https://www.divephotoguide.com/user/camorezzz1954
https://permacultureglobal.org/users/56842-danielle-hawkins
https://tubeteencam.com/user/mulantir195519891988/profile
I just couldn’t leave your web site before suggesting that I extremely enjoyed the standard info a person provide to your visitors?
Is going to be back steadily to inspect new posts
Hello i am kavin, its my first time to commenting anyplace, when i read this piece of writing
i thought i could also make comment due to this sensible post.
Great beat ! I wish to apprentice while you amend your web site, how can i subscribe for a weblog site?
The account aided me a appropriate deal. I had been tiny bit familiar of this your broadcast provided vibrant transparent
concept
I loved as much as you will receive carried out right here.
The sketch is attractive, your authored subject matter stylish.
nonetheless, you command get bought an nervousness over that you wish be
delivering the following. unwell unquestionably come
more formerly again as exactly the same nearly a
lot often inside case you shield this hike.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~powergirl19601
https://permacultureglobal.org/users/56079-marisela-peck
http://www.nfomedia.com/profile?uid=rOiWacF
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191155
https://permacultureglobal.org/users/56783-mary-montanez
Hello to every , because I am genuinely keen of reading this weblog’s post to be updated on a regular basis. It consists of nice stuff.
https://cutt.ly/LeroC4md
Hey, I think your website might be having browser compatibility issues. When I look at your blog site in Safari, it looks fine but when opening in Internet Explorer, it has some overlapping. I just wanted to give you a quick heads up! Other then that, wonderful blog!
https://cutt.ly/Aero1Iwo
Hey I know this is off topic but I was wondering if you knew of any
widgets I could add to my blog that automatically tweet my newest
twitter updates. I’ve been looking for a plug-in like this
for quite some time and was hoping maybe you would have some experience with something like
this. Please let me know if you run into anything.
I truly enjoy reading your blog and I look forward to your new updates.
Hi, this weekend is good in support of me, as
this time i am reading this enormous informative article here at my residence.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/potap4eg19581952
http://www.nfomedia.com/profile?uid=rOiUfdJ
https://onedio.ru/profile/htgcvglkf-197-2
https://imageevent.com/holydemon1998
https://www.obesityhelp.com/members/makap1969/about_me/
wonderful put up, very informative. I wonder why the opposite specialists of
this sector do not understand this. You should continue your writing.
I’m confident, you’ve a great readers’ base already!
I love your blog.. very nice colors & theme. Did you create this website yourself or did you
hire someone to do it for you? Plz answer back
as I’m looking to create my own blog and would like to find
out where u got this from. thank you
Howdy I am so thrilled I found your webpage, I really found you by accident,
while I was researching on Google for something else, Regardless I am here now and would just like to say cheers for a
incredible post and a all round enjoyable blog (I also love the theme/design), I don’t have time to browse it all at the minute but I have book-marked it and also
added in your RSS feeds, so when I have time I will be back to read a great deal
more, Please do keep up the superb work.
Hey I know this is off topic but I was wondering if you knew of any widgets I
could add to my blog that automatically tweet my
newest twitter updates. I’ve been looking for a plug-in like this for quite
some time and was hoping maybe you would have some experience with something like this.
Please let me know if you run into anything.
I truly enjoy reading your blog and I look forward to your new updates.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/sarawa585
https://oc2c21973.diary.ru/
https://www.haikudeck.com/presentations/ql9wfkUuHG
https://cannabis.net/user/148817
https://chyoa.com/user/tonik991950
After looking into a number of the blog posts on your site, I honestly
appreciate your way of writing a blog. I bookmarked it to my bookmark website
list and will be checking back soon. Take a look at my website too
and let me know your opinion.
Hello all, here every one is sharing these kinds of know-how, thus it’s fastidious to read this web site, and I used to pay a quick visit this webpage all the time.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/qpeu65gm
https://okwave.jp/profile/u3111937.html
https://launchpad.net/~angoletto19801
https://www.haikudeck.com/presentations/szeuAkwum1
https://launchpad.net/~xludoedx19561
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/oleg984er1974
https://tubeteencam.com/user/dang1973/profile
https://chyoa.com/user/lll007lll1966
https://tubeteencam.com/user/jhdswdgsd1994/profile
https://permacultureglobal.org/users/55929-larkeese-cox
There is certainly a great deal to find out about this topic.
I really like all the points you’ve made.
Do you have a spam issue on this website; I also am a blogger, and I was wondering your situation; many of us have created some nice methods and we are
looking to trade methods with other folks, please shoot me an e-mail if interested.
I think this is among the most important information for me.
And i am glad reading your article. But should remark on some general things, The site style
is wonderful, the articles is really excellent : D. Good job,
cheers
my web-site :: date night ideas
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/albatros1983
https://tubeteencam.com/user/vatsurka1975/profile
https://tubeteencam.com/user/samons1958/profile
https://www.metal-archives.com/users/mikiara1962
https://www.quia.com/profiles/patriciayo373
What’s up friends, how is all, and what you wish for
to say concerning this article, in my view its truly remarkable designed
for me.
Great post. I used to be checking constantly this blog and I am
impressed! Very helpful information specifically the closing section :
) I maintain such info much. I used to be seeking this certain info for a long time.
Thanks and best of luck.
I was recommended this website by my cousin. I am not sure whether this post is written by him as no one else know such detailed about my trouble.
You’re amazing! Thanks!
I love what you guys are up too. This type
of clever work and reporting! Keep up the superb
works guys I’ve added you guys to my blogroll.
Simply want to say your article is as astonishing.
The clearness for your publish is simply cool and i could think you’re an expert
on this subject. Fine with your permission let me to grab your
feed to stay updated with forthcoming post. Thank you one million and please continue the
enjoyable work.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/tftt9kqq
https://tubeteencam.com/user/dadangel1969/profile
https://www.divephotoguide.com/user/izunami1999
https://www.divephotoguide.com/user/deldago1989
https://cannabis.net/user/148383
If some one desires expert view regarding blogging and site-building after that i
recommend him/her to visit this webpage, Keep
up the good job.
Ab imo pectore — С полной откровенностью.
My partner and I stumbled over here coming from a different page and thought I may
as well check things out. I like what I see so now i’m following
you. Look forward to going over your web page yet again.
Great article. I am going through some of these issues as well..
I would like to thank you for the efforts you have put in writing this site.
I am hoping to view the same high-grade content by you later on as well.
In truth, your creative writing abilities has motivated me to get my own, personal blog now 😉
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/ol11969
https://permacultureglobal.org/users/56262-bridgette-satya
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3190364
https://rentry.org/crvvvtd9
https://www.quia.com/profiles/tellyru
Pretty! This has been a really wonderful post. Thank you
for supplying this information.
For newest news you have to go to see internet and on world-wide-web I found this web page as a finest web site for most recent
updates.
I like what I see so now i’m following you, I am hoping to view the same high-grade content by you later on as well. In truth, your creative writing abilities has motivated me to get my own, personal blog now.
I visited many sites however the audio quality for audio songs current at this web
site is truly wonderful.
Remarkable! Its truly remarkable article, I have got much clear idea concerning from this piece of writing.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.divephotoguide.com/user/madair1999
https://www.hentai-foundry.com/user/stalkerxx1961/profile
https://launchpad.net/~lasdafgn19731
https://tubeteencam.com/user/magester1989/profile
https://anotepad.com/notes/95c6mc54
Ad usum delphini — Для использования дофином.
When some one searches for his necessary thing, thus he/she wishes to be
available that in detail, so that thing is maintained over
here.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/cirneame
https://permacultureglobal.org/users/57190-heather-gonzalez
https://www.divephotoguide.com/user/medman1951
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191838
https://www.haikudeck.com/presentations/4RxLDiCGSV
You can definitely see your expertise in the article you
write. The world hopes for more passionate writers such as you who are not afraid to mention how they
believe. At all times follow your heart.
These are truly impressive ideas in on the topic of blogging.
You have touched some good points here. Any way keep up wrinting.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/xp38pjyy
https://onedio.ru/profile/diania-196-0
https://cannabis.net/user/149260
https://cannabis.net/user/149995
https://www.divephotoguide.com/user/elvien1982
In fact no matter if someone doesn’t be aware of after that its up to other people that they will help, so here it happens.
I used to be able to find good info from your blog posts.
Everyone loves what you guys tend to be up too.
This kind of clever work and reporting! Keep up the fantastic works guys I’ve incorporated
you guys to my own blogroll.
Because the admin of this web page is working, no hesitation very quickly it will
be well-known, due to its feature contents.
Excellent web site. A lot of helpful info here. I’m sending it to several pals ans also sharing in delicious.
And obviously, thanks on your effort!
Nice blog right here! Also your web site so much up fast!
What host are you the usage of? Can I am getting your associate link to your host?
I desire my website loaded up as fast as yours lol
buysteriodsonline.com
“…”Xiao Xiangxiang의 예쁜 얼굴이 얼어 붙었습니다. “스승님, 아프세요?”
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://hanshi1967.diary.ru/
https://onedio.ru/profile/mmmnmnmn-197-0
https://rentry.org/2khawy8w
https://cannabis.net/user/149278
http://www.babelcube.com/user/wesley-tucker
Definitely believe that which you stated. Your favorite justification appeared
to be on the net the easiest thing to be aware of.
I say to you, I definitely get irked while people consider worries that they plainly do not know about.
You managed to hit the nail upon the top as well as defined
out the whole thing without having side effect , people can take a
signal. Will probably be back to get more. Thanks
Amabile opus — Милое созданье.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/benji-french
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191702
https://permacultureglobal.org/users/56118-michelle-sharp
https://www.obesityhelp.com/members/dimkjee1982/about_me/
https://mudya1968.diary.ru/
Ad disputandum — Для обсуждения.
Hi there, I enjoy reading all of your post.
I wanted to write a little comment to support you.
В нашем обществе, где диплом – это начало успешной карьеры в любой сфере, многие стараются найти максимально простой путь получения образования. Наличие документа об образовании переоценить невозможно. Ведь диплом открывает дверь перед любым человеком, желающим вступить в профессиональное сообщество или продолжить обучение в каком-либо университете.
Мы предлагаем максимально быстро получить любой необходимый документ. Вы сможете купить диплом, и это является удачным решением для человека, который не смог завершить образование, утратил документ или желает исправить свои оценки. диплом изготавливается аккуратно, с особым вниманием ко всем деталям. В результате вы сможете получить документ, максимально соответствующий оригиналу.
Преимущество такого подхода заключается не только в том, что можно оперативно получить свой диплом. Процесс организован удобно, с профессиональной поддержкой. Начиная от выбора нужного образца до консультации по заполнению личных данных и доставки в любое место России — все находится под полным контролем качественных мастеров.
Таким образом, для тех, кто ищет максимально быстрый способ получить необходимый документ, наша услуга предлагает отличное решение. Купить диплом – значит избежать длительного процесса обучения и сразу перейти к важным целям, будь то поступление в университет или старт удачной карьеры.
http://proobzh.ru
Aut disce, aut discede — Или учись, или уходи.
В нашем обществе, где диплом – это начало удачной карьеры в любой сфере, многие пытаются найти максимально быстрый путь получения качественного образования. Важность наличия официального документа сложно переоценить. Ведь именно диплом открывает двери перед каждым человеком, который стремится вступить в сообщество профессионалов или учиться в высшем учебном заведении.
Мы предлагаем максимально быстро получить этот необходимый документ. Вы можете заказать диплом нового или старого образца, и это будет отличным решением для человека, который не смог завершить обучение, утратил документ или хочет исправить свои оценки. диплом изготавливается с особой тщательностью, вниманием к мельчайшим нюансам, чтобы на выходе получился полностью оригинальный документ.
Превосходство такого подхода состоит не только в том, что вы быстро получите диплом. Весь процесс организован удобно и легко, с профессиональной поддержкой. Начиная от выбора нужного образца документа до грамотного заполнения личных данных и доставки в любое место России — все под абсолютным контролем квалифицированных специалистов.
Всем, кто пытается найти максимально быстрый способ получения требуемого документа, наша компания предлагает выгодное решение. Купить диплом – это значит избежать долгого обучения и не теряя времени перейти к достижению собственных целей, будь то поступление в университет или начало карьеры.
http://www.proobzh.ru
В современном мире, где диплом является началом отличной карьеры в любой сфере, многие ищут максимально быстрый путь получения образования. Наличие документа об образовании трудно переоценить. Ведь именно он открывает дверь перед людьми, стремящимися вступить в сообщество квалифицированных специалистов или учиться в любом университете.
В данном контексте мы предлагаем очень быстро получить любой необходимый документ. Вы можете купить диплом старого или нового образца, что будет выгодным решением для человека, который не смог завершить образование, потерял документ или желает исправить плохие оценки. Каждый диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам. В итоге вы сможете получить документ, 100% соответствующий оригиналу.
Преимущества подобного подхода состоят не только в том, что вы сможете быстро получить свой диплом. Процесс организовывается удобно, с профессиональной поддержкой. Начиная от выбора необходимого образца диплома до правильного заполнения личных данных и доставки по России — все будет находиться под абсолютным контролем квалифицированных мастеров.
Для всех, кто ищет быстрый способ получения требуемого документа, наша услуга предлагает выгодное решение. Приобрести диплом – значит избежать длительного обучения и не теряя времени переходить к достижению своих целей, будь то поступление в ВУЗ или начало удачной карьеры.
edu-nt.ru
Aut vincere, aut mori — Или победить, или умереть.
В нашем обществе, где диплом – это начало отличной карьеры в любой области, многие стараются найти максимально быстрый и простой путь получения качественного образования. Факт наличия официального документа об образовании трудно переоценить. Ведь диплом открывает дверь перед любым человеком, который желает начать трудовую деятельность или продолжить обучение в ВУЗе.
Наша компания предлагает очень быстро получить этот важный документ. Вы можете купить диплом нового или старого образца, что становится удачным решением для всех, кто не смог завершить образование или утратил документ. дипломы выпускаются с особой тщательностью, вниманием к мельчайшим нюансам. На выходе вы получите полностью оригинальный документ.
Преимущества подобного подхода заключаются не только в том, что вы сможете быстро получить диплом. Весь процесс организован удобно, с нашей поддержкой. От выбора необходимого образца до консультаций по заполнению персональных данных и доставки в любой регион страны — все под абсолютным контролем качественных специалистов.
Всем, кто ищет быстрый и простой способ получения необходимого документа, наша услуга предлагает выгодное решение. Приобрести диплом – значит избежать долгого обучения и сразу перейти к важным целям, будь то поступление в университет или старт трудовой карьеры.
proobzh.ru
В современном мире, где диплом становится началом успешной карьеры в любом направлении, многие ищут максимально быстрый путь получения образования. Факт наличия официального документа об образовании переоценить попросту невозможно. Ведь именно диплом открывает двери перед всеми, кто хочет вступить в профессиональное сообщество или продолжить обучение в любом ВУЗе.
Наша компания предлагает оперативно получить любой необходимый документ. Вы сможете заказать диплом, что является выгодным решением для всех, кто не смог закончить обучение, утратил документ или желает исправить плохие оценки. Любой диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам. В итоге вы получите документ, 100% соответствующий оригиналу.
Плюсы данного подхода заключаются не только в том, что можно максимально быстро получить диплом. Процесс организовывается комфортно, с нашей поддержкой. Начав от выбора требуемого образца до консультаций по заполнению личных данных и доставки в любой регион страны — все под абсолютным контролем наших специалистов.
Таким образом, для тех, кто пытается найти быстрый и простой способ получить требуемый документ, наша компания предлагает отличное решение. Приобрести диплом – это значит избежать длительного обучения и сразу перейти к достижению собственных целей: к поступлению в ВУЗ или к началу трудовой карьеры.
http://www.dof-edu.ru
В нашем обществе, где диплом – это начало успешной карьеры в любой области, многие пытаются найти максимально быстрый путь получения образования. Наличие официального документа переоценить просто невозможно. Ведь диплом открывает двери перед всеми, кто собирается вступить в сообщество квалифицированных специалистов или учиться в ВУЗе.
Предлагаем максимально быстро получить любой необходимый документ. Вы имеете возможность купить диплом нового или старого образца, и это является отличным решением для всех, кто не смог закончить обучение, утратил документ или желает исправить плохие оценки. Любой диплом изготавливается аккуратно, с особым вниманием ко всем нюансам. На выходе вы сможете получить полностью оригинальный документ.
Преимущество данного подхода заключается не только в том, что вы быстро получите диплом. Весь процесс организован комфортно, с профессиональной поддержкой. Начав от выбора необходимого образца до консультаций по заполнению персональной информации и доставки в любое место России — все под полным контролем наших мастеров.
Для тех, кто ищет оперативный способ получить необходимый документ, наша компания предлагает выгодное решение. Приобрести диплом – значит избежать долгого процесса обучения и не теряя времени переходить к достижению личных целей, будь то поступление в ВУЗ или старт карьеры.
edu-nt.ru
В нашем мире, где диплом – это начало удачной карьеры в любом направлении, многие пытаются найти максимально быстрый и простой путь получения образования. Необходимость наличия официального документа трудно переоценить. Ведь диплом открывает двери перед любым человеком, который собирается вступить в сообщество профессионалов или продолжить обучение в высшем учебном заведении.
В данном контексте наша компания предлагает максимально быстро получить этот необходимый документ. Вы сможете купить диплом, что будет выгодным решением для всех, кто не смог завершить образование, потерял документ или хочет исправить свои оценки. Любой диплом изготавливается аккуратно, с максимальным вниманием к мельчайшим нюансам, чтобы на выходе получился 100% оригинальный документ.
Плюсы этого подхода состоят не только в том, что можно быстро получить диплом. Весь процесс организован удобно, с нашей поддержкой. Начиная от выбора подходящего образца до грамотного заполнения личной информации и доставки по стране — все под абсолютным контролем наших специалистов.
Для тех, кто пытается найти быстрый способ получить необходимый документ, наша компания предлагает выгодное решение. Приобрести диплом – это значит избежать длительного процесса обучения и сразу перейти к достижению собственных целей, будь то поступление в ВУЗ или старт карьеры.
http://www.dof-edu.ru
Aeternum vale — Прости навеки
В нашем обществе, где диплом является началом успешной карьеры в любой сфере, многие ищут максимально простой путь получения качественного образования. Важность наличия официального документа сложно переоценить. Ведь именно диплом открывает двери перед каждым человеком, который желает вступить в сообщество профессионалов или продолжить обучение в университете.
Предлагаем оперативно получить любой необходимый документ. Вы имеете возможность приобрести диплом, и это будет отличным решением для всех, кто не смог закончить обучение или потерял документ. диплом изготавливается аккуратно, с максимальным вниманием ко всем деталям, чтобы в результате получился 100% оригинальный документ.
Плюсы такого подхода заключаются не только в том, что можно оперативно получить диплом. Весь процесс организован просто и легко, с профессиональной поддержкой. От выбора нужного образца документа до консультации по заполнению персональной информации и доставки по России — все под абсолютным контролем квалифицированных специалистов.
Для всех, кто ищет быстрый способ получения требуемого документа, наша компания предлагает отличное решение. Купить диплом – значит избежать длительного обучения и не теряя времени перейти к личным целям, будь то поступление в университет или начало удачной карьеры.
proobzh.ru
I truly love your blog.. Great colors & theme. Did you build this web site yourself?
Please reply back as I’m hoping to create my own personal website and would love to learn where you got this from or exactly
what the theme is named. Cheers!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3112964.html
https://ellak.gr/user/sylker1968/
https://www.metal-archives.com/users/shyster1965
https://cannabis.net/user/149331
http://www.babelcube.com/user/jolett-juarez
When some one searches for his essential thing, so he/she wishes to be
available that in detail, therefore that thing is
maintained over here.
I have fun with, cause I found just what I was taking a
look for. You have ended my 4 day long hunt! God Bless you
man. Have a great day. Bye
В современном мире, где диплом – это начало удачной карьеры в любой отрасли, многие ищут максимально быстрый путь получения качественного образования. Факт наличия документа об образовании трудно переоценить. Ведь диплом открывает двери перед каждым человеком, желающим вступить в профессиональное сообщество или продолжить обучение в ВУЗе.
В данном контексте наша компания предлагает очень быстро получить этот необходимый документ. Вы сможете купить диплом нового или старого образца, и это является выгодным решением для человека, который не смог завершить образование, потерял документ или желает исправить свои оценки. диплом изготавливается аккуратно, с особым вниманием ко всем элементам, чтобы на выходе получился продукт, максимально соответствующий оригиналу.
Плюсы этого подхода состоят не только в том, что вы максимально быстро получите свой диплом. Весь процесс организован комфортно, с нашей поддержкой. От выбора нужного образца документа до консультаций по заполнению персональной информации и доставки по стране — все под полным контролем качественных специалистов.
Всем, кто ищет максимально быстрый способ получения требуемого документа, наша компания готова предложить выгодное решение. Заказать диплом – это значит избежать длительного обучения и сразу перейти к достижению своих целей, будь то поступление в ВУЗ или старт трудовой карьеры.
http://www.proobzh.ru
Confessus pro judicato habetur — прав. Сознавшийся считается осуждённым.
Neat blog! Is your theme custom made or did you download it from somewhere?
A theme like yours with a few simple adjustements would really
make my blog shine. Please let me know where you got your theme.
Appreciate it
В нашем обществе, где диплом – это начало отличной карьеры в любой отрасли, многие ищут максимально простой путь получения качественного образования. Наличие документа об образовании сложно переоценить. Ведь диплом открывает двери перед всеми, кто стремится вступить в профессиональное сообщество или учиться в ВУЗе.
Наша компания предлагает оперативно получить этот важный документ. Вы имеете возможность купить диплом нового или старого образца, что является выгодным решением для человека, который не смог завершить обучение, утратил документ или хочет исправить свои оценки. дипломы производятся с особой аккуратностью, вниманием ко всем элементам. На выходе вы сможете получить полностью оригинальный документ.
Преимущества данного подхода заключаются не только в том, что вы сможете максимально быстро получить диплом. Процесс организован просто и легко, с нашей поддержкой. От выбора подходящего образца до консультации по заполнению личных данных и доставки по стране — все находится под абсолютным контролем квалифицированных специалистов.
В итоге, для всех, кто хочет найти быстрый способ получения требуемого документа, наша услуга предлагает отличное решение. Купить диплом – значит избежать продолжительного процесса обучения и сразу переходить к своим целям: к поступлению в ВУЗ или к началу удачной карьеры.
http://www.edu-nt.ru/
What’s up every one, here every one is sharing these knowledge, therefore
it’s nice to read this webpage, and I used to visit this weblog everyday.
Ad gustum — По вкусу.
В нашем мире, где диплом становится началом успешной карьеры в любой области, многие стараются найти максимально быстрый путь получения качественного образования. Наличие официального документа об образовании переоценить просто невозможно. Ведь именно диплом открывает двери перед всеми, кто стремится вступить в профессиональное сообщество или учиться в любом университете.
В данном контексте мы предлагаем быстро получить этот важный документ. Вы можете купить диплом, что будет выгодным решением для всех, кто не смог завершить образование, потерял документ или хочет исправить свои оценки. Все дипломы изготавливаются аккуратно, с особым вниманием к мельчайшим элементам. На выходе вы получите продукт, максимально соответствующий оригиналу.
Плюсы подобного подхода заключаются не только в том, что вы сможете оперативно получить свой диплом. Весь процесс организован удобно и легко, с профессиональной поддержкой. От выбора нужного образца документа до консультаций по заполнению личных данных и доставки в любое место страны — все находится под полным контролем квалифицированных мастеров.
В итоге, для всех, кто хочет найти быстрый способ получения требуемого документа, наша услуга предлагает отличное решение. Заказать диплом – это значит избежать долгого обучения и сразу переходить к личным целям: к поступлению в университет или к началу трудовой карьеры.
http://www.proobzh.ru
Hello to every one, the contents present at this website are in fact awesome for people knowledge, well, keep up the good work fellows.
http://www.edu-digest.ru
Министерство неджентльменских дел
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiWgjK
https://www.haikudeck.com/presentations/LVEvwlB0NE
https://rentry.org/fc8yv3qf
https://tubeteencam.com/user/rus181998/profile
https://darkdesir1965.diary.ru/
Министерство неджентльменских дел
Good day! Would you mind if I share your blog with my zynga group?
There’s a lot of folks that I think would really appreciate your content.
Please let me know. Thank you
Do you mind if I quote a couple of your posts as long
as I provide credit and sources back to your site?
My blog is in the very same area of interest as yours and my users would truly benefit from some of the information you provide here.
Please let me know if this okay with you. Thanks a lot!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiVbiB
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3189243
https://haveagood.holiday/users/344755
https://koshe4k1971.diary.ru/
https://rentry.org/zobaud2s
Усик – Фьюри: смотреть онлайн-трансляцию церемонии
Усик – Фьюри: смотреть онлайн-трансляцию церемонии
Excellent post. I used to be checking constantly this weblog and I am inspired!
Very useful information particularly the last phase :
) I handle such info much. I used to be looking for this certain info for a long time.
Thanks and good luck.
Александр Усик – Тайсон Фьюри. Смотреть онлайн LIVE
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/c76ya2d5
https://chyoa.com/user/knowdeath1967
https://onedio.ru/profile/morale-195-2
https://www.obesityhelp.com/members/dredffr1987/about_me/
https://www.divephotoguide.com/user/neplach1986
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/ungry1997/profile
https://okwave.jp/profile/u3111521.html
https://launchpad.net/~ilecfg19951
http://www.nfomedia.com/profile?uid=rOiVhgJ
https://chyoa.com/user/bh4o1973
Appreciate it. Ample information.
whoah this weblog is excellent i like reading your posts.
Stay up the great work! You realize, many persons
are hunting round for this information, you could help them
greatly.
Фуриоса: Хроники Безумного Макса
Фуриоса: Хроники Безумного Макса
Woah! I’m really loving the template/theme of this website.
It’s simple, yet effective. A lot of times it’s hard to get that “perfect balance” between user
friendliness and appearance. I must say that you’ve
done a very good job with this. Additionally, the blog
loads extremely fast for me on Opera. Outstanding Blog!
Фуриоса: Хроники Безумного Макса
I am regular reader, how are you everybody? This paragraph posted at this web site is genuinely nice.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/149278
https://rentry.org/fyudozqw
https://okwave.jp/profile/u3112028.html
https://okwave.jp/profile/u3110216.html
https://www.divephotoguide.com/user/issalt1996
I am really inspired together with your writing abilities and also with the layout for your weblog.
Is that this a paid theme or did you modify it
yourself? Anyway stay up the nice high quality writing, it is uncommon to see a
nice blog like this one these days..
Thanks on your marvelous posting! I definitely enjoyed
reading it, you might be a great author. I will make sure to bookmark
your blog and may come back from now on. I want to encourage you continue your great work, have a nice morning!
В наше время, когда диплом становится началом успешной карьеры в любой сфере, многие ищут максимально быстрый и простой путь получения образования. Наличие официального документа об образовании переоценить просто невозможно. Ведь именно он открывает двери перед любым человеком, желающим вступить в сообщество профессиональных специалистов или учиться в университете.
Наша компания предлагает очень быстро получить этот важный документ. Вы можете приобрести диплом, что будет выгодным решением для всех, кто не смог завершить обучение, потерял документ или желает исправить свои оценки. дипломы изготавливаются с особой тщательностью, вниманием ко всем нюансам, чтобы на выходе получился документ, полностью соответствующий оригиналу.
Преимущества такого подхода заключаются не только в том, что можно быстро получить диплом. Процесс организовывается просто и легко, с профессиональной поддержкой. Начиная от выбора необходимого образца до консультации по заполнению личной информации и доставки по России — все под абсолютным контролем наших мастеров.
Для всех, кто хочет найти максимально быстрый способ получить требуемый документ, наша компания готова предложить выгодное решение. Заказать диплом – это значит избежать продолжительного процесса обучения и сразу перейти к достижению личных целей: к поступлению в ВУЗ или к началу удачной карьеры.
http://edu-nt.ru/
Психолог онлайн
Сегодня, когда диплом – это начало удачной карьеры в любом направлении, многие стараются найти максимально быстрый путь получения образования. Наличие официального документа об образовании переоценить просто невозможно. Ведь именно он открывает дверь перед людьми, желающими вступить в профессиональное сообщество или продолжить обучение в каком-либо ВУЗе.
В данном контексте мы предлагаем очень быстро получить любой необходимый документ. Вы сможете приобрести диплом старого или нового образца, и это будет выгодным решением для всех, кто не смог закончить обучение или потерял документ. диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам, чтобы на выходе получился документ, полностью соответствующий оригиналу.
Преимущества этого решения состоят не только в том, что можно оперативно получить свой диплом. Весь процесс организован удобно, с профессиональной поддержкой. От выбора необходимого образца до грамотного заполнения личных данных и доставки в любое место страны — все под полным контролем опытных мастеров.
Всем, кто ищет быстрый и простой способ получить необходимый документ, наша услуга предлагает отличное решение. Приобрести диплом – значит избежать длительного процесса обучения и не теряя времени переходить к достижению собственных целей: к поступлению в ВУЗ или к началу удачной карьеры.
http://www.edu-nt.ru/
В наше время, когда диплом – это начало удачной карьеры в любой области, многие пытаются найти максимально простой путь получения образования. Факт наличия официального документа об образовании переоценить невозможно. Ведь именно он открывает двери перед каждым человеком, который собирается вступить в профессиональное сообщество или продолжить обучение в любом университете.
В данном контексте мы предлагаем быстро получить любой необходимый документ. Вы сможете купить диплом старого или нового образца, что будет удачным решением для всех, кто не смог закончить обучение или потерял документ. Все дипломы изготавливаются с особой тщательностью, вниманием к мельчайшим элементам, чтобы в результате получился документ, максимально соответствующий оригиналу.
Плюсы такого решения заключаются не только в том, что можно оперативно получить диплом. Весь процесс организован комфортно, с нашей поддержкой. От выбора подходящего образца до консультации по заполнению персональных данных и доставки в любое место страны — все под абсолютным контролем квалифицированных мастеров.
Всем, кто ищет быстрый и простой способ получения необходимого документа, наша компания предлагает отличное решение. Заказать диплом – это значит избежать продолжительного обучения и сразу перейти к достижению своих целей: к поступлению в университет или к началу удачной карьеры.
http://edu-nt.ru/
Психотерапия
Психоаналитик
Hello it’s me, I am also visiting this website daily, this site is really nice and the viewers are really sharing good thoughts.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/smk4jfpd
https://www.obesityhelp.com/members/kturie1994/about_me/
https://www.hentai-foundry.com/user/ghhyk1972/profile
https://www.divephotoguide.com/user/izunami1999
https://okwave.jp/profile/u3109554.html
Please let me know if you’re looking for a author for your site.
You have some really good posts and I think I would
be a good asset. If you ever want to take some of the load off, I’d love to write some
material for your blog in exchange for a link back to mine.
Please shoot me an email if interested. Cheers!
Valuable information. Fortunate me I found your website accidentally,
and I am shocked why this accident did not came about
in advance! I bookmarked it.
I am not sure where you’re getting your info, but great topic.
I needs to spend some time learning more or understanding more.
Thanks for excellent info I was looking for this info for my mission.
This site really has all of the information and facts I needed
about this subject and didn’t know who to ask.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/rycjian-196-6
https://anotepad.com/notes/7xt93t32
https://onedio.ru/profile/hypophrenia-195-8
https://www.quia.com/profiles/andreawalls
https://bora19821953.bandcamp.com/album/mothers-day-fantasy
We’re a group of volunteers and starting a new scheme
in our community. Your website offered us with
valuable information to work on. You have done an impressive job and our entire community will be
grateful to you.
Hi there to all, since I am really keen of reading this weblog’s post to be updated daily.
It carries nice material.
Excellent site you have here.. It’s difficult to find good quality writing like yours nowadays.
I truly appreciate people like you! Take care!!
whoah this blog is wonderful i love studying your articles.
Keep up the great work! You understand, a lot of persons are hunting round
for this info, you could help them greatly.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/7xt93t32
https://cannabis.net/user/149406
https://www.metal-archives.com/users/arsenchik1960
https://www.divephotoguide.com/user/uiipo1972
https://www.hentai-foundry.com/user/alex1591999/profile
Because the admin of this web site is working, no question very
soon it will be famous, due to its feature contents.
I have been exploring for a little bit for any high quality articles or weblog posts on this kind of area .
Exploring in Yahoo I ultimately stumbled upon this site. Reading
this info So i am satisfied to convey that I have a very just right uncanny feeling I found
out exactly what I needed. I most unquestionably
will make sure to do not overlook this website and give it a look
regularly.
Amazing! Its in fact remarkable article, I have got much
clear idea regarding from this article.
I’m truly enjoying the design and layout of your blog.
It’s a very easy on the eyes which makes it much more
enjoyable for me to come here and visit more often. Did you hire out a developer to create your theme?
Excellent work!
Good day! This is my first comment here so I just wanted to give a quick shout out and tell you I truly enjoy
reading through your blog posts. Can you suggest any other blogs/websites/forums that
cover the same topics? Thanks a lot!
Hi there, this weekend is fastidious in support of me, for the reason that this moment i am reading this great informative post here at my house.
wonderful post, very informative. I ponder why the opposite specialists of this sector do not understand this.
You should continue your writing. I am confident, you’ve a great readers’ base already!
I every time emailed this webpage post page to all my contacts, as if like to read it afterward my contacts will too.
I don’t even know how I ended up here, but I thought this post was great.
I do not know who you are but definitely you’re going to a famous blogger
if you are not already 😉 Cheers!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/okudoku1954/profile
https://www.hentai-foundry.com/user/inslight1998/profile
https://anotepad.com/notes/gcppgdxf
https://www.divephotoguide.com/user/djgaren1975
https://www.obesityhelp.com/members/shane541956/about_me/
Have you ever considered about adding a little bit more than just your articles?
I mean, what you say is important and everything.
However imagine if you added some great graphics or video clips to
give your posts more, “pop”! Your content is excellent but with images and
video clips, this website could undeniably be one of the best in its niche.
Awesome blog!
Greate post. Keep writing such kind of information on your page.
Im really impressed by your blog.
Hey there, You’ve done a fantastic job. I will definitely digg it and individually suggest to my friends.
I’m confident they’ll be benefited from this web site.
Feel free to surf to my website; great post to read
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~defactori19961
https://jokey1976.bandcamp.com/album/brittany-and-kari-do-truth-or-dare
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3189089
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3191355
https://haveagood.holiday/users/348019
For newest news you have to pay a visit web and on the web I found this website as a best web page for most recent updates.
Hello to every body, it’s my first visit of this weblog;
this webpage carries remarkable and genuinely fine stuff in support of readers.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://batprime1963.diary.ru/
https://rentry.org/8rqwe9mi
https://cannabis.net/user/151076
https://chyoa.com/user/hitasua1982
https://permacultureglobal.org/users/59582-brandon-toney
I’m not that much of a internet reader to be honest but your sites
really nice, keep it up! I’ll go ahead and bookmark your website to come back later on. Many thanks
Right here is the right site for everyone who hopes to find out about
this topic. You know so much its almost tough to argue with you (not that
I really will need to…HaHa). You definitely put
a new spin on a subject that’s been written about for a long time.
Great stuff, just excellent!
Aw, this was an extremely good post. Spending some time and actual effort
to produce a very good article… but what can I
say… I hesitate a whole lot and never manage to get
nearly anything done.
Appreciation to my father who told me on the topic of this weblog, this webpage
is genuinely amazing.
Hi there I am so grateful I found your site, I really found you by mistake, while I was browsing on Yahoo for something else, Nonetheless I am here now and would just like to say thank
you for a fantastic post and a all round entertaining blog (I also love the theme/design),
I don’t have time to read it all at the moment but I have bookmarked it and also added your RSS feeds, so when I have time I will
be back to read much more, Please do keep up the excellent job.
Simply desire to say your article is as surprising.
The clarity in your post is just excellent
and i could assume you are an expert on this subject.
Fine with your permission let me to grab your RSS feed to keep up to date with forthcoming post.
Thanks a million and please keep up the rewarding work.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/chad370le
https://anotepad.com/notes/4eh56wg5
https://www.quia.com/profiles/cynthia489si
https://www.obesityhelp.com/members/dooogerl1981/about_me/
https://www.obesityhelp.com/members/denat1977/about_me/
В нашем обществе, где диплом – это начало успешной карьеры в любой сфере, многие ищут максимально быстрый путь получения образования. Наличие официального документа об образовании сложно переоценить. Ведь диплом открывает двери перед всеми, кто собирается начать трудовую деятельность или продолжить обучение в высшем учебном заведении.
В данном контексте мы предлагаем оперативно получить этот важный документ. Вы можете купить диплом, что будет удачным решением для всех, кто не смог закончить образование, утратил документ или желает исправить плохие оценки. дипломы выпускаются с особой тщательностью, вниманием ко всем элементам. На выходе вы сможете получить 100% оригинальный документ.
Преимущества данного подхода заключаются не только в том, что можно оперативно получить диплом. Процесс организован просто и легко, с нашей поддержкой. Начиная от выбора нужного образца до точного заполнения персональных данных и доставки в любой регион страны — все под полным контролем квалифицированных мастеров.
В итоге, для тех, кто ищет быстрый способ получить требуемый документ, наша компания предлагает выгодное решение. Заказать диплом – значит избежать продолжительного обучения и сразу перейти к достижению собственных целей, будь то поступление в ВУЗ или начало карьеры.
http://6landik-diploms.com/
Hmm is anyone else experiencing problems with the images on this blog loading? I’m trying to figure out if its a problem on my end or if it’s the blog. Any suggestions would be greatly appreciated.
http://idiplomanys.com/
На сегодняшний день, когда диплом является началом отличной карьеры в любой сфере, многие стараются найти максимально быстрый и простой путь получения качественного образования. Наличие документа об образовании переоценить просто невозможно. Ведь именно диплом открывает двери перед всеми, кто собирается вступить в профессиональное сообщество или продолжить обучение в любом университете.
Предлагаем оперативно получить любой необходимый документ. Вы сможете приобрести диплом старого или нового образца, и это является выгодным решением для человека, который не смог закончить образование, потерял документ или хочет исправить плохие оценки. дипломы изготавливаются аккуратно, с максимальным вниманием к мельчайшим нюансам. На выходе вы сможете получить продукт, 100% соответствующий оригиналу.
Преимущества такого решения заключаются не только в том, что можно оперативно получить диплом. Весь процесс организовывается комфортно и легко, с профессиональной поддержкой. Начиная от выбора подходящего образца до консультации по заполнению персональной информации и доставки по России — все под абсолютным контролем опытных специалистов.
Всем, кто ищет быстрый и простой способ получить необходимый документ, наша компания предлагает отличное решение. Приобрести диплом – это значит избежать длительного обучения и сразу перейти к достижению собственных целей, будь то поступление в ВУЗ или начало успешной карьеры.
http://6landik-diploms.com/
Please let me know if you’re looking for a writer for your blog.
You have some really good articles and I think I would be a good asset.
If you ever want to take some of the load off, I’d absolutely love to write
some content for your blog in exchange for a link back to mine.
Please shoot me an e-mail if interested. Regards!
Hello would you mind stating which blog platform you’re using?
I’m going to start my own blog in the near future but I’m having a
difficult time making a decision between BlogEngine/Wordpress/B2evolution and
Drupal. The reason I ask is because your design and style seems different then most blogs and I’m looking for something completely unique.
P.S Sorry for being off-topic but I had to
ask!
geinoutime.com
이 처방 국가는 홀로 두 나라에 불과합니다.
Great post. I will be facing a few of these
issues as well..
I do not even know the way I stopped up right here, but I thought
this post was good. I don’t recognise who you might be however definitely you’re going to a famous blogger in the event you aren’t already.
Cheers!
I know this if off topic but I’m looking into starting my own weblog and was wondering what all
is needed to get set up? I’m assuming having a blog like yours would
cost a pretty penny? I’m not very internet savvy so
I’m not 100% sure. Any recommendations or advice
would be greatly appreciated. Appreciate it
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/2bcsw5sn
https://imageevent.com/confessor1998
https://chyoa.com/user/angbeaver1967
http://www.babelcube.com/user/mike-rattanajatuphorn
https://chyoa.com/user/policepw1964
When I initially commented I clicked the “Notify me when new comments are added”
checkbox and now each time a comment is added I get three e-mails with the same comment.
Is there any way you can remove people from that service?
Bless you!
Thanks for one’s marvelous posting! I quite enjoyed reading it, you’re a great author.I will
make sure to bookmark your blog and definitely will
come back in the foreseeable future. I want to encourage
you to continue your great job, have a nice morning!
Does your blog have a contact page? I’m having trouble locating it but, I’d like to send you an e-mail.
I’ve got some suggestions for your blog you might be interested in hearing.
Either way, great website and I look forward to seeing it develop
over time.
naturally like your web-site but you have to check the spelling on several of your posts.
Several of them are rife with spelling issues and I find it
very bothersome to tell the reality on the other
hand I will certainly come back again.
Hello! I know this is kinda off topic however , I’d figured I’d ask.
Would you be interested in trading links or maybe
guest authoring a blog post or vice-versa? My website discusses
a lot of the same topics as yours and I think we could greatly benefit from each other.
If you’re interested feel free to shoot me an email.
I look forward to hearing from you! Superb blog by the way!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/lisionok1979/profile
https://www.quia.com/profiles/jiwelsh
https://www.hentai-foundry.com/user/alced1988/profile
https://tubeteencam.com/user/grhsr1996/profile
https://www.metal-archives.com/users/sobutilni19881994199619561955
If you are going for finest contents like I
do, only pay a visit this web page every day for the reason that it gives feature contents,
thanks
Do you have a spam problem on this website; I also am a blogger, and I was wondering your situation; many of us have developed some nice practices and we are looking to swap
strategies with other folks, please shoot me an e-mail if interested.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/dem0n1s1991
https://gvjh1974.bandcamp.com/album/doug-stories-1
https://imageevent.com/donskoi1985
https://chyoa.com/user/drag6661967
https://rentry.org/vxykp6gd
Admiring the dedication you put into your blog and detailed information you provide.
It’s awesome to come across a blog every once in a while that isn’t the same out of date rehashed material.
Fantastic read! I’ve saved your site and I’m including your RSS feeds to my Google account.
I’m gone to inform my little brother, that he should also visit this web site
on regular basis to take updated from hottest gossip.
Asking questions are in fact nice thing if you are
not understanding something fully, however this paragraph gives nice understanding
even.
I’m really loving the theme/design of your weblog.
Do you ever run into any browser compatibility problems?
A handful of my blog audience have complained about my website not working correctly
in Explorer but looks great in Opera. Do you have any solutions to help fix this problem?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/eugen851957/profile
https://launchpad.net/~linfow19841
https://www.haikudeck.com/presentations/o0Rqo2TzHW
https://permacultureglobal.org/users/59274-gilberto-barrett
https://www.quia.com/profiles/chleighton
Heya are using WordPress for your site platform? I’m new to the blog world but I’m trying to get started and create my own. Do you
require any html coding expertise to make your own blog?
Any help would be really appreciated!
Incredible points. Sound arguments. Keep up the amazing spirit.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/TfRNwyVrLw
https://tubeteencam.com/user/lulakebab1993/profile
https://okwave.jp/profile/u3117362.html
https://rentry.org/uqco5sd2
https://rentry.org/3282xrgf
Hello to every body, it’s my first visit of this website;
this web site carries amazing and actually good material in support of visitors.
Thanks for the marvelous posting! I truly enjoyed reading it,
you could be a great author.I will always bookmark your blog and will
come back in the foreseeable future. I want to encourage you to definitely continue your great job, have a nice weekend!
Magnificent beat ! I would like to apprentice while
you amend your site, how can i subscribe for a
blog website? The account helped me a acceptable deal. I had been a
little bit acquainted of this your broadcast provided bright
clear concept
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/smokemo1994
https://rentry.org/b7ags6bq
https://www.haikudeck.com/presentations/SMwBg2nEQT
https://www.obesityhelp.com/members/kolibri1958/about_me/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196274
Hi friends, how is everything, and what you desire to say
regarding this post, in my view its truly remarkable in support of me.
Hey there! Do you use Twitter? I’d like to follow you
if that would be okay. I’m undoubtedly enjoying your blog and
look forward to new updates.
Everyone loves what you guys tend to be up too. This sort of
clever work and reporting! Keep up the terrific works guys I’ve included you guys
to my blogroll.
Carpe diem
Very good information. Lucky me I recently found your blog by chance (stumbleupon).
I’ve saved as a favorite for later!
I was very pleased to find this website. I need to to thank you for
your time for this wonderful read!! I definitely loved every little bit of
it and I have you book-marked to look at new things in your web site.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiZceF
https://permacultureglobal.org/users/59804-jason-yorgason
https://rentry.org/sbqfq3po
https://kenchic1995.diary.ru/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195636
I’ll right away snatch your rss as I can not in finding your email subscription link or e-newsletter service.
Do you have any? Kindly let me recognize in order that I
may subscribe. Thanks.
What i do not understood is if truth be told how you’re no longer really a lot more
neatly-preferred than you might be now. You are very intelligent.
You already know therefore considerably when it comes to this matter, made me personally consider it from
numerous various angles. Its like men and women are not fascinated unless it is one thing
to accomplish with Girl gaga! Your own stuffs nice.
At all times maintain it up!
I have been exploring for a little bit for any high quality articles or
weblog posts in this sort of house . Exploring in Yahoo I finally
stumbled upon this site. Studying this info So i’m glad to exhibit that I have an incredibly good uncanny
feeling I came upon exactly what I needed. I most indisputably will
make sure to do not omit this web site and provides it a glance regularly.
I visited several sites but the audio feature for audio songs
existing at this site is really marvelous.
you’re really a good webmaster. The website loading pace is amazing.
It seems that you are doing any unique trick. Also, The contents are masterwork.
you’ve done a magnificent task on this matter!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rrushu1987.bandcamp.com/album/our-story-ch-7-final-chapter
https://anotepad.com/notes/5mkny2cx
https://imageevent.com/lirina1981
https://haveagood.holiday/users/350557
https://www.hentai-foundry.com/user/garuuun1966/profile
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/352183
https://chyoa.com/user/mopito1964
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195897
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196532
https://okwave.jp/profile/u3117502.html
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/diemust1953
https://jimjam1989.diary.ru/
https://haveagood.holiday/users/350076
https://rentry.org/33yxxs3v
https://www.haikudeck.com/presentations/3MyCCGAmvm
When someone writes an post he/she retains the plan of a user in his/her brain that how a user can know it.
Thus that’s why this piece of writing is perfect.
Thanks!
I’m impressed, I have to admit. Rarely do I come across a
blog that’s both educative and engaging, and let me tell you, you’ve
hit the nail on the head. The issue is something which not
enough folks are speaking intelligently about. I am very happy that I stumbled across this
during my hunt for something regarding this.
Everything is very open with a precise description of the challenges.
It was truly informative. Your site is very helpful. Thank you for sharing!
Great information. Lucky me I recently found your blog by chance (stumbleupon).
I’ve book-marked it for later!
Hello, just wanted to say, I liked this blog post. It was inspiring.
Keep on posting!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/mysti1955/profile
https://rentry.org/2uvcno5k
https://okwave.jp/profile/u3115171.html
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195347
https://imageevent.com/diableroo1953
АКТУАЛЬНЫЙ ПРОМОКОД 1x_1791223 НА 2024г
Промокоды 1xbet – это неотъемлемая часть современной жизни. Мы все стремимся сэкономить деньги и получить наибольшую выгоду от своих покупок. Исходя из этого, я хотел бы предложить вам уникальный промокод от компании 1xbet, который поможет вам насладиться азартом и выйти в плюс.
1xbet – это ведущая мировая букмекерская контора, предлагающая широкий спектр спортивных ставок и увлекательный игровой опыт. С недавних пор, компания также предлагает возможность играть в виртуальные онлайн-казино, что делает её еще более привлекательной для азартных игроков.
Итак, промокод от компании 1xbet дает вам возможность получить бонус при регистрации на сайте. Просто введите этот уникальный код при создании своего аккаунта, и вам будет начислен бонусный баланс, который вы можете использовать для различных видов ставок и игр.
Промокоды 1ХБЕТ На что вы можете потратить свой бонусный баланс? Возможностей здесь предостаточно. Вы можете делать ставки на спортивные события – футбол, теннис, хоккей, баскетбол и многое другое. Независимо от того, являетесь ли вы фанатом больших лиг или предпочитаете теннисные матчи мирового уровня, 1xbet предлагает обширный выбор категорий и событий, на которые можно сделать ставку.
Если вы больше склонны к игровым автоматам и карточным играм, то также найдете множество вариантов на сайте 1xbet. Яркие слоты, популярные игры, лотереи и многое другое – все это доступно вам с помощью бонусного баланса, который вы получите с промокодом.
Что еще привлекательно в использовании промокода? Это великолепная возможность испытать свою удачу, не тратя при этом деньги из своего собственного кошелька. Бонусный баланс предоставляет вам льготу играть на деньги конторы без риска потери личных средств. Это прекрасный способ познакомиться с азартным миром и понять, на какие игры и виды ставок вы хотите делать наибольший акцент.
Помимо этого, компания 1xbet предлагает регулярные акции и бонусы для своих постоянных игроков, что делает её еще более привлекательной для всех поклонников азартных игр.
Не упустите возможность получить уникальный промокод от 1xbet и начать свой путь в мир азартных игр с удвоенными шансами на успех! Зарегистрируйтесь прямо сейчас, используя промокод, и откройте для себя увлекательный и захватывающий мир ставок и игр на сайте 1xbet. Вам понравится!
Источник: https://mmocenter.ru/blog/promokod-1xbet/
I could not resist commenting. Very well written!
Its such as you read my mind! You seem to grasp a lot approximately this,
such as you wrote the e book in it or something. I think that you
simply could do with a few p.c. to force the message home a bit, but instead of that,
that is magnificent blog. A fantastic read. I will certainly be back.
I think this is one of the most significant info for me. And i’m glad reading your article.
But wanna remark on some general things, The web site style
is great, the articles is really nice : D. Good
job, cheers
Hey There. I found your weblog using msn. That is a really neatly written article.
I’ll make sure to bookmark it and come back to learn more of your useful
information. Thank you for the post. I will definitely comeback.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/33w8z9nk
https://okwave.jp/profile/u3116192.html
https://rentry.org/vh9ku2qf
https://drong01978.diary.ru/
https://www.hentai-foundry.com/user/gyteur1961/profile
I blog frequently and I genuinely thank you for
your content. This great article has really peaked my interest.
I am going to take a note of your blog and keep checking for new information about
once per week. I opted in for your Feed too.
Heya i am for the primary time here. I came across this board
and I find It truly useful & it helped me out much. I am hoping to provide something again and help others such as you aided me.
Hi, Neat post. There is an issue together with your site in internet explorer, might test
this? IE still is the marketplace chief and a huge section of people will miss
your fantastic writing because of this problem.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/lupita-thomas
https://www.quia.com/profiles/charlesvelasquez
https://rentry.org/4yidbn22
https://ttajia4b1996.diary.ru/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195852
Hi there, just wanted to tell you, I loved this article.
It was helpful. Keep on posting!
I like the helpful information you provide
in your articles. I’ll bookmark your blog and check again here frequently.
I’m quite certain I will learn many new stuff right here!
Good luck for the next!
I was wondering if you ever considered changing the page layout
of your website? Its very well written; I love what youve got to say.
But maybe you could a little more in the way of content so people could connect
with it better. Youve got an awful lot of text for
only having 1 or 2 pictures. Maybe you could space it out better?
Its like you read my mind! You appear to know so much about this, like you wrote the book in it or
something. I think that you could do with some pics to drive the message home a bit,
but other than that, this is excellent blog.
An excellent read. I’ll definitely be back.
animehangover.com
하지만 이 남자가 친숙할수록 팡지판은 걱정이 많다.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiYgdH
https://www.hentai-foundry.com/user/artddle1977/profile
http://www.babelcube.com/user/summer-walker
https://cannabis.net/user/150532
https://imageevent.com/alex9791952
Hi, i read your blog from time to time and i own a similar one and i was just wondering if you get a lot of spam feedback?
If so how do you reduce it, any plugin or anything you can advise?
I get so much lately it’s driving me insane so any help is very
much appreciated.
Plinko 1WIN Игра Plinko на платформе 1Win – увлекательное развлечение, которое притягивает внимание множества игроков со всех уголков мира. Plinko – классическая аркадная игра, в которой игроки пускают шарик сверху на специально размеченное поле, состоящее из рядов крючков и препятствий. Шарик, скатываясь вниз, прыгает от крючка к крючку и попадает в различные ячейки на дне поля, каждая из которых имеет свою выигрышную стоимость.
Однако Plinko на 1Win – это не только развлекательная аркада, но и способ получить реальный выигрыш. В игровом поле присутствуют различные выигрышные ячейки с разными коэффициентами, что позволяет игроку получать крупные деньги при успешном попадании.
Aviator 1WIN Aviator is an exhilarating and immersive game that takes players on a thrilling aerial adventure. Set in a visually stunning world, the game combines the thrill of flying with strategic gameplay elements, making it a captivating experience for both casual gamers and hardcore enthusiasts alike.
One of the key aspects that sets Aviator apart from other games in its genre is its realistic and dynamic flight mechanics. The game utilizes advanced physics to simulate the real-world dynamics of flying, allowing players to feel the wind in their hair and the adrenaline coursing through their veins as they soar through the sky. Whether it’s performing daring stunts or engaging in intense dogfights, the flight experience in Aviator is unparalleled.
In Aviator, players have the opportunity to take on various missions and challenges, each with its own unique objectives and rewards. From delivering important cargo to engaging in aerial combat, each mission provides a different set of challenges that require strategic thinking and precision flying skills. The game also offers a wide range of customizable aircraft, allowing players to personalize their flying experience and optimize their performance in the skies.
Lucky Jet 1WIN Мир изменился. Времена устройства постепенно ушли в прошлое, и сегодня важным является скорость и мобильность. Именно поэтому компания Lucky Jet разработала уникальный продукт – 1win.
1win – это новое поколение реальных и виртуальных развлечений, объединенных в одной платформе. Будь то спортивные ставки, казино, слоты или электронный покер, 1win предлагает всемирно известные игры в одном месте. Пользователи больше не нуждаются в установке отдельных приложений или посещении различных сайтов, все решения доступны сразу на платформе 1win.
Выбирайте из бесконечного списка событий и спортивных матчей и делайте ставки на любимых команд или игроков. Следите за результатами в режиме реального времени и получайте радость от победы. И все это в одно касание на вашем смартфоне или планшете.
Mines 1WIN Mines 1win is a thrilling game that combines the excitement of mining and the thrill of winning. In this immersive virtual world, players have the opportunity to explore an underground mine filled with precious gems and valuable resources.
As you descend deeper into the mine, you’ll encounter various challenges and obstacles that test your skills and strategy. From navigating treacherous pathways to avoiding deadly traps, every step you take is crucial and could lead to glorious riches or unfortunate demise.
The objective of the game is to mine as many gems as possible while carefully managing your resources and avoiding any potential dangers. With each successful mining expedition, you’ll earn valuable points and rewards that can be used to upgrade your equipment and improve your chances of striking it big.
But Mines 1win is not just about mining; it’s also about outsmarting your competitors. As you progress through the game, you’ll encounter rival miners who will stop at nothing to claim the gems for themselves. Engage in intense strategic battles as you compete to secure the most valuable resources and emerge as the ultimate master of the mines.
Aviator 1WIN Авиатор – захватывающая и увлекательная игра, которая позволяет погрузиться в мир авиации и стать настоящим пилотом. Разработанная командой профессиональных разработчиков, игра Aviator предлагает игрокам уникальную возможность испытать себя в роли пилота самолета.
Основное преимущество игры заключается в ее реалистичности. Все аспекты авиации, начиная от управления самолетом до выполнения сложных маневров, воссозданы с большой точностью. Реалистичная физика полета, динамические погодные условия и плавное управление делают игру максимально реалистичной.
В игре представлены разнообразные виды самолетов, от классических истребителей до грузовых и пассажирских лайнеров. Каждый самолет имеет свои характеристики и особенности, что добавляет глубины и разнообразия геймплею.
Speed Cash 1WIN Speed Cash – инновационная платформа онлайн-гемблинга, которая предлагает своим пользователям уникальный опыт игры и высокие шансы на выигрыш. Благодаря передовым технологиям и максимально удобному интерфейсу, 1win становится идеальным выбором как для новичков, так и для опытных игроков.
Сервис предлагает широкий ассортимент популярных игр, включая слоты, рулетку, покер, блэкджек и многие другие. Главное преимущество Speed Cash в том, что он предлагает мгновенное начисление выигрышей на счет игрока, что гарантирует мгновенный доступ к заработанным средствам.
Безопасность и надежность игры – основные принципы, на которых базируется Speed Cash. Платформа использует передовые технологии шифрования данных, чтобы гарантировать конфиденциальность и защиту личной информации пользователей. Кроме того, 1win сотрудничает только с проверенными провайдерами игрового контента, чтобы обеспечить честную и непредвзятую игру.
1WIN Ставки на спорт — увлекательное и захватывающее занятие, которое привлекает множество людей по всему миру. И одной из платформ, где можно оказаться в центре спортивных событий и испытать адреналин от угадывания исходов матчей, является 1win.
1win – это онлайн-букмекерская контора, которая предоставляет возможность ставить на самые популярные и востребованные виды спорта. Будь то футбол, хоккей, баскетбол или теннис, на 1win каждый найдет что-то, что вызовет его интерес.
Ключевым преимуществом 1win является удобство пользования и широкий выбор ставок. Платформа предоставляет возможность сделать как простые, так и комбинированные ставки, в зависимости от предпочтений и стратегии пользователя. Кроме того, 1win предлагает привлекательные коэффициенты, которые повышают шансы на победу и увеличивают возможные выигрыши.
Одним из важных аспектов, на который обращают внимание пользователи 1win, является безопасность и надежность платформы. 1win осуществляет лицензионную деятельность и соблюдает все необходимые стандарты безопасности, чтобы гарантировать честную игру и защиту данных клиентов.
Кроме того, 1win предлагает своим пользователям различные бонусы и акции. Новым клиентам доступен приветственный бонус, который позволяет получить дополнительные средства для ставок. Также на 1win проводятся регулярные акции, которые позволяют участникам получить дополнительные вознаграждения и повысить свои шансы на выигрыш.
Источник https://1wmri.com/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/152207
https://anotepad.com/notes/jeamcttw
https://haveagood.holiday/users/350446
https://okwave.jp/profile/u3116851.html
https://cannabis.net/user/150743
This website was… how do I say it? Relevant!!
Finally I have found something that helped me. Kudos!
Keep this going please, great job!
My brother recommended I may like this blog. He used to be totally right.
This submit actually made my day. You can not consider just how
much time I had spent for this information! Thanks!
Kudos, Loads of advice.
I was recommended this blog by my cousin. I’m not sure whether this post is written by him as nobody else know such detailed about my
problem. You are incredible! Thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/lhicks337
https://gylka1974.diary.ru/
https://imageevent.com/ambience1969
https://permacultureglobal.org/users/60306-michael-schaefer
https://cannabis.net/user/152190
Attractive component to content. I just stumbled upon your web site
and in accession capital to assert that I acquire in fact enjoyed account your blog posts.
Anyway I will be subscribing to your feeds and even I success you access consistently quickly.
you are in point of fact a excellent webmaster. The site loading velocity is incredible.
It kind of feels that you’re doing any unique trick.
Also, The contents are masterpiece. you’ve done a fantastic activity on this topic!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/58885-elizabeth-madsen
https://rentry.org/p87o28wq
https://www.hentai-foundry.com/user/stripka19821973/profile
http://www.nfomedia.com/profile?uid=rOiYahC
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196404
Can I just say what a relief to uncover a person that actually understands what they are talking about on the internet.
You definitely understand how to bring an issue
to light and make it important. A lot more people ought to
read this and understand this side of your story.
It’s surprising you aren’t more popular because you surely possess the gift.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/pethelp-196-7
https://rentry.org/hn9rxmxz
https://tubeteencam.com/user/werwerw196519591986/profile
https://www.hentai-foundry.com/user/kilka301978/profile
https://ellak.gr/user/pomaaaaaa1967/
I was wondering if you ever considered changing the layout of your blog?
Its very well written; I love what youve
got to say. But maybe you could a little more in the way of content so people could connect with it better.
Youve got an awful lot of text for only having one
or two pictures. Maybe you could space it out better?
I know this site provides quality depending articles and extra material,
is there any other web site which presents these kinds of stuff in quality?
We are a group of volunteers and opening a new scheme in our community.
Your site offered us with valuable information to work on. You have done an impressive job and our whole community will be
thankful to you.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151984
https://haveagood.holiday/users/351280
https://www.haikudeck.com/presentations/89DixF7dIH
https://www.hentai-foundry.com/user/coby20101976/profile
https://permacultureglobal.org/users/60056-keith-hoffman
Hello! I know this is kind of off topic but I was wondering if you
knew where I could locate a captcha plugin for my comment form?
I’m using the same blog platform as yours and I’m having trouble finding one?
Thanks a lot!
This is really interesting, You’re a very skilled blogger.
I’ve joined your feed and look forward to seeking more of your great post.
Also, I have shared your site in my social networks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/toewsss1982/profile
https://tubeteencam.com/user/bibossik1995/profile
https://www.hentai-foundry.com/user/oothdbgf1956/profile
https://www.hentai-foundry.com/user/sl1958/profile
https://rentry.org/yikrry5g
Промокоды играют важную роль в мире онлайн-шопинга и развлечений, и 1win не является исключением. 1win – это платформа для ставок, казино и онлайн-игр, которая предлагает ряд уникальных возможностей для своих пользователей. Одной из этих возможностей являются промокоды, которые предоставляются пользователям для получения различных преимуществ и бонусов.
Промокоды 1Win представляют собой уникальные комбинации символов, которые могут быть введены на платформе для активации различных акций и бонусов. Они позволяют пользователям получить дополнительные средства на свой игровой счет, повысить свой уровень лояльности или получить доступ к эксклюзивным предложениям и мероприятиям.
Одним из популярных промокодов, предоставляемых 1win, является “m1WIN2024”. Вводя этот промокод при регистрации, новые пользователи получают бонус на свой первый депозит. Это отличная возможность начать свой опыт на платформе с дополнительными средствами и увеличить свои шансы на успех.
Кроме того, 1win предоставляет регулярные промокоды для своих зарегистрированных пользователей. Такие промокоды могут приходить через электронную почту или быть опубликованы на официальных каналах 1win в социальных сетях. Они предлагают возможность получить дополнительные бонусы при пополнении счета, участии в определенных играх или просто за активность на платформе.
Пользуясь промокодами 1win, пользователи могут значительно обогатить свой игровой опыт и получить дополнительные возможности для выигрыша. Важно помнить, что каждый промокод имеет свои условия использования, поэтому рекомендуется внимательно ознакомиться с правилами и требованиями, прежде чем использовать какой-либо промокод.
В целом, использование промокодов 1win – это удобный и простой способ получить дополнительные бонусы и преимущества на платформе. Независимо от того, вы новый пользователь или постоянный клиент, промокоды могут стать отличной возможностью для улучшения вашего опыта игры. Убедитесь в том, что вы следите за обновлениями и акциями, чтобы не упустить свою возможность воспользоваться промокодами 1win и повысить свои шансы на выигрыш.
Источник: https://mmocenter.ru/blog/promokod-1win-promokody-1vin-pri-registracii-na-segodnya/
I’ve been exploring for a little for any high quality articles or blog posts
in this kind of house . Exploring in Yahoo I ultimately stumbled upon this site.
Studying this info So i am happy to express
that I have an incredibly excellent uncanny feeling I found out just what I needed.
I such a lot without a doubt will make certain to do
not disregard this website and give it a look regularly.
Very descriptive post, I liked that bit. Will there be a part 2?
You ought to take part in a contest for one of the finest
blogs online. I’m going to highly recommend this web site!
Woah! I’m really enjoying the template/theme of this blog.
It’s simple, yet effective. A lot of times it’s tough to get that
“perfect balance” between usability and visual appeal.
I must say you’ve done a awesome job with this.
Additionally, the blog loads super quick for me on Firefox.
Exceptional Blog!
ilogidis.com
많은 사람들이 은밀하게 유포하는 뉴스에 따르면 이들은 개인사업자일 가능성이 매우 높다.
Neat blog! Is your theme custom made or did you download it from somewhere?
A theme like yours with a few simple adjustements would really make my blog jump out.
Please let me know where you got your theme.
Thanks
This is very fascinating, You’re a very skilled blogger.
I’ve joined your feed and stay up for seeking extra of your magnificent post.
Also, I have shared your web site in my social networks
Thank you for every other informative blog. Where else could I am getting that kind of information written in such an ideal approach?
I have a venture that I am just now running on, and I have
been on the glance out for such info.
Thanks for sharing your thoughts about Ceritoto Login. Regards
What a material of un-ambiguity and preserveness of precious experience regarding
unpredicted feelings.
Wonderful blog! Do you have any helpful hints for aspiring writers?
I’m hoping to start my own website soon but I’m a little lost on everything.
Would you suggest starting with a free platform like WordPress or go for
a paid option? There are so many choices out there that I’m totally
overwhelmed .. Any suggestions? Thank you!
Hello! I just would like to give you a huge thumbs up for your excellent info you
have here on this post. I’ll be coming back to your site
for more soon.
Thanks designed for sharing such a fastidious thinking,
post is pleasant, thats why i have read it completely
This is a topic which is close to my heart… Best wishes!
Where are your contact details though?
Actually when someone doesn’t know then its up to other viewers that they
will assist, so here it takes place.
Greetings! I’ve been reading your weblog for a long time now
and finally got the bravery to go ahead and give you a shout out
from Humble Tx! Just wanted to tell you keep up the good work!
Greetings from Florida! I’m bored to death at work so I decided
to check out your website on my iphone during lunch break. I really like
the knowledge you present here and can’t wait to take a look
when I get home. I’m surprised at how quick your blog loaded
on my mobile .. I’m not even using WIFI, just 3G ..
Anyhow, amazing site!
I’m really enjoying the design and layout of your website.
It’s a very easy on the eyes which makes it much more enjoyable for me to
come here and visit more often. Did you hire out a designer to
create your theme? Exceptional work!
It is appropriate time to make a few plans for the longer term and it’s time to be
happy. I’ve learn this put up and if I may
just I want to counsel you some attention-grabbing things or tips.
Maybe you could write subsequent articles regarding this article.
I want to learn even more issues about it!
Hello there! I could have sworn I’ve been to this website before but after going through
some of the posts I realized it’s new to me. Nonetheless,
I’m certainly delighted I came across it and I’ll be bookmarking it and checking back often!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/aisonne1981/about_me/
https://mrbz1k1974.diary.ru/
https://jom1998.bandcamp.com/album/in-need-of-it
https://rentry.org/qopzmris
https://onedio.ru/profile/varchik-199-5
Simply wish to say your article is as amazing.
The clarity for your post is simply nice and i can think you are
an expert in this subject. Fine together with your permission let me to seize your
feed to stay up to date with imminent post. Thanks 1,000,000
and please keep up the gratifying work.
geinoutime.com
Fang Jifan이 다시 말했습니다. “폐하, 내 아들 아, 할 일이 하나 더 있습니다.”
I absolutely love your blog.. Excellent colors & theme.
Did you create this site yourself? Please reply back
as I’m trying to create my own site and would like to know where you got this from
or just what the theme is named. Many thanks!
Fantastic web site. A lot of helpful info here.
I am sending it to several friends ans additionally sharing in delicious.
And certainly, thanks to your sweat!
Hello, everything is going perfectly here and ofcourse every one is sharing data,
that’s in fact excellent, keep up writing.
This information is invaluable. When can I find out more?
Amazing tons of fantastic info.
You actually make it appear really easy with your presentation but I to find this topic to be really one thing which I believe I might
by no means understand. It sort of feels too complex and very extensive for me.
I am taking a look forward to your next submit, I will attempt
to get the grasp of it!
Great post but I was wanting to know if you could write
a litte more on this topic? I’d be very thankful if
you could elaborate a little bit more. Cheers!
This design is incredible! You definitely
know how to keep a reader entertained. Between your wit and your videos, I was almost moved to start my own blog (well, almost…HaHa!) Wonderful job.
I really enjoyed what you had to say, and more than that,
how you presented it. Too cool!
I’d like to find out more? I’d like to find out some additional
information.
No matter if some one searches for his required thing, so he/she desires to be available
that in detail, therefore that thing is maintained over here.
You need to be a part of a contest for one of the best sites online.
I will highly recommend this web site!
Have you ever considered about including a little bit more than just your articles?
I mean, what you say is valuable and everything.
But think about if you added some great graphics or video clips to give
your posts more, “pop”! Your content is excellent but with pics and video
clips, this site could definitely be one of the best in its field.
Amazing blog!
Good day I am so happy I found your blog page, I really found you
by error, while I was searching on Google for something else, Anyways I am here now and would
just like to say many thanks for a fantastic post and a all round entertaining blog (I also love the theme/design), I don’t have
time to look over it all at the moment but I have saved it and also added in your RSS feeds, so when I have time I will be back to read more, Please do keep
up the fantastic work.
Hi to every single one, it’s in fact a pleasant for me
to visit this web page, it includes helpful Information.
Good day! Do you know if they make any plugins to protect against
hackers? I’m kinda paranoid about losing everything I’ve worked hard on. Any suggestions?
Hey just wanted to give you a quick heads up. The words in your content seem to be
running off the screen in Firefox. I’m not
sure if this is a format issue or something to do with internet browser compatibility but I thought I’d post to let you
know. The design look great though! Hope you get the problem fixed soon. Cheers
I’m now not positive the place you’re getting your info, however good topic.
I must spend a while learning much more or figuring out more.
Thanks for fantastic information I was on the lookout
for this information for my mission.
Every weekend i used to pay a visit this web page,
for the reason that i want enjoyment, since this this web page conations truly nice funny data too.
Промокоды 1win представляют собой специальные коды, пример promokod1WINbonus, которые позволяют пользователям получить дополнительные бонусы и преимущества при использовании платформы 1win. Благодаря этим промокодам, каждый зарегистрированный пользователь может получить дополнительные средства на свой игровой счет, бесплатные ставки или уникальные предложения.
Чтобы воспользоваться промокодами 1win, необходимо выполнить несколько простых шагов. Во-первых, необходимо зарегистрироваться на платформе 1win, создав личный аккаунт и указав все необходимые данные. После успешной регистрации, пользователь получает доступ к разнообразным промокодам, которые могут быть получены внутри системы или с помощью специальных акций и предложений.
Для активации промокода, пользователю необходимо перейти в свой личный кабинет на платформе 1win. Там требуется найти соответствующую вкладку или раздел, где можно ввести промокод. После ввода кода и нажатия на кнопку активации, система проверяет его на корректность и применяет соответствующие бонусы к аккаунту пользователя.
Промокоды 1win предоставляют широкий спектр возможностей для игроков. Некоторые промокоды предлагают дополнительные финансовые средства, которые можно использовать для ставок на спорт, казино или покер. Другие коды могут давать право на бесплатные ставки, бездепозитные бонусы или фриспины в слот-играх.
Однако, важно отметить, что промокоды 1win могут иметь определенные условия использования. Например, некоторые коды могут иметь ограниченный срок действия, требовать выполнения определенных условий или иметь минимальные требования к размеру ставки.
В целом, использование промокодов 1win – это отличная возможность получить дополнительные бонусы и преимущества на платформе. Они позволяют увеличить шансы на выигрыш и получить больше удовольствия от использования игровой платформы 1win. Не упустите возможность воспользоваться этими промокодами и получить максимальную выгоду от своей игры на 1win!
Источник https://promokod-1win-bonus.ru/
I know this if off topic but I’m looking into starting my own weblog and was
wondering what all is required to get set up?
I’m assuming having a blog like yours would cost a pretty penny?
I’m not very web savvy so I’m not 100% positive. Any tips or advice would be greatly appreciated.
Cheers
It’s going to be finish of mine day, but before
end I am reading this impressive piece of writing to increase my experience.
What i do not realize is in fact how you are not actually much more neatly-liked than you may be right
now. You are so intelligent. You recognize therefore significantly when it comes to this subject, made me for
my part consider it from a lot of numerous angles.
Its like men and women are not involved unless it is something
to do with Girl gaga! Your personal stuffs excellent.
Always handle it up!
No matter if some one searches for his vital thing, therefore he/she desires to be available that in detail, thus that
thing is maintained over here.
It’s not my first time to visit this web site, i am browsing this web page
dailly and take fastidious data from here daily.
We are a group of volunteers and starting a new scheme in our community.
Your web site provided us with valuable information to work on. You’ve done an impressive job and our whole community will be grateful to you.
Hello there, I found your site by the use of Google
even as looking for a comparable topic, your site got
here up, it seems great. I have bookmarked it in my google bookmarks.
Hi there, simply became aware of your weblog thru Google, and located that it is truly informative.
I am going to be careful for brussels. I’ll
appreciate if you happen to continue this in future. Lots
of folks might be benefited out of your writing. Cheers!
I am truly thankful to the owner of this web page who has shared this
enormous piece of writing at at this time.
I simply couldn’t depart your site before suggesting that I actually enjoyed the
standard information a person supply to your visitors? Is going to be again often to check
out new posts
Excellent blog here! Also your website loads up very fast!
What host are you using? Can I get your affiliate link to
your host? I wish my site loaded up as fast as yours lol
What’s up to every single one, it’s truly
a pleasant for me to visit this site, it includes precious Information.
I am in fact glad to glance at this blog posts which contains lots of useful facts, thanks for providing
these statistics.
Terrific work! That is the kind of info that are
meant to be shared across the net. Shame on the seek engines for now not positioning this post higher!
Come on over and discuss with my site . Thank you =)
bookmarked!!, I like your website!
Hey! I know this is kinda off topic but I was wondering if you knew where I could find a captcha plugin for my comment form?
I’m using the same blog platform as yours and I’m having difficulty
finding one? Thanks a lot!
Hi there mates, how is all, and what you want to say on the topic of this article, in my view its genuinely awesome in support of me.
https://www.bignewsnetwork.com/news/274382311/combining-modern-technology-and-a-creative-approach-to-business-with-del-mar-energy|
https://gb.kompass.com/p/kompass-posts/ua103618/business-of-the-future-investments-and-innovations-with-del/08959a1b-800b-4587-a919-628e6a117550/|
https://techduffer.com/business-of-the-future-investments-and-innovations-with-del-mar-energy/|
https://wikitechlibrary.com/del-mar-energy-successful-investments-in-the-american-oil-and-gas-sector/
Can I simply say what a relief to find someone
who actually knows what they’re talking about on the internet.
You definitely realize how to bring a problem to light and make
it important. More and more people have to read this and understand this side of
the story. I was surprised that you’re not more popular since you certainly have the gift.
spam konten and adult konten
this spam konten and adult konten
spam konten and adult konten
Hi, i think that i saw you visited my site thus i came to “return the favor”.I am attempting to find things to improve my site!I suppose its ok to use some of your ideas!!
Superb, what a website it is! This blog gives helpful facts to us, keep it up.
It’s a shame you don’t have a donate button! I’d definitely donate to this superb blog!
I suppose for now i’ll settle for book-marking and adding your RSS feed to my Google
account. I look forward to fresh updates and will talk about this website with my Facebook group.
Talk soon!
geinoutime.com
Alfonso는 일어서서 수행원을 바라보았습니다. “그는 어디에 있습니까?”
If you desire to grow your experience just keep visiting this web site and be updated with the most
up-to-date news update posted here.
This is really interesting, You’re a very skilled blogger.
I have joined your feed and look forward to seeking more of your
excellent post. Also, I’ve shared your website in my social networks!
This design is spectacular! You most certainly know
how to keep a reader amused. Between your wit and your videos, I was almost moved
to start my own blog (well, almost…HaHa!) Great job. I really enjoyed what you had to say,
and more than that, how you presented it. Too cool!
Hey I know this is off topic but I was wondering if
you knew of any widgets I could add to my blog that automatically tweet my newest twitter updates.
I’ve been looking for a plug-in like this for quite some time and was hoping maybe
you would have some experience with something like this.
Please let me know if you run into anything. I
truly enjoy reading your blog and I look forward
to your new updates.
I’m not sure why but this weblog is loading incredibly slow for me.
Is anyone else having this issue or is it a issue on my end?
I’ll check back later and see if the problem still exists.
Диплом О Высшем Образовании
Диплом О Высшем Образовании
Аттестат о среднем образовании приобретают при поступлении в колледжи, лицеи, ведь есть шанс перейти сразу на второй курс при наличии документа. Не важно, в какое учебное заведение вы решите поступить или на какую работу устроиться. Многие задаются вопросом, что больше всего пользуется спросом: сертификат Вуза или аттестаты. Во избежание возможных конфликтов с клиентом, мы предварительно все согласовываем по почте или другим удобным способом. Выиграть в конкурентной борьбе на рынке труда в Москве гораздо проще людям, у которых имеется оригинал документа с проведением о получении образования в ВУЗе. Обидно уступать хорошее место другим претендентам, зная, что великолепно справились бы с требованиями, указанными в вакансии.
http://https://russkiy365-diploms-srednee.ru/
Доверенность По Загранпаспорту
Если купить диплом фармацевта и стать дипломом О Высшем Образовании в этой отрасли, вы даже сможете создать свой собственный лекарственным препарат и найти способ лучше справляться с существующими заболеваниями с меньшим количеством побочных эффектов. Прежде всего, нужно отметить, что свидетельство о смерти, считается юридическим документом, который выдаётся, как подтверждающая выписка о смерти человека. Большинство боятся сделать ответственный шаг, продолжая трудится на отведенных местах без перспектив к росту. Иногда люди желают идти работать и получать знания непосредственно в процессе работы.
Зачетная Книжка Купить Москва
Покупателю следует ответственно подойти к выбору диплома, ведь он позволяет освоить новые сферы деятельности. Если вы до сих пор не знаете, где купить диплом УФ СГА, то мы к вашим услугам. Если у вас имеется диплом об окончании института, вам обеспечена должность с высокой заработной платой, “почитание” в кругу коллег и близких.
If some one needs expert view about blogging then i recommend him/her
to pay a visit this website, Keep up the good job.
Wow, fantastic weblog structure! How long have you ever been blogging for?
you made running a blog glance easy. The total look of your website is wonderful, let alone the content material!
http://henryfirearmsshop.com/where-you-can-buy-perfectmoney-reliable-platforms-and-exchanges/
Проверить Диплом О Высшем Профессиональном Образовании
Проверить Диплом О Высшем Профессиональном Образовании
Можно не сомневаться, сделка с незнакомым человеком не принесет ничего хорошего, максимум, на что можно рассчитывать так это ксерокопия оригинального бланка, а порой и всего лишь самодельный бланк, который с типографским не имеет ничего общего. Многие программисты, дизайнеры и журналисты самостоятельно прошли курс профессионального образованья в интернете, они много занимались и изучали информацию со всего мира. Это та же форма обучения, что и в ВУЗах, но только дома за компьютером без посещения занятий. В случае возникновения вопросов можно связаться с консультантом, который подробно и грамотно ответит на ваши вопросы. Мы сегодня стараемся сделать для своих клиентов всё возможное, чтобы помочь реализовать поставленные цели, независимо от места проживания и социального положения.
https://gruppa365-diploms-srednee.ru
Академический Международный Институт Ами
Покупка каждой разновидности бланка – специфична, обращайте на это внимание при выборе компании для сотрудничества. На рынке представлены бланки, наличие которых юридически подтверждает получение среднего полного, специального или высшего образования. Строганова Заказать МГЮА – Московская государственная юридическая академия им. Это дополнительная мера безопасности для наших пользователей, которая позволяет им быть уверенными в качестве предоставляемых услуг.
Купить Диплом О Высшем Образовании В Украине
Также, это может быть вариант для людей, которые уже имеют определенные знания и навыки в определенной области, но не имеют официального документа об образовании. 1 Трудового Кодекса РФ, квалификация работника – уровень знаний, умений, профессиональных навыков и опыта работы работника. Заказчикам все документы пересылаются по почте, а затем оплачиваются наложенным платежом.
Very quickly this website will be famous among all blog viewers, due to it’s fastidious content
Доверенность На Автомобиль Цена
К сожалению, не все могут похвастаться доверенностей, что имеют его, но получить диплом не так просто, как кажется на первый взгляд, по крайней мере, в государственных учебных заведениях. Нормативный срок обучения бакалавров в России составляет 4 года. Гарантии, которые мы предоставляем, дают возможность не переживать о соответствии автомобиля утвержденным законодательным нормам. И это открывает новые возможности для получения автомобиля Цена о высшем образовании. При этом соискатель даже может обладать нужными знаниями, а нередко уже и некоторым опытом работы в данной сфере. Что касается видов дипломов, то здесь доступны совершенно разные уровни, и диплом техникума и ВУЗа.
http://www.russkiy365-diploms-srednee.ru
Генеральная Доверенность На Машину Цена
Воспользоваться этой бесплатной услугой намного быстрее и эффективнее традиционного метода. При изготовлении дипломов используются только качественные бланки. Как обратить на себя внимание и выделиться из толпы уже при доверенности На Автомобиль Цена заявки на открытую вакансию. Итак, если выбор сделан и решено купить диплом высшего образования занесением в реестр, следует подумать, где лучше это сделать.
Купить Диплом О Среднем Специальном Образовании В Оренбурге
Мы понимаем, что для многих людей важно, чтобы их личная информация не попала в неправильные руки. Военкоматы, как правило, дают направления на медицинское обследование в согласованные медицинские доверенности На Автомобиль Цена. Купить диплом университета в Норильске государственный ПГУАС, а так же любого другого вуза можно на нашем сайте быстро и недорого.
This post will help the internet visitors
for setting up new weblog or even a blog from start to end.
Академ Справка
Академ Справка
Образование играет значительную роль в жизни, ведь без него непросто найти справку, выстроить карьеру и стать успешным. Вы станете владельцем абсолютно оригинальных “корочек”, которые всегда проходят любые проверки подлинности. Ускоренный формат – получите новую профессию фронтенд-разработчика. Такие Аттестат 2022 содержат все элементы защиты, и заполняются нашими сотрудниками по примерам официально выданных образцов. И тут нет ничего предосудительного, так как большинство из тех, кто закончили высшее учебное заведение и получили диплом, очень редко разбираются в специальности, которая указана на их корочке. Заказчик может выбрать срок гарантии, в течение которого сервис обязуется внести исправления в работу или вернуть денежные средства в случае неудовлетворительного качества выполнения заказа. Приобрести, настоящий диплом техникума Казахстана, человек, окончивший техникум, востребован как никогда, В сложной справки можно купить диплом техникума в Москве, рабочий человек с трудом представляет себя за партой, заказать диплом любого техникума россии с доставкой.
http://www.russkiy365-diploms-srednee.ru
Проверка Дипломов
В качестве источников каких-то фундаментальных знаний они подходят, но для цитирования нет. Множественные предложения по продаже официальных дипломов чаще всего не соответствуют действительности. Диплом государственного образца – это документ, который выдается после окончания высшего учебного заведения. Купить диплом специалиста, магистра или бакалавра – это выбор, который зависит от ваших целей и потребностей.
Купить Диплом О Высшем Образовании В Кирове
А что делать, если корочка о знаниях вдруг стала крайне необходима. При этом мы гарантируем справка наших отношений с клиентами. Важно проверять, чтобы все диагнозы, если есть непризывные диагнозы, были четко внесены в карту осмотра.
I’m not sure exactly why but this website is loading extremely slow for me.
Is anyone else having this issue or is it a issue on my end?
I’ll check back later on and see if the problem still exists.
Фоллаут 1 сезон смотреть онлайн
Hello just wanted to give you a quick heads up.
The text in your article seem to be running off the screen in Internet explorer.
I’m not sure if this is a format issue or something to
do with web browser compatibility but I thought I’d post to let
you know. The design look great though! Hope
you get the issue fixed soon. Many thanks
Today, I went to the beachfront with my kids. I found a sea shell and gave it to my 4 year
old daughter and said “You can hear the ocean if you put this to your ear.” She placed the
shell to her ear and screamed. There was a hermit crab
inside and it pinched her ear. She never wants to go back!
LoL I know this is entirely off topic but I had to tell someone!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196758
https://tubeteencam.com/user/darkevil31998/profile
https://permacultureglobal.org/users/58783-andy-globa
https://www.metal-archives.com/users/juyhtgrf1995197819981964
https://rentry.org/r6cits6t
I’m impressed, I must say. Rarely do I encounter
a blog that’s both equally educative and interesting,
and let me tell you, you’ve hit the nail on the head.
The issue is something which not enough people are speaking intelligently about.
I’m very happy I came across this in my hunt for something relating to this.
Right here is the perfect web site for anyone who would like to find
out about this topic. You understand so much its almost
hard to argue with you (not that I actually will need to…HaHa).
You certainly put a new spin on a topic which has
been written about for many years. Excellent stuff, just great!
I really love your blog.. Excellent colors & theme. Did you create this
site yourself? Please reply back as I’m planning to create my own personal
site and would love to know where you got this from or just what the theme is named.
Thanks!
Wow! This blog looks just like my old one! It’s on a entirely different topic but it has pretty much the same layout and design. Wonderful choice of colors!
Психолог 2024
Since the admin of this website is working, no hesitation very shortly it will be well-known, due to
its quality contents.
It’s an awesome paragraph in support of all the web viewers; they
will take advantage from it I am sure.
Amazing issues here. I am very glad to see your article. Thank you a lot and I’m taking a look forward to contact you. Will you kindly drop me a e-mail?
http://1g1552.mybb.ru/viewtopic.php?id=599#p708
https://goool.forumbb.ru/viewtopic.php?id=1007#p2266
http://kh.txi.ru/forum/index.php?
http://abroad.ekafe.ru/viewtopic.php?f=5&t=1787
http://kutejnikovo-61.ru/forum/messages/forum1/topic181/message180/?result=new#message180
For most recent news you have to pay a visit internet and
on internet I found this web site as a best website for newest updates.
Hi, this weekend is fastidious in support of me, because
this point in time i am reading this great educational article here
at my residence.
Hmm is anyone else having problems with the images on this blog loading? I’m trying to figure out if its a problem on my end or if it’s the blog. Any responses would be greatly appreciated.
http://enriquerp.listbb.ru/viewtopic.php?f=4&t=310
http://itbitgroup.ru/poluchite-diplom-prosto-byistro-nadezhno
http://x91392sl.beget.tech/2024/04/27/diplomy-na-zakaz-nasha-cel-vash-uspeh.html
http://dominion.listbb.ru/viewtopic.php?f=11&t=295
http://poltavagok.ru/poluchenie-poddelnyih-udostovereniy-o-vyisshem-obrazovanii/
Please let me know if you’re looking for a writer for your blog.
You have some really good articles and I feel I
would be a good asset. If you ever want to take some
of the load off, I’d really like to write some content for your blog in exchange for a
link back to mine. Please shoot me an e-mail if interested.
Many thanks!
I have read several just right stuff here. Definitely worth bookmarking for revisiting.
I surprise how a lot effort you put to create this kind of fantastic informative
site.
Thank you for the auspicious writeup. It in reality used to be a amusement account it. Look complicated to more introduced agreeable from you! By the way, how could we keep up a correspondence?
http://avrorp.getbb.ru/viewtopic.php?f=183&t=588
http://nnars.ru/forum/suggestion-box/1082-пїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅ#74468
https://www.import-moto.com/users/88
http://o91746bp.beget.tech/2024/04/27/diplomy-na-vybor-vash-put-k-professionalnomu-rostu.html
http://umorfishki.ru/izmenite-svoyu-zhizn-kupite-diplom-i-otkroyte-novyie-gorizontyi
My partner and I absolutely love your blog and find many of your post’s to be what precisely I’m
looking for. Do you offer guest writers to write content for
you personally? I wouldn’t mind creating a post or elaborating on some of the subjects you write about here.
Again, awesome blog!
Normally I do not read post on blogs, however I would like to say that
this write-up very forced me to take a look at and do so!
Your writing taste has been amazed me. Thank you, quite nice article.
For hottest news you have to pay a quick visit world-wide-web and on internet I
found this web page as a best website for most recent
updates.
We stumbled over here coming from a different page and thought I should check things out.
I like what I see so now i am following you. Look forward to finding out about your web page again.
Hello there, You have done a fantastic job. I’ll definitely digg it and personally suggest to my friends. I am sure they will be benefited from this web site.
http://forums.worldsamba.org/showthread.php?tid=992935
http://skazka.g-talk.ru/viewtopic.php?f=1&t=1641
http://forum.lazarevskaya.ru/showthread.php?p=294218#post294218
http://crimea.bestbb.ru/viewtopic.php?id=524#p658
http://shoptema.ru/forum/topic/31905/
Attractive section of content. I just stumbled upon your website and in accession capital to assert that I acquire in fact enjoyed account your blog posts. Anyway I’ll be subscribing to your feeds and even I achievement you access consistently rapidly.
http://www.kupitediplom0029.ru
My spouse and I stumbled over here by a different website
and thought I should check things out. I like what I see so i am just following you.
Look forward to checking out your web page for a second time.
Hello my loved one! I want to say that this article is awesome, great written and include
almost all significant infos. I would like to peer extra posts like
this .
Hello! This is my 1st comment here so I just wanted to give a quick shout out and tell you I truly enjoy reading your blog posts. Can you suggest any other blogs/websites/forums that cover the same topics? Thanks a ton!
kupitediplom0029.ru/
Thanks , I’ve just been searching for information approximately this topic for a long time and yours is the best I’ve discovered till now.
But, what about the bottom line? Are you positive
concerning the source?
This is my first time visit at here and i am really happy to read everthing at alone place.
https://www.kupitediplom0029.ru/
Today, I went to the beachfront with my children. I found a sea shell and gave it to my 4 year old daughter and said “You can hear the ocean if you put this to your ear.” She placed the shell
to her ear and screamed. There was a hermit crab inside and it pinched her ear.
She never wants to go back! LoL I know this is entirely
off topic but I had to tell someone!
Hi there everyone, it’s my first visit at this site, and post is truly fruitful
in support of me, keep up posting these types of content.
You are so interesting! I don’t believe I have read a single thing like that
before. So great to discover somebody with some
unique thoughts on this subject. Really.. many thanks for starting this up.
This website is one thing that is needed on the
web, someone with a little originality!
I have been surfing online more than 2 hours today, yet I never found any interesting article like yours.
It is pretty worth enough for me. In my view, if all site owners and bloggers made
good content as you did, the internet will be a lot
more useful than ever before.
Do you have a spam problem on this site; I also am a blogger, and I was wondering your situation; we have developed some nice procedures and we are looking to trade solutions with others, why not shoot me an e-mail if interested.
Thanks for the auspicious writeup. It if truth be told was a entertainment
account it. Glance complicated to far introduced agreeable from you!
However, how could we keep in touch?
I know this site provides quality dependent posts and extra data, is there any other web site which
gives such stuff in quality?
Fantastic post but I was wanting to know if you could write a litte more on this topic? I’d be very grateful if you could elaborate a little bit further. Kudos!
https://vip.7bb.ru/viewtopic.php?id=2683#p6562
http://testforum.1stbb.ru/viewtopic.php?f=3&t=332
https://mymink.5bb.ru/viewtopic.php?id=8136#p454028
http://forum.lazarevskaya.ru/showthread.php?p=294218#post294218
https://wegug.in/blogs/327/пїЅпїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅ-пїЅ-пїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅ-пїЅпїЅпїЅпїЅпїЅпїЅпїЅ
Hi to every body, it’s my first visit of this webpage; this webpage consists of remarkable and genuinely fine information in support of visitors.
https://www.diplomdoc.ru/
Great site you have here but I was wanting to know if you knew of any message boards that cover the same topics discussed here? I’d really like to be a part of group where I can get advice from other experienced individuals that share the same interest. If you have any suggestions, please let me know. Appreciate it!
http://https://okdiplom.com/
Great post. I was checking continuously this blog and I’m impressed!
Very helpful information particularly the last
part 🙂 I care for such information a lot. I was seeking this particular
info for a very long time. Thank you and good luck.
I appreciate, lead to I discovered just what I was looking
for. You have ended my 4 day long hunt! God Bless you
man. Have a great day. Bye
Hey there, You have done a great job. I’ll certainly digg it and personally suggest to my
friends. I am confident they will be benefited from this website.
These are in fact wonderful ideas in concerning blogging.
You have touched some nice factors here. Any way keep up wrinting.
Wow, awesome weblog layout! How long have you ever been running a
blog for? you made blogging look easy. The full look of your web site is great, as neatly as the content!
I enjoy what you guys are up too. This sort of clever work and
reporting! Keep up the great works guys I’ve added
you guys to my personal blogroll.
Pretty component to content. I simply stumbled upon your blog and in accession capital to say that I get actually
loved account your weblog posts. Any way I will be subscribing to your feeds or even I achievement
you get right of entry to constantly quickly.
Yesterday, while I was at work, my cousin stole my iphone and tested to see if it can survive a thirty foot drop, just so she can be a youtube sensation. My iPad is now broken and she has 83 views. I know this is totally off topic but I had to share it with someone!
http://asg-amt.phorum.pl/viewtopic.php?p=31684#31684
https://delta72.at.ua/forum/61-25662-1
https://forum.shvedun.ru/ucp.php?mode=login
http://dachkanews.ru/kupit-diplom-vash-bilet-k-luchshemu-budushhemu/
http://forum.safe-animals.ru/index.php?showtopic=26448
My partner and I absolutely love your blog and find a lot of your post’s to be exactly what I’m looking for. Would you offer guest writers to write content for you? I wouldn’t mind producing a post or elaborating on many of the subjects you write in relation to here. Again, awesome web site!
diplompro.ru
I am regular visitor, how are you everybody? This post posted at
this site is truly fastidious.
What’s up to every one, the contents present at this site are genuinely remarkable for people experience, well, keep up the nice work fellows.
https://diplompro.ru/
Today, I went to the beach front with my children. I found a sea shell and gave it to my 4 year old daughter and said “You can hear the ocean if you put this to your ear.” She placed the shell to her ear and screamed. There was a hermit crab inside and it pinched her ear. She never wants to go back! LoL I know this is entirely off topic but I had to tell someone!
https://magazin-diplomov.ru/
В современном мире, где диплом – это начало удачной карьеры в любой отрасли, многие стараются найти максимально быстрый и простой путь получения образования. Наличие документа об образовании сложно переоценить. Ведь диплом открывает двери перед любым человеком, желающим начать профессиональную деятельность или учиться в ВУЗе.
Предлагаем быстро получить этот важный документ. Вы сможете приобрести диплом старого или нового образца, что становится отличным решением для всех, кто не смог закончить обучение, потерял документ или хочет исправить свои оценки. Любой диплом изготавливается с особой аккуратностью, вниманием к мельчайшим элементам, чтобы на выходе получился полностью оригинальный документ.
Превосходство подобного решения заключается не только в том, что можно быстро получить свой диплом. Процесс организовывается комфортно, с нашей поддержкой. От выбора необходимого образца до точного заполнения личной информации и доставки в любой регион России — все под абсолютным контролем опытных мастеров.
В итоге, всем, кто ищет быстрый и простой способ получения требуемого документа, наша компания предлагает выгодное решение. Приобрести диплом – значит избежать долгого процесса обучения и сразу перейти к личным целям, будь то поступление в ВУЗ или старт карьеры.
http://www.sh5novosel.ru
Its such as you learn my thoughts! You seem to know so much about this, like
you wrote the e-book in it or something. I feel that you could do with
some p.c. to pressure the message home a bit, however instead
of that, that is excellent blog. A great read.
I’ll certainly be back.
I’ve been browsing online more than 4 hours today, yet I never found any interesting article like yours.
It is pretty worth enough for me. In my view, if all site owners
and bloggers made good content as you did, the web will be a lot more
useful than ever before.
Exceptional post however I was wondering if you could write a litte more on this subject? I’d be very grateful if you could elaborate a little bit further. Bless you!
institute-diplom.ru
Hi there friends, pleasant post and nice arguments commented at this place, I am genuinely enjoying by these.
institute-diplom.ru
Hi there, after reading this awesome paragraph i am as well delighted to share my know-how here with colleagues.
https://www.bisness-diplom.ru/
Howdy! This post couldn’t be written any better! Reading this post reminds me of my good old room mate! He always kept talking about this. I will forward this post to him. Fairly certain he will have a good read. Thanks for sharing!
http://www.zakaz-na-diplom.ru
I’m not sure why but this weblog is loading incredibly slow for me.
Is anyone else having this issue or is it a issue on my end?
I’ll check back later and see if the problem still exists.
Its like you read my mind! You appear to know so much about this, like you wrote the book in it or something. I think that you could do with some pics to drive the message home a bit, but other than that, this is wonderful blog. A great read. I will certainly be back.
http://www.diplom-city24.ru
What’s up mates, pleasant post and nice arguments commented at this place, I am genuinely enjoying by these.
http://https://peoplediplom.ru/
At this time it seems like BlogEngine is the top blogging platform available right now. (from what I’ve read) Is that what you are using on your blog?
https://peoplediplom.ru/
Hello Dear, are you genuinely visiting this website daily, if so after that you will absolutely take fastidious know-how.
http://https://diplom45.ru/
A Sobreezybabe company.
Explore Clean Glam’s range of clean beauty cosmetics and natural skincare products.
From organic skincare routines to vegan beauty products, we lead the clean beauty market with eco-friendly, non-toxic solutions.
Learn how to create effortless Clean Glam Makeup looks with our step by step tutorials!
Discover holistic beauty solutions for a radiant you!
Hello, I would like to subscribe for this webpage to take most recent updates, therefore where can i do it please help out.
http://www.1magistr.ru
Thanks for the good writeup. It in truth was a amusement account it. Glance advanced to more introduced agreeable from you! However, how could we keep up a correspondence?
http://https://kdiplom.ru/
obviously like your website but you have to take a look at
the spelling on several of your posts. Several of them
are rife with spelling problems and I find it very troublesome to tell the truth on the other hand
I will certainly come back again.
Hey I know this is off topic but I was wondering if you knew of any
widgets I could add to my blog that automatically tweet my newest
twitter updates. I’ve been looking for a plug-in like this for quite some time and was hoping maybe you would have some experience with something like
this. Please let me know if you run into anything.
I truly enjoy reading your blog and I look forward to your new updates.
all the time i used to read smaller posts which as
well clear their motive, and that is also happening with this article which I am reading at this time.
I all the time used to read piece of writing in news papers but now as
I am a user of web thus from now I am using net for articles
or reviews, thanks to web.
Hey! Do you know if they make any plugins to assist with Search Engine Optimization? I’m trying to get my blog to rank for
some targeted keywords but I’m not seeing very good gains.
If you know of any please share. Kudos!
Unleash your wanderlust! Explore the world with Travel in Beautiful Fashion, your ultimate travel blog.
Compare airfares on over 700+ budget airlines, book unique rental cars, and discover cultural immersion or cultural travel hidden gems with our
expert-curated travel guides and insider travel tips.
Browse our Wanderlust digital nomad Travel blog for articles on Eco-tourism, solo travel, flashpacking, responsible travel,
beautiful camping and more!
I really like it when people come together and share ideas.
Great website, stick with it!
Superb website you have here but I was wondering if you knew of any message boards that
cover the same topics discussed here? I’d really like to be a part of group where I can get advice from other experienced people that share the
same interest. If you have any recommendations, please let me know.
Cheers!
I do believe all of the concepts you have offered to your post.
They are very convincing and will definitely work.
Nonetheless, the posts are very quick for newbies.
May just you please prolong them a little from subsequent
time? Thank you for the post.
Great goods from you, man. I have understand your stuff previous to
and you’re just too great. I really like what you’ve acquired here, certainly like what you’re saying and the way in which you say
it. You make it entertaining and you still care for to keep it
sensible. I cant wait to read far more from you. This
is actually a terrific site.
I’ve been browsing online more than 4 hours today, yet I never found any interesting article like yours.
It is pretty worth enough for me. In my view, if
all web owners and bloggers made good content as you
did, the net will be a lot more useful than ever before.
Yesterday, while I was at work, my cousin stole my iPad and tested to see if it can survive a 25 foot drop, just so she can be a youtube sensation. My iPad is now broken and she has 83 views. I know this is completely off topic but I had to share it with someone!
https://www.diplom-city24.ru/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/e985s6hn
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195979
https://cannabis.net/user/151878
https://www.quia.com/profiles/gillho
https://haveagood.holiday/users/351204
Great blog you have here but I was wondering if you knew of any discussion boards that cover the same topics discussed here? I’d really like to be a part of community where I can get opinions from other experienced people that share the same interest. If you have any suggestions, please let me know. Many thanks!
https://www.zakaz-na-diplom.ru/
This is really interesting, You are a very skilled blogger. I’ve joined your feed and look forward to seeking more of your fantastic post. Also, I’ve shared your web site in my social networks!
https://institute-diplom.ru/
Hi, everything is going fine here and ofcourse every one is sharing data, that’s really excellent, keep up writing.
https://medium.com/@oximus777/диплом-по-вашему-выбору-гарантированное-качество-и-конфиденциальность-40f9f0a1683e
http://forum.vorchun.ru/viewtopic.php?f=2&t=249627
https://szr-coll-isk.ru/forum/messages/forum1/topic309/message379/?result=new#message379
The other day, while I was at work, my sister stole my apple ipad and tested to see if it can survive a thirty foot drop, just so she can be a youtube sensation. My iPad is now broken and she has 83 views. I know this is completely off topic but I had to share it with someone!
https://www.diplom-kuplu.ru/
Amazing! Its genuinely amazing paragraph, I have got much clear idea about from this piece of writing.
https://diplom-kuplu.ru/
geinoutime.com
마음에 도를 갖고 마음으로 도를 섬기는 모든 기술을 배우십시오.
Thank you for any other informative blog. Where else may I am getting that kind of info written in such an ideal means? I’ve a challenge that I’m simply now running on, and I have been at the look out for such info.
https://www.hashtap.com/@artur.sh/путь-к-профессиональному-успеху-приобрети-диплом-и-стань-экспертом-2WgOB7y3NJpm
https://www.neymarfootballforum.com/read-blog/1279
http://ya.bestbb.ru/viewtopic.php?id=2783#p6269
It’s an amazing paragraph designed for all the online visitors; they will get benefit from
it I am sure.
Hi there, just became alert to your blog through
Google, and found that it’s truly informative. I’m
gonna watch out for brussels. I will appreciate if you continue this in future.
Numerous people will be benefited from your writing.
Cheers!
whoah this weblog is excellent i really like studying your posts.
Keep up the great work! You know, a lot of persons are
searching around for this info, you could help them greatly.
For most recent news you have to pay a quick visit the web and on the web I found this web page as a best web page for newest updates.
https://www.easyinsurancehub.co.uk/forum/thread-5609.html
http://technoevents.ru/gotovsya-k-vyizovam-ryinka-priobreti-diplom-i-stan-konkurentosposobnyim/
http://www.les-ailes-chalaisiennes.com/forum/topic/купить-диплом-ваш-ключ-к-новым-возможн/#postid-18641
I like the helpful info you provide in your articles.
I will bookmark your weblog and check again here regularly.
I am quite sure I will learn many new stuff right here!
Good luck for the next!
Remarkable issues here. I am very satisfied to see your article. Thanks a lot and I am having a look ahead to contact you. Will you kindly drop me a e-mail?
http://animalplanetnews.ru/diplom-po-zhelaniyu-priobreti-obrazovanie-kotoroe-sootvetstvuet-tvoim-interesam
http://www.rrsclub.ru/showthread.php?p=41971#post41971
https://vip.7bb.ru/viewtopic.php?id=2705#p6597
I enjoy what you guys are up too. This sort of clever work and coverage!
Keep up the fantastic works guys I’ve included you guys to our
blogroll.
No matter if some one searches for his required thing, therefore he/she wishes to be available
that in detail, so that thing is maintained over here.
Good day! This is my first visit to your blog!
We are a team of volunteers and starting a new project in a community
in the same niche. Your blog provided us valuable information to work on. You have done a marvellous
job!
Heya i’m for the first time here. I found this board
and I in finding It really useful & it helped me out much.
I hope to offer one thing back and help others like you aided me.
If some one needs to be updated with latest technologies then he must be
pay a quick visit this site and be up to date all the time.
Howdy, i read your blog occasionally and i own a similar one and i was just
curious if you get a lot of spam remarks? If so how do you stop it,
any plugin or anything you can advise? I get so much lately it’s driving me crazy
so any assistance is very much appreciated.
At this time it looks like Movable Type is the best blogging platform out there right now. (from what I’ve read) Is that what you are using on your blog?
http://sfera100.ru/forum/user/2471/
http://censornet.ru/put-k-professionalnomu-uspehu-priobreti-diplom-i-stan-ekspertom/
http://kids-news.ru/byistryiy-put-k-karere-kak-kupit-diplom-i-dostich-uspeha
Hello it’s me, I am also visiting this site regularly, this
web page is genuinely pleasant and the visitors are actually
sharing pleasant thoughts.
Wonderful blog! I found it while surfing around on Yahoo News.
Do you have any suggestions on how to get listed in Yahoo News?
I’ve been trying for a while but I never seem to get there!
Appreciate it
Oh my goodness! Incredible article dude! Many thanks, However
I am experiencing troubles with your RSS. I don’t know why I am unable to
join it. Is there anyone else getting identical RSS problems?
Anyone that knows the answer will you kindly respond?
Thanks!!
I’ve been exploring for a bit for any high-quality articles or weblog
posts in this kind of space . Exploring in Yahoo I eventually
stumbled upon this website. Reading this information So i am satisfied to show that
I have an incredibly just right uncanny feeling I discovered just what I needed.
I such a lot indubitably will make sure to don?t
overlook this website and provides it a glance on a constant basis.
Superb website you have here but I was curious about if you knew of
any message boards that cover the same topics talked
about here? I’d really like to be a part of community where I can get advice from other knowledgeable individuals that share
the same interest. If you have any recommendations,
please let me know. Thanks a lot!
I have been browsing online more than three hours today, yet I
never found any interesting article like yours. It is pretty
worth enough for me. In my opinion, if all web owners and
bloggers made good content as you did, the net will be a lot more
useful than ever before.
Pretty! This has been an incredibly wonderful post. Thanks for providing these details.
http://forexgroupx.ru/kupit-diplom-pervyiy-shag-k-uspeshnoy-karere
https://etkinigoster.com/blogs/238501/ упить-диплом-ваш-шанс-получить-высококлассное-образование-без-лишних-хлопот
https://rodme.ru/diplom-mechty-vash-put-k-professionalnomu-priznaniyu-i-uvajeniyu-t12181.html
Hey there! I know this is kinda off topic however , I’d figured I’d ask.
Would you be interested in trading links or maybe guest writing a blog post or vice-versa?
My blog goes over a lot of the same subjects as yours and I think we
could greatly benefit from each other. If
you happen to be interested feel free to send me an e-mail.
I look forward to hearing from you! Awesome blog by the way!
Wow, incredible blog layout! How long have you been blogging for?
you make blogging look easy. The overall look of your website is fantastic, let alone the content!
I do not even know the way I finished up here,
however I thought this publish was good. I do not understand who you might be but certainly you
are going to a famous blogger in the event you aren’t already.
Cheers!
I am not certain the place you are getting your info, but good topic.
I must spend a while studying more or working out more.
Thanks for fantastic information I used to be searching for this info for my mission.
This design is spectacular! You certainly
know how to keep a reader entertained. Between your wit and your
videos, I was almost moved to start my own blog (well, almost…HaHa!) Wonderful job.
I really enjoyed what you had to say, and more than that, how
you presented it. Too cool!
This is my first time pay a quick visit at here and i am in fact impressed to read everthing at single place.
http://superforum.spybb.ru/viewtopic.php?id=2480#p50275
http://www.markusragger.at/chess/index.php/kforum/jm-news-pro-module/633978
http://newfinbiz.ru/diplom-s-garantiey-kachestva-gde-priobresti-dokument-kotoryiy-otvechaet-vyisokim-standartam
Hello there, just became aware of your blog through Google,
and found that it’s truly informative. I’m gonna
watch out for brussels. I will be grateful if you continue this in future.
Numerous people will be benefited from your writing.
Cheers!
always i used to read smaller articles or reviews which also clear their motive, and that is also happening with this post which
I am reading now.
Nice post. I was checking continuously this blog and I am impressed!
Very helpful info specifically the last part 🙂 I care for such information a lot.
I was seeking this certain info for a very long time. Thank you
and best of luck.
whoah this blog is magnificent i really like reading your articles.
Keep up the great work! You know, many people are hunting around
for this information, you can help them greatly.
В нашем мире, где диплом – это начало успешной карьеры в любом направлении, многие пытаются найти максимально простой путь получения качественного образования. Наличие документа об образовании сложно переоценить. Ведь именно диплом открывает двери перед всеми, кто хочет вступить в сообщество профессионалов или продолжить обучение в ВУЗе.
Наша компания предлагает очень быстро получить любой необходимый документ. Вы сможете заказать диплом старого или нового образца, и это является выгодным решением для человека, который не смог закончить обучение или потерял документ. Каждый диплом изготавливается с особой аккуратностью, вниманием ко всем элементам, чтобы в результате получился полностью оригинальный документ.
Преимущества подобного подхода заключаются не только в том, что можно максимально быстро получить диплом. Процесс организован удобно и легко, с нашей поддержкой. Начиная от выбора подходящего образца диплома до точного заполнения личных данных и доставки в любое место России — все находится под абсолютным контролем качественных специалистов.
Всем, кто ищет быстрый способ получить необходимый документ, наша услуга предлагает выгодное решение. Приобрести диплом – это значит избежать долгого обучения и не теряя времени перейти к достижению своих целей, будь то поступление в ВУЗ или старт успешной карьеры.
http://forumcar.mybb.ru/viewtopic.php?id=66#p67
http://fiatforum.5bb.ru/viewtopic.php?id=920#p5342
http://ukrainaru.bbok.ru/viewtopic.php?id=6875#p6923
http://www.pets-ural.ru/content/forum/viewtopic.php?p=57109#57109
https://socialcompare.com/en/member/josearnold2-7i45t61v
I’m not sure exactly why but this weblog is loading extremely slow for me.
Is anyone else having this problem or is it a problem on my
end? I’ll check back later on and see if the problem still exists.
Very quickly this website will be famous amid all blog visitors, due to
it’s nice posts
Excellent blog you have here but I was wondering if you knew of any message boards that cover the
same topics talked about in this article? I’d really love to be a part of online community where I can get feed-back from other knowledgeable people that share the same
interest. If you have any recommendations, please let me know.
Thanks a lot!
Undeniably believe that which you said. Your favorite justification appeared to be on the net the easiest thing to be
aware of. I say to you, I certainly get irked while people consider worries
that they just don’t know about. You managed to hit
the nail upon the top and defined out the whole thing without having
side-effects , people can take a signal. Will probably be back to get more.
Thanks
Hi there, for all time i used to check website posts here in the early hours in the dawn, since i like
to find out more and more.
Does your site have a contact page? I’m having a tough time locating it but, I’d like
to send you an email. I’ve got some ideas for your blog you might be interested in hearing.
Either way, great website and I look forward to seeing it grow over time.
My partner and I absolutely love your blog and find nearly all of your post’s to be what precisely I’m looking for. Would you offer guest writers to write content in your case? I wouldn’t mind writing a post or elaborating on most of the subjects you write regarding here. Again, awesome weblog!
https://www.lffix.dk/revolutionize-your-business-with-our-cutting-edge/
http://mykinotime.ru/page/3/
http://www.uinvestor.ru/
http://www.realtalkwithnthabi.co.za/2017/11/17/alatos-performs-time-seminar/
http://www.ajeci.com.br/2017/09/28/palestra-solucoes-tigre-ads-em-tubos-corrugados-para-drenagem-e-saneamento/
Excellent article. I certainly love this site. Continue the good
work!
This article provides clear idea designed for the new viewers of blogging, that genuinely how
to do blogging and site-building.
I all the time used to study piece of writing in news papers but now as I am a
user of internet thus from now I am using net for articles or reviews,
thanks to web.
Hi! This is my first comment here so I just wanted to give
a quick shout out and say I genuinely enjoy reading your articles.
Can you recommend any other blogs/websites/forums that deal with the same topics?
Thank you so much!
Greetings! I know this is kinda off topic but I was wondering if you knew where I could find a captcha plugin for my comment form?
I’m using the same blog platform as yours and I’m having
difficulty finding one? Thanks a lot!
레프리칸 리치스
Ouyang Zhi는 약간 눈살을 찌푸렸습니다. “내시 양, Jinzhou는 우리가 여기 있다는 것을 어떻게 압니까?”
Saved as a favorite, I really like your web site!
I’m really enjoying the theme/design of your website.
Do you ever run into any internet browser compatibility issues?
A number of my blog visitors have complained about my blog not operating correctly in Explorer but looks great in Chrome.
Do you have any solutions to help fix this problem?
Thanks for your marvelous posting! I certainly enjoyed reading it, you can be
a great author.I will make certain to bookmark your blog
and will often come back from now on. I want to encourage yourself to continue your great writing,
have a nice holiday weekend!
Very good post. I’m going through a few of these issues as
well..
I seriously love your website.. Very nice colors & theme.
Did you make this amazing site yourself? Please reply back as I’m attempting to create my own website and want to know where you got this from
or just what the theme is named. Many thanks!
I’ve been surfing online more than 3 hours today, yet I never
found any interesting article like yours. It is pretty worth
enough for me. Personally, if all webmasters and bloggers made good content
as you did, the net will be a lot more useful than ever
before.
situs bokep spam xnxx
It is appropriate time to make some plans for the future and it’s time to be happy.
I have read this post and if I could I want to suggest you
some interesting things or advice. Perhaps you could write next articles referring to this article.
I want to read even more things about it!
Hi, i believe that i noticed you visited my website thus i got
here to go back the favor?.I am attempting to
find issues to enhance my web site!I suppose its good enough to make use of a few of your ideas!!
Hey! I just wanted to ask if you ever have any issues with hackers?
My last blog (wordpress) was hacked and I ended up losing months of hard work due to no
back up. Do you have any solutions to protect against hackers?
I needed to thank you for this excellent read!! I certainly enjoyed every bit
of it. I have you book-marked to look at new stuff
you post…
If you want to increase your knowledge simply keep visiting
this website and be updated with the latest news update posted here.
You made some good points there. I looked on the web for more info about the
issue and found most people will go along with your views on this site.
Pretty! This has been a really wonderful post. Many thanks for supplying these details.
We absolutely love your blog and find nearly all of your post’s to be precisely what I’m
looking for. Do you offer guest writers to write content for
yourself? I wouldn’t mind writing a post or elaborating on a
number of the subjects you write related to
here. Again, awesome weblog!
Hi this is kind of of off topic but I was wondering if blogs use WYSIWYG
editors or if you have to manually code with HTML. I’m starting a blog soon but have
no coding skills so I wanted to get guidance from someone with experience.
Any help would be enormously appreciated!
certainly like your web site however you have to check the spelling on quite a few of
your posts. A number of them are rife with spelling problems and I to find it very troublesome to tell the reality
however I will definitely come again again.
Great post. I was checking constantly this weblog and I am impressed!
Extremely useful information specifically the last phase :
) I maintain such info much. I was looking for this particular info for a very long time.
Thank you and good luck.
The other day, while I was at work, my sister stole
my apple ipad and tested to see if it can survive a forty foot drop,
just so she can be a youtube sensation. My iPad is now broken and she has 83 views.
I know this is entirely off topic but I had to share it with someone!
After I initially commented I appear to have clicked the -Notify me when new comments are added- checkbox and now
each time a comment is added I get 4 emails with the same comment.
There has to be a means you are able to remove me from that service?
Thanks a lot!
I do agree with all of the ideas you’ve presented for your post.
They are really convincing and will certainly work.
Nonetheless, the posts are very brief for beginners. Could you
please lengthen them a little from subsequent time? Thank you for
the post.
This website definitely has all the information I needed concerning this subject and didn’t know who to ask.
We are a group of volunteers and opening a brand new scheme in our community.
Your site offered us with helpful info to work on. You’ve
performed a formidable activity and our entire neighborhood might be thankful to you.
Hello there, just became alert to your blog through Google, and found that it’s really informative.
I’m going to watch out for brussels. I will appreciate
if you continue this in future. A lot of people will be benefited from your writing.
Cheers!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/terocona1986/about_me/
https://haveagood.holiday/users/352199
http://www.nfomedia.com/profile?uid=rOiZejB
https://permacultureglobal.org/users/59121-sarah-johnson
http://www.babelcube.com/user/alicia-williams
Good day! Do you know if they make any plugins to help with Search Engine Optimization?
I’m trying to get my blog to rank for some targeted keywords but I’m not
seeing very good gains. If you know of any please
share. Thanks!
Quality content is the secret to invite the viewers to pay
a visit the site, that’s what this site is providing.
Сегодня, когда диплом – это начало отличной карьеры в любой области, многие ищут максимально быстрый и простой путь получения образования. Наличие официального документа переоценить невозможно. Ведь диплом открывает дверь перед любым человеком, который желает начать трудовую деятельность или учиться в высшем учебном заведении.
Предлагаем быстро получить любой необходимый документ. Вы сможете заказать диплом, и это является выгодным решением для человека, который не смог завершить образование или утратил документ. диплом изготавливается с особой аккуратностью, вниманием к мельчайшим нюансам, чтобы в результате получился документ, 100% соответствующий оригиналу.
Преимущества подобного решения состоят не только в том, что вы быстро получите диплом. Процесс организовывается удобно, с профессиональной поддержкой. Начав от выбора требуемого образца документа до консультаций по заполнению личной информации и доставки в любое место страны — все будет находиться под полным контролем опытных мастеров.
Для тех, кто хочет найти максимально быстрый способ получения необходимого документа, наша компания предлагает отличное решение. Купить диплом – значит избежать долгого процесса обучения и сразу перейти к своим целям, будь то поступление в ВУЗ или начало удачной карьеры.
http://devushka.winbb.ru/viewtopic.php?id=684#p3322
http://fkvrn.webtalk.ru/viewtopic.php?id=696#p21973
https://sponforum.ixbb.ru/viewtopic.php?id=12316#p42418
http://fabrikaglamura.webtalk.ru/viewtopic.php?id=771#p39764
http://www.rrsclub.ru/showthread.php?p=38647#post38647
В наше время, когда диплом является началом удачной карьеры в любом направлении, многие ищут максимально простой путь получения качественного образования. Наличие официального документа переоценить невозможно. Ведь именно он открывает дверь перед всеми, кто собирается вступить в сообщество профессионалов или продолжить обучение в ВУЗе.
В данном контексте наша компания предлагает быстро получить этот важный документ. Вы имеете возможность купить диплом, что является выгодным решением для человека, который не смог закончить обучение или потерял документ. дипломы производятся аккуратно, с особым вниманием к мельчайшим деталям. На выходе вы сможете получить документ, 100% соответствующий оригиналу.
Преимущество такого подхода состоит не только в том, что вы максимально быстро получите диплом. Весь процесс организован просто и легко, с профессиональной поддержкой. От выбора необходимого образца документа до консультации по заполнению персональной информации и доставки по России — все под абсолютным контролем квалифицированных мастеров.
Всем, кто ищет быстрый способ получить требуемый документ, наша компания предлагает выгодное решение. Приобрести диплом – это значит избежать продолжительного обучения и не теряя времени перейти к достижению своих целей, будь то поступление в ВУЗ или начало карьеры.
Купить сертификат специалиста
each time i used to read smaller content which also
clear their motive, and that is also happening with this post which I am
reading at this time.
What’s up, yup this article is truly good and
I have learned lot of things from it on the topic of blogging.
thanks.
I love your blog.. very nice colors & theme. Did you design this website yourself
or did you hire someone to do it for you? Plz respond as I’m looking to construct my own blog and would like to know where u got
this from. many thanks
situsxnxx
situsxnxx
situsxnxx
situsxnxx
Seriously a lot of terrific info!
What a data of un-ambiguity and preserveness of valuable experience about unpredicted emotions.
geinoutime.com
다행스럽게도 Li Chaowen은 피부가 두껍고 모두의 동정적인 시선에 눈을 멀게했습니다.
situs bokep
situs bokep
situs bokep
Hi are using WordPress for your site platform?
I’m new to the blog world but I’m trying to get started and create my
own. Do you need any coding expertise to make your own blog?
Any help would be greatly appreciated!
Pretty section of content. I just stumbled upon your web site and in accession capital to assert that
I acquire actually enjoyed account your blog posts.
Anyway I will be subscribing to your augment and even I achievement you access consistently rapidly.
An impressive share! I have just forwarded this onto a co-worker who had been conducting a little research on this. And he actually bought me lunch due to the fact that I discovered it for him… lol. So allow me to reword this…. Thanks for the meal!! But yeah, thanks for spending the time to talk about this issue here on your internet site.
https://opensocialfactory.com/story16458078/%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC-%D1%81%D0%BF%D0%B5%D1%86%D0%B8%D0%B0%D0%BB%D0%B8%D1%81%D1%82%D0%B0
https://thenextnovator.com.vn/
https://www.360nhadep.com/
https://politictoday.ru/kupit-diplom-legkiy-put-k-uspeshnoy-karere
http://aboutallfinance.ru/page/3
It’s awesome to go to see this site and reading
the views of all colleagues regarding this post, while I
am also zealous of getting know-how.
I absolutely love your blog and find most of your post’s to be just what I’m looking for.
can you offer guest writers to write content to suit your needs?
I wouldn’t mind producing a post or elaborating on a number of the subjects you write in relation to here.
Again, awesome weblog!
Heya are using WordPress for your blog platform?
I’m new to the blog world but I’m trying to get started and set up
my own. Do you require any coding expertise to make your own blog?
Any help would be really appreciated!
Thanks for every other informative website. The place else could I
get that type of information written in such a perfect means?
I’ve a undertaking that I am just now operating on, and I’ve been at
the glance out for such information.
Hi there! I know this is kind of off topic but I was wondering
which blog platform are you using for this site? I’m getting fed up of WordPress because
I’ve had issues with hackers and I’m looking at options for another platform.
I would be awesome if you could point me in the direction of
a good platform.
Hi are using WordPress for your blog platform?
I’m new to the blog world but I’m trying to get started and set up my own. Do you need any
html coding expertise to make your own blog? Any help would
be greatly appreciated!
I was able to find good information from your blog posts.
В современном мире, где диплом является началом успешной карьеры в любом направлении, многие пытаются найти максимально быстрый путь получения качественного образования. Факт наличия официального документа сложно переоценить. Ведь диплом открывает дверь перед каждым человеком, который хочет начать трудовую деятельность или продолжить обучение в высшем учебном заведении.
Мы предлагаем быстро получить этот необходимый документ. Вы сможете заказать диплом нового или старого образца, и это становится удачным решением для всех, кто не смог закончить обучение, потерял документ или хочет исправить плохие оценки. диплом изготавливается аккуратно, с особым вниманием ко всем деталям, чтобы в итоге получился продукт, полностью соответствующий оригиналу.
Превосходство подобного подхода заключается не только в том, что можно оперативно получить диплом. Процесс организован удобно, с профессиональной поддержкой. Начиная от выбора требуемого образца диплома до консультации по заполнению личных данных и доставки в любое место России — все под абсолютным контролем опытных мастеров.
Всем, кто ищет оперативный способ получить необходимый документ, наша компания предлагает отличное решение. Заказать диплом – это значит избежать продолжительного процесса обучения и сразу перейти к достижению собственных целей, будь то поступление в университет или начало трудовой карьеры.
diplomsagroups.com
В современном мире, где диплом становится началом удачной карьеры в любой области, многие стараются найти максимально простой путь получения образования. Важность наличия документа об образовании трудно переоценить. Ведь диплом открывает двери перед каждым человеком, желающим вступить в сообщество профессиональных специалистов или учиться в университете.
Наша компания предлагает максимально быстро получить этот важный документ. Вы сможете приобрести диплом нового или старого образца, что будет выгодным решением для человека, который не смог закончить обучение или утратил документ. Любой диплом изготавливается аккуратно, с особым вниманием ко всем деталям. В итоге вы сможете получить документ, максимально соответствующий оригиналу.
Преимущества такого решения заключаются не только в том, что вы максимально быстро получите диплом. Процесс организован комфортно, с нашей поддержкой. От выбора требуемого образца документа до консультаций по заполнению личных данных и доставки в любой регион России — все под полным контролем наших мастеров.
В результате, всем, кто ищет оперативный способ получить требуемый документ, наша компания предлагает выгодное решение. Заказать диплом – значит избежать длительного обучения и не теряя времени перейти к достижению собственных целей, будь то поступление в ВУЗ или старт карьеры.
https://www.gimnazija-ivanjica.edu.rs/mobilearn/claroline/phpbb/viewtopic.php?topic=259&cidReset=true&cidReq=INF1
https://www.mapleprimes.com/users/JoseArnold2
https://znayvse.mybb.ru/viewtopic.php?id=13464#p29870
http://secondwife.mybb.ru/viewtopic.php?id=942#p240095
https://lwccareers.lindsey.edu/profiles/4687735-lonnie-kelley
Hey just wanted to give you a quick heads up.
The words in your post seem to be running off the screen in Chrome.
I’m not sure if this is a format issue or something
to do with browser compatibility but I thought I’d post to let you know.
The layout look great though! Hope you get the issue solved soon. Cheers
It’s amazing designed for me to have a website, which is good in favor of my know-how. thanks admin
https://censornet.ru/page/6/
https://bookmarkspecial.com/story17167418/%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC-%D0%BE-%D0%B2%D1%8B%D1%81%D1%88%D0%B5%D0%BC-%D0%BE%D0%B1%D1%80%D0%B0%D0%B7%D0%BE%D0%B2%D0%B0%D0%BD%D0%B8%D0%B8
https://beaukvwvt.dsiblogger.com/59592722/the-definitive-guide-to-marketing
http://fabnews.ru/forum/showthread.php?p=76080&mode=threaded
https://socialbuzzmaster.com/story2551710/%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC-%D1%81%D1%82%D0%BE%D0%BC%D0%B0%D1%82%D0%BE%D0%BB%D0%BE%D0%B3%D0%B0
situs xnxx spam adult
I enjoy what you guys are up too. Such clever work and exposure!
Keep up the fantastic works guys I’ve you guys to my own blogroll.
WOW just what I was looking for. Came here by searching for https://glints.com/vn/profile/public/018d8d3d-ce51-4621-855f-76c10518b61f
I am regular reader, how are you everybody? This paragraph posted at this web site
is truly pleasant.
I just could not depart your web site prior to suggesting that I extremely
enjoyed the standard information an individual supply for your visitors?
Is going to be again continuously in order
to check up on new posts
I’ll immediately clutch your rss feed as I can’t to find your e-mail subscription link or e-newsletter service.
Do you have any? Please permit me know so that I may just subscribe.
Thanks.
It’s not my first time to visit this site, i am
browsing this web site dailly and get pleasant facts from here everyday.
I loved as much as you’ll receive carried out right here. The sketch is tasteful, your authored subject matter stylish.
nonetheless, you command get got an impatience over that you wish be delivering the following.
unwell unquestionably come further formerly again as exactly the same
nearly very often inside case you shield this hike.
Hello i am kavin, its my first occasion to commenting anyplace,
when i read this article i thought i could also create comment due to this good article.
First of all I want to say superb blog! I had a quick question that
I’d like to ask if you don’t mind. I was curious to find out how you center yourself and clear
your mind prior to writing. I’ve had a difficult time clearing
my thoughts in getting my ideas out there. I truly do enjoy writing but it just seems like the first 10 to 15 minutes tend to be lost simply just trying
to figure out how to begin. Any suggestions or tips?
Many thanks!
Thanks for finally talking about > Fornix (crus)/
Stria Terminalis – fxst | Whole Brain Protocol for Tractography with Empirical MRI < Loved it!
After I initially left a comment I seem to have clicked the -Notify me
when new comments are added- checkbox and now every time a comment
is added I receive four emails with the same comment.
There has to be a means you are able to remove me from that service?
Thanks!
It’s going to be finish of mine day, but before finish I am reading
this fantastic piece of writing to improve my experience.
Generally I do not read article on blogs, however I wish to
say that this write-up very forced me to try and do it! Your writing style has
been amazed me. Thank you, quite nice article.
terbukti main di Aladdin138 situs game online paling mudah menghasilkan cuan tambahan harian, semua game yang dimainkan sangat mudah dan gampang dimenangkan untuk menghasilkan uang jajan lebih
LOGIN — https://wiganutc.org
Today, while I was at work, my cousin stole my iPad and tested to see if it can survive a thirty foot drop,
just so she can be a youtube sensation. My iPad is now broken and she has 83 views.
I know this is totally off topic but I had to share it with someone!
Excellent weblog right here! Also your website rather a lot up fast!
What web host are you the use of? Can I am getting your
affiliate hyperlink for your host? I desire my website loaded
up as quickly as yours lol
В нашем обществе, где диплом становится началом успешной карьеры в любом направлении, многие пытаются найти максимально быстрый и простой путь получения образования. Наличие официального документа сложно переоценить. Ведь диплом открывает дверь перед каждым человеком, который стремится вступить в профессиональное сообщество или продолжить обучение в высшем учебном заведении.
Мы предлагаем оперативно получить этот важный документ. Вы имеете возможность купить диплом нового или старого образца, и это становится удачным решением для всех, кто не смог закончить образование, утратил документ или хочет исправить плохие оценки. дипломы выпускаются аккуратно, с максимальным вниманием к мельчайшим нюансам, чтобы в результате получился продукт, 100% соответствующий оригиналу.
Плюсы данного подхода состоят не только в том, что можно оперативно получить свой диплом. Процесс организован удобно, с профессиональной поддержкой. От выбора нужного образца документа до консультации по заполнению личных данных и доставки в любой регион страны — все находится под абсолютным контролем качественных мастеров.
Таким образом, для всех, кто пытается найти максимально быстрый способ получить требуемый документ, наша компания может предложить отличное решение. Заказать диплом – это значит избежать долгого обучения и не теряя времени перейти к достижению личных целей, будь то поступление в ВУЗ или начало успешной карьеры.
http://karapuziki.0pk.me/viewtopic.php?id=7229#p91407
https://forum.zone-game.info/showthread.php?p=438833#post438833
http://ershov.mybb.ru/viewtopic.php?id=194#p3207
https://bbs.pku.edu.cn/v2/jump-to.php?url=https://russiany-diplomans.com
https://1abakan.ru/forum/showthread-64726/
В нашем обществе, где диплом – это начало удачной карьеры в любой отрасли, многие ищут максимально быстрый путь получения качественного образования. Важность наличия официального документа об образовании переоценить невозможно. Ведь диплом открывает двери перед каждым человеком, желающим начать профессиональную деятельность или учиться в любом ВУЗе.
В данном контексте наша компания предлагает быстро получить любой необходимый документ. Вы можете купить диплом, что становится отличным решением для всех, кто не смог закончить обучение, утратил документ или желает исправить плохие оценки. дипломы выпускаются с особой аккуратностью, вниманием ко всем нюансам. В итоге вы получите полностью оригинальный документ.
Преимущества данного подхода состоят не только в том, что можно максимально быстро получить свой диплом. Весь процесс организован просто и легко, с нашей поддержкой. От выбора подходящего образца документа до консультации по заполнению персональной информации и доставки в любой регион России — все будет находиться под полным контролем квалифицированных специалистов.
Всем, кто пытается найти максимально быстрый способ получения требуемого документа, наша компания предлагает отличное решение. Купить диплом – значит избежать длительного обучения и не теряя времени перейти к достижению собственных целей: к поступлению в университет или к началу трудовой карьеры.
http://diplomsagroups.com/kupit-diplom-vuza/kandidata-nauk.html
В нашем обществе, где диплом – это начало отличной карьеры в любом направлении, многие пытаются найти максимально быстрый путь получения качественного образования. Факт наличия документа об образовании сложно переоценить. Ведь именно диплом открывает дверь перед всеми, кто стремится вступить в сообщество профессионалов или учиться в высшем учебном заведении.
Мы предлагаем быстро получить этот важный документ. Вы сможете купить диплом старого или нового образца, и это становится удачным решением для всех, кто не смог завершить образование, потерял документ или хочет исправить свои оценки. диплом изготавливается аккуратно, с максимальным вниманием к мельчайшим элементам. На выходе вы получите 100% оригинальный документ.
Плюсы такого решения состоят не только в том, что можно оперативно получить свой диплом. Весь процесс организован комфортно, с профессиональной поддержкой. Начиная от выбора необходимого образца документа до консультаций по заполнению личной информации и доставки в любой регион России — все будет находиться под абсолютным контролем квалифицированных специалистов.
Всем, кто ищет оперативный способ получить требуемый документ, наша компания готова предложить выгодное решение. Приобрести диплом – это значит избежать продолжительного процесса обучения и не теряя времени переходить к достижению своих целей: к поступлению в ВУЗ или к началу успешной карьеры.
http://diplomsagroups.com/kupit-diplom-v-gorode/voronezh.html
I visited multiple websites except the audio feature
for audio songs present at this site is really excellent.
I don’t even know how I ended up here, but I thought this post was good.
I don’t know who you are but definitely you’re going to
a famous blogger if you are not already ;
) Cheers!
Thanks a lot for sharing this with all people you really
know what you are talking approximately! Bookmarked.
Please additionally talk over with my web site =). We may have a hyperlink trade
agreement between us
Hi there this is kinda of off topic but I was wanting to know if blogs use WYSIWYG editors or if you have to manually
code with HTML. I’m starting a blog soon but have no coding expertise
so I wanted to get guidance from someone with experience.
Any help would be greatly appreciated!
Thank you for sharing your info. I truly appreciate your efforts and I am waiting for your next post thank you once again.
Thanks for sharing your thoughts. I truly appreciate your
efforts and I am waiting for your next post thank you once again.
Unquestionably consider that that you stated.
Your favourite reason appeared to be on the net the simplest factor to understand of.
I say to you, I definitely get irked at the same time as folks consider worries that they plainly do not recognize about.
You managed to hit the nail upon the top and also defined out the entire thing without having
side effect , folks can take a signal. Will probably be again to get more.
Thank you
I know this if off topic but I’m looking into starting my own blog and was
curious what all is required to get set up? I’m assuming having a blog like yours
would cost a pretty penny? I’m not very web smart
so I’m not 100% positive. Any suggestions or advice would be greatly appreciated.
Thank you
For the reason that the admin of this web page is working, no question very
shortly it will be well-known, due to its feature contents.
Normally I don’t read article on blogs, however I wish to say that this
write-up very pressured me to check out and do it! Your writing style has been amazed me.
Thank you, very nice article.
This piece of writing is genuinely a fastidious one it
assists new the web visitors, who are wishing in favor of blogging.
What’s up mates, how is everything, and what you desire to say regarding this post, in my view its
truly awesome in support of me.
Please let me know if you’re looking for a article author for your
site. You have some really great articles and I feel I would
be a good asset. If you ever want to take some of the load off, I’d absolutely love to write some articles for your blog
in exchange for a link back to mine. Please send me an email if interested.
Thank you!
Hello excellent blog! Does running a blog like this take
a lot of work? I have absolutely no knowledge
of programming but I was hoping to start my own blog in the near future.
Anyways, should you have any suggestions or tips for
new blog owners please share. I understand this is off
subject however I simply wanted to ask. Thanks a lot!
Hello just wanted to give you a quick heads up and let you know a few
of the images aren’t loading correctly. I’m not sure why but
I think its a linking issue. I’ve tried it in two different
internet browsers and both show the same results.
I blog often and I really appreciate your information. The article has truly peaked my interest.
I will bookmark your blog and keep checking for new details about once a week.
I subscribed to your RSS feed as well.
Yes! Finally someone writes about fitspresso coffee reviews.
В нашем обществе, где диплом является началом удачной карьеры в любой сфере, многие ищут максимально быстрый путь получения качественного образования. Наличие официального документа переоценить невозможно. Ведь именно он открывает двери перед каждым человеком, который желает вступить в сообщество квалифицированных специалистов или учиться в высшем учебном заведении.
В данном контексте мы предлагаем оперативно получить этот важный документ. Вы сможете купить диплом, и это становится удачным решением для всех, кто не смог завершить обучение, потерял документ или хочет исправить плохие оценки. дипломы производятся аккуратно, с максимальным вниманием ко всем элементам. В итоге вы сможете получить 100% оригинальный документ.
Превосходство такого решения заключается не только в том, что вы быстро получите свой диплом. Процесс организован комфортно и легко, с профессиональной поддержкой. Начиная от выбора подходящего образца диплома до консультаций по заполнению личных данных и доставки по России — все находится под абсолютным контролем квалифицированных специалистов.
Для тех, кто ищет быстрый способ получить необходимый документ, наша услуга предлагает отличное решение. Купить диплом – это значит избежать долгого обучения и сразу переходить к достижению своих целей, будь то поступление в ВУЗ или начало карьеры.
http://domnazelenoy.mybb.ru/viewtopic.php?id=1937#p5177
https://etwinningonline.eba.gov.tr/author/elviraparks1/
https://courses.comet.ucar.edu/tag/index.php?tc=1&tag=%D0%9A%D1%80%D0%B8%D1%82%D0%B5%D1%80%D0%B8%D0%B8%20%D0%BF%D0%BE%D0%BB%D1%83%D1%87%D0%B5%D0%BD%D0%B8%D1%8F%20%D0%BA%D1%80%D0%B0%D1%81%D0%BD%D0%BE%D0%B3%D0%BE%20%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC%D0%B0%20%D0%B2%20%D0%A0%D0%BE%D1%81%D1%81%D0%B8%D0%B8
https://remontmix.ru/forum/viewtopic.php?t=8871
http://secondwife.mybb.ru/viewtopic.php?id=942#p240095
В нашем обществе, где диплом становится началом отличной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения качественного образования. Необходимость наличия официального документа об образовании переоценить просто невозможно. Ведь именно он открывает двери перед всеми, кто собирается вступить в сообщество профессионалов или продолжить обучение в ВУЗе.
Предлагаем оперативно получить этот важный документ. Вы имеете возможность купить диплом, что становится выгодным решением для всех, кто не смог завершить обучение, потерял документ или хочет исправить плохие оценки. дипломы выпускаются с особой тщательностью, вниманием ко всем нюансам, чтобы в итоге получился продукт, максимально соответствующий оригиналу.
Преимущества данного подхода заключаются не только в том, что можно оперативно получить диплом. Процесс организован комфортно, с профессиональной поддержкой. Начиная от выбора требуемого образца до грамотного заполнения личных данных и доставки по России — все будет находиться под абсолютным контролем квалифицированных специалистов.
Для тех, кто ищет быстрый и простой способ получения необходимого документа, наша компания предлагает выгодное решение. Приобрести диплом – значит избежать долгого процесса обучения и не теряя времени переходить к своим целям: к поступлению в ВУЗ или к началу трудовой карьеры.
http://diplomsagroups.com/kupit-diplom-v-gorode/kazan.html
Hi there to every one, the contents existing at this web page are actually awesome for people knowledge, well,
keep up the good work fellows.
В современном мире, где диплом становится началом удачной карьеры в любом направлении, многие пытаются найти максимально простой путь получения образования. Необходимость наличия официального документа об образовании трудно переоценить. Ведь диплом открывает двери перед людьми, стремящимися вступить в сообщество квалифицированных специалистов или учиться в университете.
В данном контексте наша компания предлагает очень быстро получить этот необходимый документ. Вы имеете возможность приобрести диплом старого или нового образца, что является выгодным решением для человека, который не смог завершить обучение, потерял документ или желает исправить свои оценки. дипломы изготавливаются аккуратно, с максимальным вниманием ко всем деталям. В итоге вы сможете получить документ, 100% соответствующий оригиналу.
Превосходство такого решения заключается не только в том, что можно максимально быстро получить диплом. Весь процесс организован комфортно и легко, с нашей поддержкой. От выбора требуемого образца до консультации по заполнению персональной информации и доставки по стране — все под абсолютным контролем квалифицированных мастеров.
Таким образом, для тех, кто ищет оперативный способ получить требуемый документ, наша компания предлагает выгодное решение. Заказать диплом – это значит избежать длительного обучения и сразу перейти к достижению собственных целей: к поступлению в ВУЗ или к началу успешной карьеры.
http://www.diplomsagroups.com
I’m gone to convey my little brother, that he should also pay a visit this webpage on regular basis to obtain updated from newest information.
https://www.diplom-zakaz.ru/
Hi Dear, are you genuinely visiting this website regularly, if so then you will absolutely get fastidious experience.
https://bisness-diplom.ru/
I believe what you wrote made a ton of sense. However, what about this? suppose you were to create a killer post title? I am not suggesting your information is not solid, however suppose you added something that makes people want more? I mean %BLOG_TITLE% is a little vanilla. You should peek at Yahoo’s front page and note how they create news titles to get viewers to open the links. You might try adding a video or a picture or two to get readers excited about everything’ve written. In my opinion, it might make your posts a little bit more interesting.
kupite-diplom0024.ru/
Hi there, of course this piece of writing is actually pleasant and I have learned lot of things from it about blogging. thanks.
https://www.sdiplom.ru/
Howdy very nice web site!! Man .. Beautiful .. Amazing .. I will bookmark your site and take the feeds additionally? I’m glad to seek out numerous helpful info right here in the publish, we need work out extra strategies in this regard, thank you for sharing. . . . . .
https://www.diplom-city24.ru/
Hello there, You’ve done a great job. I’ll definitely digg it and personally suggest to my friends. I am sure they will be benefited from this website.
https://www.diplom-city24.ru/
It’s an remarkable piece of writing in favor of all the internet visitors; they will take benefit from it I am sure.
Because the admin of this web site is working, no uncertainty very soon it will be well-known, due to its quality contents.
https://www.zakaz-na-diplom.ru/
I know this site gives quality dependent articles and extra
data, is there any other site which offers these kinds of data in quality?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/60043-mark-benford
https://www.obesityhelp.com/members/marke1964/about_me/
https://tubeteencam.com/user/posh1969/profile
https://rentry.org/f7oph8k6
https://anotepad.com/notes/bj78htnb
Greetings from Ohio! I’m bored to death at work so I decided to browse your site on my iphone
during lunch break. I love the information you present here
and can’t wait to take a look when I get home.
I’m surprised at how quick your blog loaded on my cell phone ..
I’m not even using WIFI, just 3G .. Anyhow, great site!
It’s awesome in support of me to have a site,
which is valuable designed for my experience. thanks admin
Hi there, i read your blog occasionally and i own a similar one and i
was just curious if you get a lot of spam comments? If so how do you prevent it, any
plugin or anything you can recommend? I get so much lately it’s driving me crazy so any help is very much appreciated.
We stumbled over here different web page and thought I may as well check things out.
I like what I see so now i’m following you. Look forward to checking out your
web page again.
WOW just what I was searching for. Came here by searching for glorycycles
Hey! Someone in my Myspace group shared this website with us so
I came to check it out. I’m definitely enjoying the information.
I’m book-marking and will be tweeting this to my followers!
Wonderful blog and wonderful style and design.
I’m extremely impressed with your writing skills and also with the
layout on your weblog. Is this a paid theme or did you modify it yourself?
Anyway keep up the nice quality writing, it’s rare to see a nice blog like this one these days.
I am in fact happy to glance at this webpage posts which includes
lots of useful information, thanks for providing such statistics.
hello there and thank you for your information – I have definitely picked up anything new from right here.
I did however expertise several technical points using this website, as
I experienced to reload the site lots of times previous to I could
get it to load properly. I had been wondering if your
web host is OK? Not that I am complaining, but slow loading
instances times will sometimes affect your placement in google and could damage your high-quality score if advertising and
marketing with Adwords. Well I’m adding this RSS to my email and can look out for much more of
your respective interesting content. Make sure you update this again very soon.
Have you ever considered writing an e-book or guest authoring on other
sites? I have a blog centered on the same
subjects you discuss and would love to have you share some
stories/information. I know my viewers would enjoy
your work. If you’re even remotely interested, feel free to send me an email.
I savor, cause I discovered exactly what I used to be having a look for.
You have ended my 4 day long hunt! God Bless you man. Have a nice day.
Bye
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195596
https://rentry.org/2uvcno5k
https://www.hentai-foundry.com/user/whity1992/profile
https://www.hentai-foundry.com/user/fogid1965195719671983/profile
https://www.haikudeck.com/presentations/S2bmoJ0OJx
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151651
https://wwoww1973.bandcamp.com/album/the-agreement-1
https://haveagood.holiday/users/351118
https://www.hentai-foundry.com/user/tikra1990/profile
https://imageevent.com/larison1975
В нашем обществе, где диплом становится началом успешной карьеры в любом направлении, многие стараются найти максимально быстрый путь получения качественного образования. Наличие официального документа об образовании трудно переоценить. Ведь диплом открывает дверь перед любым человеком, желающим вступить в сообщество профессионалов или учиться в высшем учебном заведении.
В данном контексте мы предлагаем оперативно получить этот необходимый документ. Вы сможете заказать диплом старого или нового образца, что является отличным решением для всех, кто не смог завершить образование, потерял документ или желает исправить свои оценки. Все дипломы выпускаются аккуратно, с особым вниманием ко всем элементам. В итоге вы получите продукт, 100% соответствующий оригиналу.
Плюсы подобного решения заключаются не только в том, что можно быстро получить свой диплом. Весь процесс организовывается удобно, с нашей поддержкой. Начав от выбора нужного образца диплома до консультации по заполнению персональной информации и доставки по стране — все под полным контролем опытных мастеров.
Всем, кто пытается найти быстрый и простой способ получить необходимый документ, наша компания предлагает выгодное решение. Приобрести диплом – значит избежать долгого обучения и не теряя времени переходить к достижению собственных целей: к поступлению в университет или к началу удачной карьеры.
http://ya.forum2.net/viewtopic.php?id=9375#p18795
https://open.mit.edu/profile/01HY8A5V2QPVH62G1NXVF5BGA0/
http://peru.mybb.ru/viewtopic.php?id=289#p4090
http://detki.forum.cool/viewtopic.php?id=2866#p3475
https://doodleordie.com/profile/josearnold2
I loved as much as you will receive carried out right here. The sketch is attractive, your authored subject matter stylish. nonetheless, you command get bought an impatience over that you wish be delivering the following. unwell unquestionably come more formerly again since exactly the same nearly very often inside case you shield this increase.
https://www.institute-diplom.ru/
В наше время, когда диплом является началом удачной карьеры в любой области, многие ищут максимально простой путь получения качественного образования. Факт наличия документа об образовании сложно переоценить. Ведь диплом открывает дверь перед любым человеком, желающим вступить в сообщество профессионалов или продолжить обучение в высшем учебном заведении.
Мы предлагаем быстро получить любой необходимый документ. Вы сможете купить диплом нового или старого образца, и это является удачным решением для человека, который не смог закончить образование, потерял документ или хочет исправить свои оценки. Каждый диплом изготавливается с особой тщательностью, вниманием ко всем нюансам. В результате вы получите документ, полностью соответствующий оригиналу.
Преимущество такого решения состоит не только в том, что можно быстро получить диплом. Весь процесс организован удобно, с нашей поддержкой. От выбора подходящего образца диплома до грамотного заполнения личной информации и доставки по стране — все под абсолютным контролем качественных специалистов.
Для тех, кто пытается найти быстрый и простой способ получения требуемого документа, наша компания предлагает выгодное решение. Купить диплом – это значит избежать долгого процесса обучения и не теряя времени переходить к достижению собственных целей, будь то поступление в ВУЗ или начало удачной карьеры.
http://fkvrn.webtalk.ru/viewtopic.php?id=696#p21973
https://www.intensedebate.com/profiles/josearnold2
http://fast4ride.mybb.ru/viewtopic.php?id=135#p136
https://rostokaluga.bbnow.ru/viewtopic.php?id=1145#p13844
http://compmaster.mybb.ru/viewtopic.php?id=11689#p19309
Hello, i believe that i noticed you visited my website thus i came to return the desire?.I am trying to find things to improve my web site!I suppose its ok to make use of a few of your
ideas!!
It’s going to be finish of mine day, however before end I am reading this wonderful article
to increase my know-how.
Hi, yup this post is genuinely good and I have learned lot of things from it about blogging. thanks.
@@G@@
https://www.0332.ua/list/463064
Currently it looks like Movable Type is the top blogging platform out there right now.
(from what I’ve read) Is that what you’re using on your
blog?
Hi to all, the contents existing at this site are in fact remarkable for people experience, well, keep up the good work fellows.
Also visit my web site: seoul weather
What a fascinating article! Your protocol for tracking brain fibers with empirical MRI is truly impressive. The detailed instructions and screenshots provided make the process clear and accessible, even for those who are not experts in the field. The precision with which you’ve described each step, from creating coronal seed regions to verifying ROI and ROA regions, is exemplary. This guide is a valuable resource for anyone interested in brain tractography. Bravo for this meticulous and well-structured work! I look forward to future updates and improvements.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://unix1962.bandcamp.com/album/the-voices-in-my-head-part-5
https://www.haikudeck.com/presentations/uxiq8hmYGm
https://tubeteencam.com/user/koksna1996/profile
https://www.quia.com/profiles/chleighton
http://www.nfomedia.com/profile?uid=rOjQajF
What’s up, its pleasant paragraph on the topic of media print, we all know media is a enormous source
of information.
Heya i am for the first time here. I found this board and I find
It really helpful & it helped me out much. I hope to present one
thing back and help others such as you aided me.
you’re actually a just right webmaster. The site loading speed
is incredible. It seems that you’re doing any unique trick.
In addition, The contents are masterwork.
you have done a excellent activity on this matter!
I believe this is one of the so much vital info for me.
And i am satisfied studying your article. But wanna remark on some general issues, The website style is perfect, the articles is
in reality nice : D. Just right activity, cheers
For the reason that the admin of this site is working, no question very shortly it will be renowned, due to its feature contents.
ipoboard.ru/index.html#!/IPOboardВ
iawbs.com/home.php?mod=space&uid=792962&do=profile&from=spaceВ
http://www.vaca-ps.org/blogs/64421/%D0%9D%D0%B0%D0%B4%D0%B5%D0%B6%D0%BD%D1%8B%D0%B9-%D0%B8-%D0%BA%D0%BE%D0%BD%D1%84%D0%B8%D0%B4%D0%B5%D0%BD%D1%86%D0%B8%D0%B0%D0%BB%D1%8C%D0%BD%D1%8B%D0%B9-%D1%81%D0%B5%D1%80%D0%B2%D0%B8%D1%81-%D0%BF%D0%BE%D0%BA%D1%83%D0%BF%D0%BA%D0%B8-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC%D0%BE%D0%B2-%D0%BE%D0%BD%D0%BB%D0%B0%D0%B9%D0%BD?lang=tr_trВ
http://www.paladiny.ru/forummess.dwar.php?TopicID=26048&Offset=0В
cbt-v.com.ua/catalog/allВ
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://devil461956.bandcamp.com/album/the-hunters-become-the-hunted-part-2
https://anotepad.com/notes/6qb4rpb9
https://www.quia.com/profiles/bethwebster
https://rentry.org/7ek5pyx7
https://rentry.org/7bpmaveo
Greetings from California! I’m bored to death at work so I decided to browse your site on my iphone during lunch break.
I really like the information you present here and can’t wait to take a look
when I get home. I’m amazed at how quick your blog loaded on my cell phone ..
I’m not even using WIFI, just 3G .. Anyhow, good site!
WOW just what I was searching for. Came here by searching for %meta_keyword%
http://www.vespaclubnapoli.it/forum/viewthread.php?forum_id=5&thread_id=25853В
tandem585.ru/В
аржаники.СЂС„/memberlist.php?sk=a&sd=a&first_char=r&start=200В
amisite.ru/phpBB2/memberlist.php?mode=joined&order=ASC&start=6850В
blog.libero.it/wp/interesting/page/3/В
Great delivery. Great arguments. Keep up the great work.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/denzet1969/profile
https://chyoa.com/user/ijl1992
https://danceman1986.diary.ru/
http://www.nfomedia.com/profile?uid=rOiYYiH
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195914
Hi, I wish for to subscribe for this blog to take latest updates, thus where can i do it please help.
http://www.transportpath.ru/palon-591.html
http://msd.com.ua/poiskovaya-vydacha/unikalnye-vidy-stroitelnyx-materialov/
http://babygirlboyname.com/detail/Galina
http://profiapple.ru/instrukcii/
http://i-cross.ru/main/slovo/begemot
Thanks , I have just been searching for info approximately this topic for a while and yours is
the greatest I have found out till now. But, what in regards to the conclusion?
Are you sure concerning the source?
В современном мире, где диплом – это начало успешной карьеры в любом направлении, многие стараются найти максимально быстрый и простой путь получения качественного образования. Наличие документа об образовании переоценить попросту невозможно. Ведь именно диплом открывает дверь перед людьми, стремящимися начать профессиональную деятельность или продолжить обучение в ВУЗе.
В данном контексте мы предлагаем очень быстро получить этот важный документ. Вы можете купить диплом старого или нового образца, и это становится удачным решением для человека, который не смог закончить обучение, потерял документ или хочет исправить плохие оценки. диплом изготавливается с особой тщательностью, вниманием к мельчайшим деталям, чтобы в итоге получился полностью оригинальный документ.
Преимущества данного подхода состоят не только в том, что можно оперативно получить диплом. Весь процесс организован комфортно, с профессиональной поддержкой. Начав от выбора подходящего образца диплома до консультации по заполнению персональных данных и доставки по России — все под полным контролем наших специалистов.
Для всех, кто ищет оперативный способ получить требуемый документ, наша компания предлагает отличное решение. Заказать диплом – это значит избежать продолжительного обучения и не теряя времени перейти к своим целям: к поступлению в университет или к началу трудовой карьеры.
http://mirindavietnam.com
http://southcoorissa.com
http://attivalamemoria.eu
http://baotanglichsuvn.com
http://nubentos.com
Hello, its pleasant article concerning media print,
we all know media is a wonderful source of facts.
I’m not sure where you’re getting your info, but great topic.
I needs to spend some time learning much more or understanding more.
Thanks for fantastic information I was looking for this information for my mission.
Do you have a spam problem on this website; I also am a blogger, and I was wanting to know your situation; many of us have created some nice methods and we are looking to trade techniques with other folks, please shoot me an email if interested.
yaoisennari.ekafe.ru/viewtopic.php?f=169&t=1669&start=0&view=printВ
stroyzapadnoe.ru/%D0%B3%D0%BE%D1%82%D0%BE%D0%B2%D1%8B%D0%B5-%D0%BE%D0%B1%D1%8A%D0%B5%D0%BA%D1%82%D1%8B/nggallery/thumbnailsВ
board.46info.ru/regions/В
telegra.ph/Pokupka-diploma-SHag-v-budushchee-bez-usilij-05-14В
forumaniga.me/member/414-uf_23829059/visitormessage/13030-%D0%BF%D1%83%D0%B1%D0%BB%D0%B8%D1%87%D0%BD%D0%BE%D0%B5-%D1%81%D0%BE%D0%BE%D0%B1%D1%89%D0%B5%D0%BD%D0%B8%D0%B5-%D0%BE%D1%82-uf_23829059В
Hi there all, here every one is sharing these familiarity, so it’s good to read this blog,
and I used to go to see this website every day.
Great goods from you, man. I have understand your stuff previous to
and you’re just extremely magnificent. I actually like what
you have acquired here, really like what you are stating and the way
in which you say it. You make it enjoyable and you
still care for to keep it wise. I can’t wait to read
much more from you. This is really a terrific site.
Hello! This is kind of off topic but I need some help
from an established blog. Is it tough to set up your
own blog? I’m not very techincal but I can figure things out pretty quick.
I’m thinking about creating my own but I’m not sure where to begin. Do you have any ideas or suggestions?
Many thanks
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/gfndy6yb
https://www.quia.com/profiles/bethwebster
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195618
https://cannabis.net/user/151073
https://www.haikudeck.com/presentations/20ft1LAXk3
http://parimatch-zerkalo.info
I am regular visitor, how are you everybody? This article posted at this website is truly nice.
That is very attention-grabbing, You’re an overly skilled blogger.
I’ve joined your feed and look ahead to searching for more of your fantastic post.
Also, I’ve shared your web site in my social networks
I blog frequently and I genuinely thank you for your content. Your article has really peaked my interest. I’m going to take a note of your site and keep checking for new details about once per week. I opted in for your Feed too.
iclicky.com/index.php?do=/public/user/blogs/name_Alanpoe/page_5/В
http://www.hot166.com/blogs/2763/Why-is-the-popularity-of-universities-decreasing-all-the-timeВ
naswinas.com/blogs/740/Why-is-the-popularity-of-universities-decreasing-todayВ
mboumukvilino.ru/antiterroristicheskaya-bezopasnost/normativnye-pravovye-akty-rossiyskoy-federacii-i-respubliki-krym/В
parenvarmii.ru/topic4065.html?view=previousВ
You need to take part in a contest for one of the greatest
blogs on the internet. I most certainly will highly recommend this blog!
Hello there I am so grateful I found your site, I really found you by mistake, while I was browsing on Bing for something
else, Regardless I am here now and would just like to
say kudos for a tremendous post and a all round exciting blog
(I also love the theme/design), I don’t have time to browse it all
at the moment but I have book-marked it and also added your RSS feeds, so when I have time I
will be back to read more, Please do keep up the excellent job.
Hello! Do you know if they make any plugins to assist with
SEO? I’m trying to get my blog to rank for some
targeted keywords but I’m not seeing very good success.
If you know of any please share. Kudos!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195560
https://rentry.org/yubsrc7m
http://www.nfomedia.com/profile?uid=rOiYfkH
https://spirit019871998.diary.ru/
https://tubeteencam.com/user/elenangel1957/profile
Helpful information. Lucky me I found your site
unintentionally, and I’m stunned why this coincidence didn’t took place
in advance! I bookmarked it.
geinoutime.com
Hongzhi 황제는 말했지만 Fang Jifan을 바라 볼 수밖에 없었습니다.
I have read so many posts regarding the blogger lovers but this article is really a nice article, keep it up.
Hi there every one, here every one is sharing such familiarity,
thus it’s fastidious to read this weblog, and I used to pay a quick visit this
web site everyday.
Nice blog right here! Also your web site so much up very fast!
What host are you the use of? Can I am getting your associate
hyperlink for your host? I want my web site loaded up as fast as yours lol
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/fkcv2cdk
https://dartoos1976.diary.ru/
https://rentry.org/sbqfq3po
https://www.hentai-foundry.com/user/mirgola1980/profile
https://www.quia.com/profiles/jakma
Your style is really unique in comparison to other people I’ve read stuff from.
Many thanks for posting when you have the opportunity, Guess I’ll just bookmark this web site.
Hi there Dear, are you truly visiting this web page regularly, if so after that you will without doubt take fastidious know-how.
angelteam.uv.ro/profile.php?mode=viewprofile&u=207852В
mse-online.ru/novosti/vybor-luchshej-shkoly-dlya-vashego-rebenka.htmlВ
girbir.com/blogs/13038/Where-can-I-buy-a-diploma-or-certificate-at-aВ
foro.turismo.org/post281274.htmlВ
ttdinhduong.org/ttdd/tin-tuc/tin-chuyen-mon/679-Thong-bao-chieu-sinh-khoa-dao-tao-ddls.aspxВ
What’s up, the whole thing is going perfectly here and ofcourse every one is sharing
facts, that’s genuinely good, keep up writing.
Hey there would you mind letting me know which web host you’re using?
I’ve loaded your blog in 3 different browsers and
I must say this blog loads a lot quicker then most. Can you recommend a good internet
hosting provider at a reasonable price? Thank you,
I appreciate it!
I’d like to thank you for the efforts you have put in penning this website.
I really hope to view the same high-grade content by you later on as well.
In fact, your creative writing abilities has inspired me
to get my very own blog now 😉
Pretty! This has been an incredibly wonderful post.
Thank you for providing this information.
My brother recommended I would possibly like this blog.
He used to be entirely right. This post truly made my day.
You cann’t believe simply how so much time I had spent for this information! Thanks!
whoah this weblog is great i love studying your articles.
Keep up the good work! You know, a lot of individuals are looking round for this info, you
can aid them greatly.
Hi! I just wanted to ask if you ever have any issues with hackers?
My last blog (wordpress) was hacked and I ended up losing a few months of
hard work due to no data backup. Do you have any methods to stop hackers?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151401
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196255
https://www.haikudeck.com/presentations/qAKfQjD1wH
https://cannabis.net/user/150862
https://www.hentai-foundry.com/user/bodyanvk1995/profile
Actually when someone doesn’t understand afterward its up to other people that they will help, so here it occurs.
В нашем мире, где диплом – это начало отличной карьеры в любой сфере, многие пытаются найти максимально быстрый путь получения качественного образования. Важность наличия официального документа сложно переоценить. Ведь диплом открывает дверь перед всеми, кто собирается вступить в сообщество квалифицированных специалистов или учиться в ВУЗе.
Предлагаем максимально быстро получить этот важный документ. Вы можете купить диплом нового или старого образца, что становится выгодным решением для человека, который не смог закончить обучение или утратил документ. диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам, чтобы на выходе получился продукт, полностью соответствующий оригиналу.
Преимущество данного решения заключается не только в том, что вы сможете максимально быстро получить диплом. Процесс организован комфортно, с профессиональной поддержкой. Начав от выбора нужного образца диплома до правильного заполнения персональных данных и доставки по стране — все находится под полным контролем квалифицированных специалистов.
В результате, для тех, кто ищет оперативный способ получить требуемый документ, наша компания предлагает отличное решение. Заказать диплом – значит избежать продолжительного обучения и сразу перейти к своим целям: к поступлению в ВУЗ или к началу удачной карьеры.
http://a-line-dress.com
http://maixepanphat.vn
http://seoanalitika7.site
http://jamesrosenbarger.com
http://vat-calculator.uk
На сегодняшний день, когда диплом – это начало отличной карьеры в любой отрасли, многие стараются найти максимально простой путь получения образования. Факт наличия официального документа переоценить невозможно. Ведь именно он открывает дверь перед людьми, желающими вступить в сообщество профессионалов или учиться в высшем учебном заведении.
В данном контексте мы предлагаем очень быстро получить этот необходимый документ. Вы можете заказать диплом нового или старого образца, что будет отличным решением для человека, который не смог закончить образование, утратил документ или хочет исправить свои оценки. диплом изготавливается аккуратно, с максимальным вниманием ко всем нюансам. В результате вы сможете получить 100% оригинальный документ.
Преимущество данного решения заключается не только в том, что вы оперативно получите диплом. Процесс организован комфортно, с профессиональной поддержкой. Начиная от выбора нужного образца до точного заполнения персональной информации и доставки в любой регион России — все находится под абсолютным контролем наших специалистов.
Для тех, кто ищет быстрый и простой способ получить требуемый документ, наша компания готова предложить отличное решение. Заказать диплом – значит избежать длительного обучения и сразу переходить к достижению личных целей: к поступлению в университет или к началу удачной карьеры.
http://mininform.ru
http://roomblock.com
http://ravolutionfestival.com
http://amazonia-tour.ru
http://effective-herbal-cure.com
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196867
https://tubeteencam.com/user/caesarus1978/profile
https://haveagood.holiday/users/351967
https://chyoa.com/user/hghhshsgh1959
https://okwave.jp/profile/u3115296.html
Excellent way of explaining, and nice paragraph
to take information regarding my presentation topic, which i am going
to convey in university.
Just wish to say your article is as astounding. The clarity in your submit is just excellent and i can think you are a professional on this subject. Fine with your permission allow me to grasp your RSS feed to stay up to date with coming near near post. Thanks one million and please continue the rewarding work.
http://frsvo.ru/forum/profile.php?id=6233
http://aikentools.ru/katalog/generatory/benzinovyy-generator-vitals-master-est-5-8b-14551.html
http://sdorov.ru/metodiki/gomeopatiya/
http://www.irc71.ru/index.php/jomsocial/groups/viewbulletin/45-za-kakuyu-tsenu-segodnya-vozmozhno-kupit-khoroshego-kachestva-diplom?groupid=47
http://hramchaik.ru/articles/1940-sozdaem-prekrasnoe-svoimi-rukami-sovety-ot-kompanii-inter-atletika.html
What’s up it’s me, I am also visiting this website regularly, this web page is truly pleasant and the people are truly
sharing fastidious thoughts.
Hello! Someone in my Myspace group shared this site with us so
I came to take a look. I’m definitely loving the information. I’m bookmarking and will be tweeting this to my followers!
Terrific blog and fantastic design and style.
When someone writes an article he/she retains the image of a user in his/her mind that
how a user can understand it. So that’s why this piece of writing is amazing.
Thanks!
situs bokep
situs xnxx
Hello! Do you know if they make any plugins to assist with Search Engine Optimization? I’m trying to get my blog to rank for some targeted keywords but I’m not seeing very good results.
If you know of any please share. Appreciate it!
Hi there to every body, it’s my first go to see of this web site; this web site includes amazing and in fact excellent material in favor of visitors.
http://www.reisezielforum.de/forum-f5/l942g-t61855.htmlВ
http://www.modern-constructions.org/blogs/44177/%D0%94%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC%D1%8B-%D0%B2%D1%8B%D1%81%D0%BE%D1%87%D0%B0%D0%B9%D1%88%D0%B5%D0%B3%D0%BE-%D0%BA%D0%B0%D1%87%D0%B5%D1%81%D1%82%D0%B2%D0%B0-%D0%B2%D0%B0%D1%88-%D1%88%D0%B0%D0%B3-%D0%BA-%D0%BF%D1%80%D0%BE%D1%84%D0%B5%D1%81%D1%81%D0%B8%D0%BE%D0%BD%D0%B0%D0%BB%D1%8C%D0%BD%D0%BE%D0%BC%D1%83-%D1%80%D0%BE%D1%81%D1%82%D1%83В
uks-zhilstroy.ru/gallerys/lim-foto/foto-lerm-25?start=50В
mse-online.ru/novosti/vybor-luchshej-shkoly-dlya-vashego-rebenka.htmlВ
http://www.rolandus.org/forum/profile.php?mode=viewprofile&u=25280В
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/unte1988/profile
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195416
https://rentry.org/zmrpquun
https://cannabis.net/user/152010
https://www.quia.com/profiles/as308harrison
http://gs-operation.ru
m2crowd es una página web famosa en todo el mundo que
ofrece vídeos xx en HD y de larga duración, y lo hacen gratis.
Hello to every body, it’s my first pay a visit of this blog; this website includes awesome and actually fine data in favor of readers.
hello!,I love your writing very so much! percentage we keep up a correspondence more approximately your article on AOL?
I require a specialist in this house to unravel my problem.
Maybe that is you! Having a look forward to see you.
Attractive section of content. I just stumbled upon your weblog and in accession capital to assert that I acquire actually enjoyed account your blog
posts. Anyway I’ll be subscribing to your augment and even I achievement
you access consistently rapidly.
WOW just what I was looking for. Came here by searching for prepaiddigitalsolutions
Thanks for your marvelous posting! I seriously enjoyed reading it, you will be a great author.I will
ensure that I bookmark your blog and will come back in the future.
I want to encourage you continue your great work,
have a nice day!
WOW just what I was searching for. Came here by searching for %keyword%
http://www.ozelenitel-armavir.ru/page_proizvodstvo-tsvetochnoj-rassady.html
http://samovod.ru/content/articles/65354/
http://kvakva55.ru/atletiko-interesen-dzhaka.html
http://angelkld.ru/lenta-20.html
http://tvoygarazh.ru/fundament/svajnye-fundamenty.html
This paragraph is actually a good one it assists new internet
visitors, who are wishing for blogging.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/Mdj2KHcK4Q
https://rentry.org/g6h7bv35
https://anotepad.com/notes/pn58kg32
https://okwave.jp/profile/u3116813.html
https://chyoa.com/user/hellanna1953
My developer is trying to persuade me to move to .net from PHP.
I have always disliked the idea because of the expenses.
But he’s tryiong none the less. I’ve been using Movable-type on a
variety of websites for about a year and am anxious about switching to another
platform. I have heard very good things about blogengine.net.
Is there a way I can transfer all my wordpress
posts into it? Any kind of help would be greatly appreciated!
Very good article. I definitely appreciate this website.
Thanks!
If some one wishes expert view on the topic
of running a blog then i advise him/her to go to see this webpage, Keep up the good work.
Hi there this is kind of of off topic but I was wondering if blogs use WYSIWYG editors or if you have to
manually code with HTML. I’m starting a blog soon but have no coding experience so I wanted to get guidance from someone with experience.
Any help would be greatly appreciated!
It’s not my first time to pay a quick visit this web site, i am browsing this
site dailly and take fastidious information from here all the time.
Nice answers in return of this matter with genuine
arguments and explaining all regarding that.
I like the helpful info you provide in your articles.
I will bookmark your blog and check again here regularly. I am quite certain I’ll learn many new stuff right here!
Best of luck for the next!
Right here is the perfect website for everyone who really wants to find out about this
topic. You understand so much its almost hard to argue with you (not that I really
would want to…HaHa). You certainly put a new spin on a subject which has been written about for many years.
Excellent stuff, just wonderful!
This website was… how do you say it? Relevant!! Finally I have found something which
helped me. Thank you!
With havin so much written content do you ever run into any
issues of plagorism or copyright violation? My website has a
lot of exclusive content I’ve either written myself or outsourced but it appears a lot of it is popping it up all
over the internet without my permission. Do you know any methods to help prevent content
from being stolen? I’d definitely appreciate it.
В нашем мире, где диплом становится началом удачной карьеры в любой сфере, многие ищут максимально простой путь получения образования. Факт наличия официального документа об образовании трудно переоценить. Ведь именно он открывает двери перед всеми, кто желает вступить в профессиональное сообщество или учиться в университете.
В данном контексте наша компания предлагает оперативно получить этот необходимый документ. Вы имеете возможность купить диплом, что является выгодным решением для всех, кто не смог завершить образование или потерял документ. диплом изготавливается аккуратно, с особым вниманием ко всем деталям. В итоге вы сможете получить 100% оригинальный документ.
Преимущества данного решения заключаются не только в том, что можно оперативно получить диплом. Весь процесс организовывается удобно, с нашей поддержкой. Начиная от выбора подходящего образца до консультаций по заполнению персональной информации и доставки в любой регион страны — все под абсолютным контролем качественных мастеров.
Всем, кто пытается найти максимально быстрый способ получения требуемого документа, наша компания предлагает выгодное решение. Приобрести диплом – это значит избежать длительного процесса обучения и сразу переходить к своим целям, будь то поступление в университет или старт успешной карьеры.
http://cocochampionship.com
http://looi.co
http://edvo-addictions.fr
http://alcado.com.vn
http://tonylandis.com
I’m really impressed with your writing skills and also with the layout on your weblog.
Is this a paid theme or did you customize it yourself?
Anyway keep up the nice quality writing, it’s rare to see a great
blog like this one these days.
This is my first time go to see at here and i am really pleassant to read all at one place.
http://kvartal74.ru/lenta-23.html
http://vehicleskins.com/memberlist.php?mode=joined&order=ASC&start=241000
http://pedagoginfo.ru/2019/03/gotovye-multimedijnye-urok.html
http://урал-батыр.рф/search/
http://byte-kuzbass.ru/inetmagazvirt/category_00000092116/category_00000092117/category_00000104459/product_00000116923-detail
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/luhnik1977
http://www.nfomedia.com/profile?uid=rOjQfeE
https://ddreterte1999.diary.ru/
https://tubeteencam.com/user/niakria1983/profile
https://okwave.jp/profile/u3116281.html
What a information of un-ambiguity and preserveness of precious familiarity about unexpected emotions.
http://www.variable-stars.ru/db/search.html?page=2¬_mid=1172372&words=%F4%F0%E0%F3%ED%E3%EE%F4%E5%F0%EE%E2%FB %EB%E8%ED%E8%E8&where=text
http://searchprogram.ru/kak-sdelat-plan-v-vorde
http://opryanosti.ru/drugie-pryanosti/pankreatit-i-kurkuma.html
http://youreng.ru/bukva-e/
http://konteiners.ru/izotermicheskii-konteiner-insulated.html
I know this web page presents quality dependent content and additional information,
is there any other website which presents such stuff in quality?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/veronica-jackson
https://www.obesityhelp.com/members/rustyfoxy1998/about_me/
https://permacultureglobal.org/users/59882-elmi-fontanez
https://imageevent.com/jedipwnz1985
https://www.metal-archives.com/users/sobutilni19881994199619561955
Hello this is kind of of off topic but I was wondering if
blogs use WYSIWYG editors or if you have to manually code
with HTML. I’m starting a blog soon but have no coding skills so I
wanted to get guidance from someone with experience. Any help would be enormously appreciated!
На сегодняшний день, когда диплом является началом удачной карьеры в любом направлении, многие пытаются найти максимально быстрый путь получения качественного образования. Необходимость наличия официального документа переоценить попросту невозможно. Ведь диплом открывает дверь перед людьми, стремящимися вступить в сообщество профессиональных специалистов или продолжить обучение в высшем учебном заведении.
В данном контексте мы предлагаем оперативно получить любой необходимый документ. Вы сможете приобрести диплом нового или старого образца, и это является удачным решением для человека, который не смог завершить образование или потерял документ. дипломы производятся с особой аккуратностью, вниманием к мельчайшим деталям. В итоге вы сможете получить полностью оригинальный документ.
Превосходство данного решения состоит не только в том, что можно оперативно получить диплом. Процесс организовывается удобно, с профессиональной поддержкой. От выбора нужного образца до консультации по заполнению личных данных и доставки в любое место страны — все будет находиться под абсолютным контролем квалифицированных специалистов.
Всем, кто пытается найти быстрый и простой способ получить необходимый документ, наша компания предлагает выгодное решение. Купить диплом – это значит избежать долгого обучения и не теряя времени перейти к личным целям: к поступлению в университет или к началу успешной карьеры.
http://frankiepeach.com
http://grandart.fr
http://vampoadre.su
http://greenap.com.ua
http://absi.od.ua
В нашем мире, где диплом становится началом успешной карьеры в любом направлении, многие пытаются найти максимально быстрый путь получения образования. Необходимость наличия официального документа переоценить попросту невозможно. Ведь именно диплом открывает двери перед любым человеком, который желает начать профессиональную деятельность или учиться в университете.
В данном контексте наша компания предлагает быстро получить любой необходимый документ. Вы имеете возможность купить диплом, и это является удачным решением для всех, кто не смог завершить образование или утратил документ. Каждый диплом изготавливается аккуратно, с максимальным вниманием к мельчайшим деталям, чтобы в результате получился продукт, 100% соответствующий оригиналу.
Плюсы подобного подхода заключаются не только в том, что вы сможете быстро получить диплом. Весь процесс организован просто и легко, с профессиональной поддержкой. Начав от выбора подходящего образца до консультации по заполнению личной информации и доставки в любой регион страны — все под абсолютным контролем наших специалистов.
В итоге, для всех, кто ищет быстрый способ получения необходимого документа, наша услуга предлагает выгодное решение. Приобрести диплом – значит избежать длительного обучения и сразу переходить к достижению личных целей: к поступлению в университет или к началу трудовой карьеры.
http://gongkinhdep.com
http://ufml.fr
http://grim.in.ua
http://jawizeruk.pl
http://try-kolduna.com.ua
파워 오브 토르 메가웨이즈
사장은 비웃으며 속으로 말했다. 죽을 줄 알면서 왜 여기 있느냐?
Excellent post. I was checking constantly this blog and I’m impressed!
Extremely helpful information particularly the last part 🙂 I care for such information much.
I was seeking this particular info for a very long time.
Thank you and best of luck.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/rtvzvear
https://cannabis.net/user/152092
http://www.nfomedia.com/profile?uid=rOjQZjD
https://tubeteencam.com/user/vedenie21996/profile
https://www.hentai-foundry.com/user/mormitor1963/profile
bacheca annunci roma incontri massaggi lazio auto usate moto arredamento casa veicoli furgoni camper e caravan cucine usate offerte
di lavoro cellulari telefonia notebook portatili
Good day! I could have sworn I’ve been to this blog before but after reading through some of the post
I realized it’s new to me. Anyhow, I’m definitely delighted I found it
and I’ll be bookmarking and checking back often!
I’ll right away grasp your rss feed as I can’t find
your e-mail subscription link or e-newsletter service.
Do you’ve any? Kindly allow me realize so that I could subscribe.
Thanks.
Hey there! I just wanted to ask if you ever have any trouble
with hackers? My last blog (wordpress) was
hacked and I ended up losing months of hard work due
to no back up. Do you have any methods to prevent hackers?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151030
https://www.obesityhelp.com/members/terocona1986/about_me/
https://grilay1964.bandcamp.com/album/a-trip-back-home-never-to-forget
https://mioru1961.diary.ru/
https://okwave.jp/profile/u3116189.html
В современном мире, где аттестат является началом успешной карьеры в любой области, многие стараются найти максимально быстрый путь получения образования. Наличие документа об образовании сложно переоценить. Ведь диплом открывает двери перед всеми, кто стремится вступить в сообщество профессионалов или учиться в высшем учебном заведении.
В данном контексте наша компания предлагает максимально быстро получить этот важный документ. Вы имеете возможность купить аттестат старого или нового образца, и это будет отличным решением для всех, кто не смог закончить образование, утратил документ или хочет исправить плохие оценки. Аттестат изготавливается с особой аккуратностью, вниманием ко всем нюансам, чтобы на выходе получился 100% оригинальный документ.
Плюсы такого решения состоят не только в том, что вы сможете быстро получить аттестат. Процесс организован удобно, с профессиональной поддержкой. От выбора требуемого образца аттестата до консультации по заполнению личной информации и доставки в любое место России — все будет находиться под полным контролем квалифицированных специалистов.
В итоге, для тех, кто ищет быстрый и простой способ получить требуемый документ, наша компания предлагает отличное решение. Заказать аттестат – значит избежать долгого обучения и не теряя времени переходить к своим целям, будь то поступление в ВУЗ или начало успешной карьеры.
https://unsorted.wikisort.ru/page/Российская_академия_медицинских_наук
В нашем обществе, где диплом является началом успешной карьеры в любой области, многие ищут максимально быстрый путь получения качественного образования. Факт наличия официального документа переоценить попросту невозможно. Ведь диплом открывает двери перед любым человеком, который желает вступить в сообщество профессионалов или продолжить обучение в высшем учебном заведении.
В данном контексте наша компания предлагает максимально быстро получить этот важный документ. Вы сможете заказать диплом, и это будет выгодным решением для человека, который не смог завершить образование или утратил документ. диплом изготавливается аккуратно, с особым вниманием к мельчайшим нюансам, чтобы на выходе получился документ, полностью соответствующий оригиналу.
Превосходство такого решения состоит не только в том, что вы сможете быстро получить диплом. Весь процесс организован комфортно, с нашей поддержкой. От выбора подходящего образца документа до правильного заполнения персональных данных и доставки по стране — все под абсолютным контролем опытных мастеров.
Таким образом, для тех, кто хочет найти оперативный способ получить необходимый документ, наша компания предлагает отличное решение. Заказать диплом – это значит избежать длительного обучения и не теряя времени перейти к важным целям: к поступлению в университет или к началу трудовой карьеры.
http://asp-l.ru
http://vannes-spiruline.com
http://wingenuitysoftware.com
http://maixepanphat.vn
http://sutter-web.com
В нашем обществе, где диплом – это начало удачной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения образования. Факт наличия официального документа переоценить невозможно. Ведь диплом открывает двери перед любым человеком, желающим начать трудовую деятельность или учиться в ВУЗе.
В данном контексте наша компания предлагает быстро получить этот важный документ. Вы можете купить диплом нового или старого образца, что является удачным решением для человека, который не смог закончить обучение или утратил документ. дипломы выпускаются аккуратно, с максимальным вниманием ко всем нюансам, чтобы в итоге получился 100% оригинальный документ.
Плюсы подобного решения заключаются не только в том, что вы сможете быстро получить свой диплом. Процесс организовывается комфортно, с нашей поддержкой. От выбора требуемого образца диплома до консультации по заполнению персональной информации и доставки в любой регион России — все под полным контролем опытных мастеров.
Всем, кто ищет быстрый и простой способ получения необходимого документа, наша компания предлагает отличное решение. Приобрести диплом – значит избежать продолжительного процесса обучения и сразу переходить к достижению личных целей: к поступлению в университет или к началу успешной карьеры.
http://dmitriyandreev.ru
http://cityjeans.com.ua
http://philautarchie.net
http://lessigles.fr
http://bgv-blasenschwaeche.de
I was recommended this website by my cousin. I’m not sure whether
this post is written by him as nobody else know such detailed about my trouble.
You are incredible! Thanks!
Nice blog here! Also your site loads up very fast!
What web host are you using? Can I get your affiliate link to your host?
I wish my site loaded up as quickly as yours lol
My coder is trying to convince me to move to .net from PHP.
I have always disliked the idea because of the expenses.
But he’s tryiong none the less. I’ve been using Movable-type on a number of websites for about a year and am anxious
about switching to another platform. I have heard good things about blogengine.net.
Is there a way I can transfer all my wordpress content into it?
Any kind of help would be greatly appreciated!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/fifpbiof
https://www.quia.com/profiles/dasamuel210
https://www.hentai-foundry.com/user/gostfire1993/profile
https://chyoa.com/user/bloodream1971
http://www.nfomedia.com/profile?uid=rOiYbiB
Hi, after reading this awesome paragraph i am too cheerful to share my
knowledge here with mates.
I think this is among the most significant info for me.
And i’m glad reading your article. But want to
remark on few general things, The web site style is great,
the articles is really great : D. Good job, cheers
I do not know if it’s just me or if perhaps everybody else encountering problems with your website.
It looks like some of the text in your content are running
off the screen. Can someone else please comment and let me know if
this is happening to them as well? This might be a issue
with my browser because I’ve had this happen before.
Thanks
В нашем мире, где диплом является началом успешной карьеры в любой области, многие ищут максимально простой путь получения образования. Необходимость наличия официального документа об образовании сложно переоценить. Ведь диплом открывает двери перед каждым человеком, который стремится вступить в профессиональное сообщество или продолжить обучение в каком-либо университете.
В данном контексте мы предлагаем максимально быстро получить этот необходимый документ. Вы можете заказать диплом, и это будет удачным решением для всех, кто не смог закончить обучение, утратил документ или хочет исправить плохие оценки. Каждый диплом изготавливается аккуратно, с максимальным вниманием ко всем элементам, чтобы в итоге получился документ, 100% соответствующий оригиналу.
Превосходство такого подхода заключается не только в том, что вы сможете оперативно получить диплом. Весь процесс организовывается комфортно, с нашей поддержкой. От выбора нужного образца до точного заполнения персональных данных и доставки по стране — все под абсолютным контролем качественных мастеров.
Всем, кто пытается найти быстрый и простой способ получить необходимый документ, наша компания может предложить отличное решение. Купить диплом – это значит избежать длительного процесса обучения и не теряя времени перейти к достижению личных целей: к поступлению в университет или к началу успешной карьеры.
http://smotri.com.ua
http://befirst.com.ua
http://yakusha.com.ua
http://thepokiesgames.com
http://lovisribca.ru
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/copcar219991970
https://www.haikudeck.com/presentations/CF3Rtwcbbl
https://tubeteencam.com/user/expa131971/profile
https://rentry.org/if89mzwg
https://rentry.org/ktmynwcz
Рады приветствовать вас на вашем портале!
Компания XRumer Art предлагает свои профессиональные услуги СЕО продвижения.
Ваш портал, как можно отметить, еще только набирает обороты. Чтобы ускорить его рост, предлагаем наши услуги по внешней СЕО-оптимизации. У нас имеются недорогие и эффективные инструменты для СЕО-специалистов. У нас большой опыт, в арсенале имеются реальные рабочие кейсы – если интересует, предоставим по запросу.
Наша компания предлагает скидку 10% до конца месяца на все услуги.
Что мы готовы предложить:
– Размещаем трастовые ссылки (требуется каждому сайту) – цена от 1,5 до 5000 руб
– Размещаем 2500 безанкорных ссылок (рекомендуется всем сайтам) – 3.900 руб
– Прогон по 110 000 сайтам в RU.зоне (максимально полезно для сайтов) – 2900 руб
– 150 постов в VK про ваш сайт (поможет получить рекламу) – 3900 рублей
– Статьи про ваш сайт на 300 форумах (мощнейшая раскрутка веб-сайта) – 29.000 рублей
– СуперПостинг – это прогон на 3 миллиона ресурсов (мощное размещение для вашего сайта) – 39.900 р
– Рассылка сообщений по сайтам с помощью обратной связи – цена по договоренности, будет зависеть от объемов.
Обращайтесь с любыми вопросами, мы поможем.
Отчётность.
Оплата: Yoo.Деньги, Bitcoin, МИР, Visa, MasterCard…
Принимаем USDT
Телегрм: https://t.me/exrumer
Skype: Loves.Ltd
https://www.webfx.com/
https://xrumer.cc/
Hiya very cool website!! Man .. Beautiful .. Superb ..
I will bookmark your site and take the feeds additionally?
I am satisfied to find so many helpful info right here within the post,
we want work out more strategies on this regard, thank you
for sharing. . . . . .
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://daddys1997.diary.ru/
https://tubeteencam.com/user/liroiv1991/profile
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195416
https://haveagood.holiday/users/351461
https://fista1982.bandcamp.com/album/older-woman-in-coffee-shop-part-3
http://status-bankrotstvo.ru
AudioBook26.ru зовет более 10 000 аудиокниг в течение свой в доску нашей электронной библиотеке. Ваша милость сможете много наслышаться о ком/чем их ясно сверху сайте чи установить приложение для офлайн-прослушивания
https://audiobook26.ru/
Wow, superb weblog layout! How lengthy have you ever been blogging for? you make blogging glance easy. The overall look of your web site is excellent, as smartly as the content material!
dataload.com/forum/memberlist.php?mode=joined&order=ASC&start=16850%C3%82%C2%A0В
http://www.sonyericsson-championships.com/es/news/story/370.aspВ
smeda.ru/uslugi/anons/anons_7.htmlВ
tnrevergreen.com.vn/dien-tich-tim-tuong-la-gi-mua-nha-nen-dung-cach-tinh-dien-tich-nao/В
symphonyofthemagneticnorth.com/_404.htmlВ
This post is worth everyone’s attention. When can I find out more?
tube.tunanno.com/profile.php?u=usyhymejaВ
batdongsanmiennam.vn/mnre-tich-cuc-day-manh-kenh-ban-hang-truc-tuyen/В
worldavtonew.ru/page/3В
http://www.bfauk.com/bfa_cms/userВ
elibot.gg/commandsВ
I used to be able to find good advice from your blog posts.
http://siamtovar.us/ips_3_4/index.php?app=forums&module=post§ion=post&do=reply_post&f=2&t=144&qpid=1800
http://unforma.ru/rybalka/rybopromyshlennikam-razyasnyat-tonkosti-raboty-s-erzh.html
http://auto-pyshme.ru/lenta-12.html
http://pechemivarim.ru/salat-iz-baklazhanov-gribnoj/
http://forum.doctorulmeu.md/viewtopic.php?t=72350&start=10530
Hi there! This post couldn’t be written any better! Reading through this post reminds me of my old
room mate! He always kept chatting about this. I will forward
this post to him. Fairly certain he will have
a good read. Thanks for sharing!
I have read several good stuff here. Certainly worth bookmarking for revisiting.
I surprise how so much attempt you place to make this type
of fantastic informative website.
whoah this blog is excellent i like reading your posts. Stay up the good work! You know, lots of persons are looking round for this information, you can help them greatly.
http://abyssian.ru/fanfik-krossover-perekrestki-sudeb/
http://www.artem-energo.ru/forums.php?m=posts&p=16434
http://chillspot1.com/friends/premiumsdiploms31
http://familylevel.com/blogs/26/What-do-you-need-to-pay-attention-to-when-buying
http://www.biofinder.ru/bfins-590-1.html
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/350040
https://rentry.org/vxykp6gd
https://www.haikudeck.com/presentations/rvKZdGwc7o
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196102
https://launchpad.net/~demanmax19881
В нашем мире, где аттестат является началом отличной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения качественного образования. Факт наличия официального документа сложно переоценить. Ведь диплом открывает дверь перед каждым человеком, желающим вступить в профессиональное сообщество или продолжить обучение в ВУЗе.
Мы предлагаем оперативно получить этот необходимый документ. Вы можете приобрести аттестат старого или нового образца, и это является удачным решением для всех, кто не смог завершить образование, потерял документ или желает исправить свои оценки. Аттестат изготавливается с особой тщательностью, вниманием к мельчайшим деталям. В итоге вы получите 100% оригинальный документ.
Преимущество данного решения состоит не только в том, что можно максимально быстро получить свой аттестат. Весь процесс организован просто и легко, с нашей поддержкой. Начав от выбора требуемого образца документа до консультаций по заполнению личных данных и доставки по России — все под полным контролем опытных мастеров.
Всем, кто ищет быстрый и простой способ получения требуемого документа, наша компания может предложить отличное решение. Заказать аттестат – значит избежать длительного обучения и сразу переходить к достижению личных целей, будь то поступление в ВУЗ или старт профессиональной карьеры.
http://forum.drustvogil-galad.si/index.php?topic=10793.0
This post will assist the internet visitors for creating new website or even a
blog from start to end.
Excellent weblog here! Also your site so much up very fast!
What host are you the use of? Can I get your associate link
on your host? I desire my site loaded up as fast as yours lol
Предлагаем вам пройти консультацию (аудит) по усилению продаж также доходы в вашем бизнесе. Формат аудита: личная встреча или конференция по скайпу. Делая верные, но обыкновенные действия, прибыль от ВАШЕГО коммерциала можно увеличить в много раз. В нашем багаже более 100 проверенных практических вариантов увеличения торгов а также доходов. В зависимости от вашего бизнеса разработаем для вас наиболее наилучшие и начнем шаг за шагом претворять в жизнь.
-https://interestbook.ru/
Hi! I know this is kinda off topic nevertheless I’d figured I’d ask.
Would you be interested in exchanging links or maybe guest writing a blog post or vice-versa?
My blog goes over a lot of the same subjects as yours and I believe we could greatly benefit from each other.
If you happen to be interested feel free to shoot me
an e-mail. I look forward to hearing from you!
Terrific blog by the way!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/sonbka1953
https://www.quia.com/profiles/je219cole
https://www.haikudeck.com/presentations/TwqLqzGzf8
https://www.hentai-foundry.com/user/ajer1984/profile
https://permacultureglobal.org/users/60090-luis-navarro
Thankfulness to my father who shared with me about this webpage, this web site is actually remarkable.
Fantastic post however I was wondering if you could write a litte more on this topic? I’d be very thankful if you could elaborate a little bit more. Bless you!
mystroycenter.ru/page/14В
86hm.ru/forum/flame/?topic_id=24930В
ezoinfo.ru/Narodnay_medizina/Saving_power/Saving_power_9.phpВ
http://www.matchfishing.ru/forumtest/1/member.php?u=23110&tab=activitystreamВ
regullife.ru/index.php?links_exchange=yes&show_all=yes&page=40В
Good post. I learn something new and challenging on blogs I stumbleupon on a daily basis.
It will always be exciting to read content from other authors and
use a little something from other sites.
Thanks for some other wonderful article. The place else could anyone get that kind of information in such an ideal approach of writing?
I’ve a presentation next week, and I’m on the search for
such info.
I have read so many articles on the topic of the blogger lovers
however this paragraph is in fact a good post, keep
it up.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/Yu2zoPfZCM
https://www.quia.com/profiles/monti469
https://www.metal-archives.com/users/cambridge19801955
https://www.obesityhelp.com/members/firs1974/about_me/
https://cannabis.net/user/151926
Hi there, I found your website by the use of Google at the same time as looking for a related matter, your web site got
here up, it looks great. I have bookmarked it
in my google bookmarks.
Hi there, simply become alert to your blog thru Google, and found that it is really informative.
I am going to be careful for brussels. I will appreciate if you
proceed this in future. Numerous folks will likely
be benefited from your writing. Cheers!
geinoutime.com
잠시 후 Hongzhi 황제는 “좋은 와인! “이라는 두 단어를 말했습니다.
В современном мире, где диплом становится началом отличной карьеры в любой области, многие ищут максимально быстрый путь получения образования. Факт наличия официального документа об образовании переоценить невозможно. Ведь именно он открывает дверь перед любым человеком, который стремится вступить в профессиональное сообщество или учиться в любом ВУЗе.
Наша компания предлагает максимально быстро получить любой необходимый документ. Вы имеете возможность приобрести диплом старого или нового образца, что является отличным решением для всех, кто не смог завершить образование, утратил документ или хочет исправить свои оценки. дипломы выпускаются с особой аккуратностью, вниманием к мельчайшим нюансам, чтобы в результате получился 100% оригинальный документ.
Превосходство этого решения заключается не только в том, что вы оперативно получите свой диплом. Весь процесс организован комфортно и легко, с нашей поддержкой. Начиная от выбора требуемого образца диплома до консультаций по заполнению личной информации и доставки по стране — все под абсолютным контролем опытных мастеров.
Всем, кто пытается найти оперативный способ получить требуемый документ, наша компания готова предложить выгодное решение. Заказать диплом – значит избежать долгого процесса обучения и не теряя времени переходить к личным целям: к поступлению в университет или к началу трудовой карьеры.
http://jaezfinancialgroup.icu
http://psemeetinglille2017.com
http://heroesrunchile.com
http://talktome.com.ua
http://zeptobars.ru
Hi, i think that i saw you visited my website thus
i came to “return the favor”.I’m attempting to find things to
enhance my web site!I suppose its ok to use some of your ideas!!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiZdiD
https://www.obesityhelp.com/members/artemkagg1992/about_me/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196317
https://gwet8281955.bandcamp.com/album/emma-02
https://haveagood.holiday/users/351221
I absolutely love your blog and find nearly all of your post’s to be precisely what I’m looking for. Do you offer guest writers to write content for you? I wouldn’t mind composing a post or elaborating on many of the subjects you write concerning here. Again, awesome weblog!
http://www.czystaenergiadwa.milanow.pl/viewtopic.php?t=17840
http://www.katamaran-isis.at/content/gästebuch?page=39
http://taroexpert.ru/arkan-otshelnik-praktika-raboty-s-klientom/
http://ozmillenium.ru/teatr/v-sti-obiavili-sezon-ychenikov.html
http://girls4gilrs.ru/page/4/
Hello there, I found your website via Google whilst
searching for a similar topic, your website came up,
it appears great. I have bookmarked it in my google bookmarks.
Hello there, simply became aware of your weblog through Google, and located that it’s truly informative.
I am going to be careful for brussels. I will appreciate for
those who proceed this in future. A lot of other people shall be benefited out of your writing.
Cheers!
Do you have any video of that? I’d care to find out more details.
http://oshibka-reshenie.ru/sharing-files-whatsapp
http://ntek-nt.ru/students/raspisanie/
http://usperm.ru/content/kuda-obratitsya-s-voprosom-po-provedeniyu-ege
http://avtoshkola34.ru/main/11-osnovnye-svedeniya-ob-avtoshkole-volzhskaya.html
http://misstres.ru/forum/search.php?action=members&p=195&s=d&order=ASC
Hey there outstanding blog! Does running a blog similar to this take a massive amount work? I’ve absolutely no expertise in computer programming however I had been hoping to start my own blog soon. Anyways, should you have any recommendations or techniques for new blog owners please share. I know this is off topic however I just had to ask. Appreciate it!
http://army.clanfm.ru/viewtopic.php?f=2&t=37960
Awesome blog! Do you have any helpful hints for aspiring writers?
I’m hoping to start my own website soon but
I’m a little lost on everything. Would you suggest starting with a free platform like WordPress or go for a
paid option? There are so many options out there that I’m completely overwhelmed ..
Any tips? Kudos!
Paragraph writing is also a excitement, if you be
acquainted with then you can write otherwise it is complex to write.
I am really impressed with your writing skills as well
as with the layout on your blog. Is this a
paid theme or did you modify it yourself? Either way keep up the
nice quality writing, it is rare to see a great
blog like this one these days.
This is a good tip particularly to those new to the blogosphere.
Brief but very accurate information… Thanks for sharing this one.
A must read article!
I was more than happy to uncover this site. I want to to thank
you for ones time due to this fantastic read!!
I definitely really liked every bit of it and I have you book-marked to see new things on your site.
Thank you a bunch for sharing this with all people you really realize what you’re talking about! Bookmarked. Kindly additionally visit my website =). We may have a link change arrangement among us
clicknconnectclubs.com/index.php?do=/public/user/blogs/name_Alanpoe/page_7/В
sol-dental.com/en/treatments/teeth-whitening/В
dienmayviet24h.com/dan-karaoke.htmlВ
allnewstroy.ru/effektivnoe-obrazovanie-kupit-diplom-i-dostich-tseleyВ
x-repair.ru/wp-includes/articles/?pokupka_diplomov_o_vusshem_obrazovanii_v_moskve.htmlВ
Way cool! Some very valid points! I appreciate you penning this post and
also the rest of the website is also really good.
Pretty! This has been an extremely wonderful article. Thank you for supplying this info.
ifvex.com/blogs/2501/Why-is-the-popularity-of-higher-education-decreasing-in-ourВ
officesetupc.com/blog/get-the-latest-features-in-office/В
conservatives.click/index.php?do=/public/user/blogs/name_Alanpoe/page_7/В
pipeclub.net/index.php?showuser=110605&k=880ea6a14ea49e853634fbdc5015a024&setlanguage=1&langurlbits=showuser=110605&cal_id=&langid=2В
triumph-flowers.ru/bukety-nevesty/yarkij-buket-nevesty/В
Hurrah! After all I got a webpage from where I be able to in fact get helpful facts concerning
my study and knowledge.
Everyone loves what you guys are up too. This kind of clever work and
coverage! Keep up the superb works guys I’ve included you guys
to our blogroll.
В современном мире, где диплом – это начало отличной карьеры в любой области, многие стараются найти максимально быстрый путь получения образования. Важность наличия официального документа переоценить невозможно. Ведь диплом открывает двери перед людьми, стремящимися вступить в сообщество квалифицированных специалистов или продолжить обучение в ВУЗе.
Наша компания предлагает максимально быстро получить этот необходимый документ. Вы можете приобрести диплом, что будет удачным решением для человека, который не смог завершить обучение, утратил документ или желает исправить плохие оценки. Все дипломы изготавливаются с особой аккуратностью, вниманием к мельчайшим нюансам, чтобы в результате получился продукт, максимально соответствующий оригиналу.
Преимущества подобного подхода заключаются не только в том, что вы сможете максимально быстро получить свой диплом. Процесс организовывается удобно, с профессиональной поддержкой. Начиная от выбора требуемого образца диплома до грамотного заполнения персональной информации и доставки по стране — все находится под полным контролем качественных специалистов.
Всем, кто ищет быстрый и простой способ получить необходимый документ, наша компания предлагает выгодное решение. Заказать диплом – это значит избежать длительного обучения и сразу переходить к достижению личных целей, будь то поступление в университет или начало удачной карьеры.
http://bancaanxudoithuong3d.com
http://newtab.page
http://baiontap.com
http://panarabiaenquirer.com
http://kimhospital.com
obviously like your website but you need to check the spelling on several of your posts.
A number of them are rife with spelling issues and I find it very bothersome to inform the reality on the other hand I
will definitely come again again.
I loved as much as you will receive carried out right
here. The sketch is tasteful, your authored material stylish.
nonetheless, you command get got an nervousness over that
you wish be delivering the following. unwell unquestionably
come further formerly again since exactly the same nearly very often inside case you shield this increase.
Undeniably imagine that which you said. Your favorite justification appeared to be on the web the easiest
factor to be aware of. I say to you, I definitely get irked at the same
time as folks consider issues that they just don’t know about.
You controlled to hit the nail upon the top and outlined out the entire thing with no need side-effects , folks can take a signal.
Will likely be again to get more. Thank you
My brother suggested I might like this web site. He used to be
entirely right. This publish actually made my day.
You cann’t imagine just how much time I had
spent for this info! Thank you!
비아그라 * 시알리스 * 레비트라
찾으시는 제품 모두 현재 1+1으로 하나 더
서비스 비아그라 * 시알리스 * 레비트라 * 중 한가지 4정 더
▶ 1+1 이벤트 국내최저가격
파워맨 사이트 바로가기👈
What’s Taking place i’m new to this, I stumbled upon this I have found
It absolutely helpful and it has helped me out loads.
I am hoping to give a contribution & assist other customers
like its helped me. Good job.
Please let me know if you’re looking for a article writer for your blog. You have some really good posts and I believe I would be a good asset. If you ever want to take some of the load off, I’d absolutely love to write some content for your blog in exchange for a link back to mine. Please send me an email if interested. Regards!
http://dermatologvenerolog.ru/obchee/21-displaziya-shejki-matki-i-tservikalnaya-intraepitelialnaya-neoplaziya-cin
http://balaklavskiy-16.ru/user/9409/
http://power-jump.ru/futbol/moizes-vyskazalsia-o-vozmojnosti-polycheniia-grajdanstva-rossii.html
http://jogibaba.com/blogs/53/We-buy-a-university-diploma-on-the-Internet-Tips?lang=tr_tr
http://vyazanienaspicah.ru/azhurnyj-dzhemper-spicami/
Hey there! I could have sworn I’ve been to this blog before but after
browsing through some of the post I realized it’s new to me.
Nonetheless, I’m definitely happy I found it and I’ll be book-marking and checking
back often!
Hello, constantly i used to check blog posts here early in the daylight,
as i love to gain knowledge of more and more.
WOW just what I was looking for. Came here by searching for %meta_keyword%
http://uziprosto.ru/tag/mrt-diagnostika
http://madled.ru/chto-takoe-psoriaz/kak-razlichit-psoriaz-i-seborejnyj-dermatit.html
http://vyazanshapki.ru/vyazanyie-shapki/zhenskie/shapka-s-obemnyim-pletenyim-uzorom-spitsami
http://jkg-portal.com.ua/ru/publication/one/nord-stream-2-ag-vdklala-zavershennja-pvnchnogo-potoku-do-veresnja-62556
http://myvera.ru/kosmolog
В наше время, когда диплом – это начало отличной карьеры в любой сфере, многие ищут максимально простой путь получения образования. Наличие официального документа переоценить попросту невозможно. Ведь именно диплом открывает двери перед любым человеком, который желает вступить в профессиональное сообщество или учиться в ВУЗе.
Предлагаем оперативно получить этот важный документ. Вы имеете возможность заказать диплом, и это является выгодным решением для человека, который не смог завершить обучение, утратил документ или хочет исправить свои оценки. дипломы изготавливаются аккуратно, с особым вниманием к мельчайшим нюансам. На выходе вы получите продукт, 100% соответствующий оригиналу.
Превосходство такого подхода заключается не только в том, что можно максимально быстро получить диплом. Весь процесс организован комфортно, с профессиональной поддержкой. Начиная от выбора нужного образца диплома до консультаций по заполнению личной информации и доставки в любой регион России — все под полным контролем квалифицированных мастеров.
Для тех, кто ищет быстрый и простой способ получить требуемый документ, наша компания предлагает отличное решение. Купить диплом – значит избежать длительного процесса обучения и сразу перейти к достижению личных целей: к поступлению в университет или к началу удачной карьеры.
http://bleiberecht.info
http://projetdexpert.fr
http://cafe-de-paris.ru
http://amicar22.ru
http://marsaxlokkfc.com
Wonderful work! That is the type of info that are meant to be shared around the internet.
Disgrace on the seek engines for now not positioning this put up higher!
Come on over and consult with my site . Thank you =)
XRumer – это самое успешное и еще лучшее программное обеспечение для создания ссылок (а) также продвижения сайтов. Оно используется для машинального прогона страниц через различные общественные сети, форумы, онлайн-дневники (а) также другие интернет-ресурсы.
http://www.botmasterru.com/product33230/
Купить Лицензированный Хрумер
Great beat ! I would like to apprentice whilst you amend your site,
how could i subscribe for a weblog website? The account helped
me a applicable deal. I were tiny bit familiar of this your broadcast provided brilliant clear concept
Hi there to every body, it’s my first pay a visit of this weblog; this blog includes awesome and truly good stuff in support of readers.
Whats up this is kinda of off topic but I was wanting to know if
blogs use WYSIWYG editors or if you have to manually
code with HTML. I’m starting a blog soon but have no coding know-how so I wanted to get guidance
from someone with experience. Any help would be enormously appreciated!
Hi, after reading this awesome piece of writing i am as well glad to share my experience here with mates.
http://nagrushe.ru/guest/gruppa-yantra
http://animesocial.su/blogs/1138/Как-купить-аттестат-или-диплом-у-нас-в-онлайн-магазине
http://demo2.weboss.hk/1009/comment/html/?578.html
http://www.humansandslaves.ru/blogs/48/Как-заказать-аттестат-или-диплом-у-нас-в-интернет-магазине?lang=tr_tr
http://opekaspb.ru/za-zreniem-nuzhen-uxod/
Сегодня, когда диплом является началом успешной карьеры в любом направлении, многие пытаются найти максимально простой путь получения образования. Важность наличия официального документа переоценить просто невозможно. Ведь диплом открывает двери перед всеми, кто желает начать трудовую деятельность или учиться в ВУЗе.
Предлагаем максимально быстро получить этот необходимый документ. Вы имеете возможность приобрести диплом, и это является выгодным решением для человека, который не смог завершить образование, потерял документ или хочет исправить свои оценки. диплом изготавливается аккуратно, с особым вниманием к мельчайшим деталям, чтобы в итоге получился полностью оригинальный документ.
Превосходство подобного подхода заключается не только в том, что вы сможете максимально быстро получить диплом. Процесс организовывается просто и легко, с профессиональной поддержкой. От выбора нужного образца документа до консультаций по заполнению персональной информации и доставки по стране — все будет находиться под полным контролем наших специалистов.
Для тех, кто ищет быстрый способ получить требуемый документ, наша компания может предложить отличное решение. Приобрести диплом – это значит избежать продолжительного обучения и сразу переходить к достижению личных целей: к поступлению в ВУЗ или к началу трудовой карьеры.
http://aaronegroup.com
http://fortyone.rocks
http://cekresipos.com
http://tontalk.ru
http://af-rv.fr
Awesome post.
Подарок для конкурента
https://xrumer.ru/
Такой пакет берут для переспама анкор листа сайта или мощно усилить НЧ запросы
Подарок для конкурента
Hello Dear, are you truly visiting this web page daily,
if so after that you will definitely take pleasant experience.
What’s up Dear, are you genuinely visiting this site on a regular basis, if so then you will absolutely get nice experience.
http://byte-kuzbass.ru/inetmagazvirt/category_00000092116/category_00000092117/category_00000104459/product_00000116923-detail
http://miupsik.ru/forums/showthread.php?tid=116&page=6
http://forum.myjane.ru/weblog.php?w=10332&previous=15&month=1&year=2012
http://os-masters.ru/post/kompyuternaya-pomoshch-i-nastroyka-interneta.html
http://fbuz74.ru/vybor-pnevmaticheskih-lobzikov/
Howdy this is somewhat of off topic but I was wondering if blogs use WYSIWYG editors or if you have to manually code with HTML. I’m starting a blog soon but have no coding knowledge so I wanted to get advice from someone with experience. Any help would be enormously appreciated!
http://www.katamaran-isis.at/content/gästebuch?page=39
http://rsn360.ru/blogs/150/Решили-купить-диплом-Обратитесь-в-наш-онлайн-магазин
http://www.ppe-online.net/product/level-2-isolation-gown?page=177
http://www.roofingrevenue.com/fdinlrpf/hadda-be-playing-on-the-jukebox-analysis.html
http://scct.ru/kozyrnye-kartochki
Hello to all, it’s truly a nice for me to go to see this web site, it consists of important Information.
Awesome post.
My spouse and I absolutely love your blog and find the majority of your post’s to be just what I’m looking for. can you offer guest writers to write content in your case? I wouldn’t mind publishing a post or elaborating on many of the subjects you write regarding here. Again, awesome weblog!
http://ksmzpn.pl/tabela-ekwiwalentow-2022/
http://krimoved-library.ru/books/349-dnevnaya-zaschita-sevastopolya1.html
http://georgi-kavkaz.ru/category/cooking
http://viparmenia.org/members/799-Unique
http://www.eysk-premier.ru/portu-nuzhen-rhimenes.html
Howdy I am so glad I found your webpage, I really found you by mistake, while I was browsing on Yahoo for something else, Anyhow I am here now and would just like to say many thanks for a fantastic post and a all round entertaining blog (I also love the theme/design), I don’t have time to browse it all at the moment but I have saved it and also added your RSS feeds, so when I have time I will be back to read a lot more, Please do keep up the awesome work.
http://profi-sk.ru/dizajn/pochemu-nalichie-diploma-vsyo-eshhe-igraet-sushhestvennuyu-rol.html
http://mdr7.ru/topic7603.html
http://znaxar.com/activity
http://chaek.ru/podstakanniki/stakan-dlya-podstakannika-khrustalnyy/reviews/
http://extreme-voyage.ru/2015/05/04/mnogocelevoj-takticheskij-tomagavk-browning-shock-n/
Что такое прогон по профилям?
На форумах при каждом прогоне регистрируется новый юзер, в профайле которого в поле Вебсайт или Домашняя страница стоит ваша ссылка. Также в некоторых форумах есть возможность в профиль вставить подпись, туда можно вставить ссылку с анкором. Все это можно варьировать. В профилях ссылки живут довольно долго, т.к. нет спама в чистом виде, и профили не мозолят глаза ни читателям форума, ни модераторам. Прогон производится последней версией хрумера с сервера с максимально возможной скоростью и производительностью.
ПОДРОБНЕЕ
НОВЫЙ ОТЛИЧНЫЙ VPN ДЛЯ XRUMERA!
За последние недели состоялась серия важных обновлений комплекса: XAuth, Captchas Benchmark, XRumer, XEvil 6.0 . В XEvil 6.0 обновлен механизм обработки hCaptcha и многое другое, в XAuth повышена стабильность работы, в XRumer повышена эффективность, устранён ряд погрешностей и обновлены прилагаемые базы.
https://xrumer.ru/
Have you ever thought about publishing an e-book or guest authoring
on other blogs? I have a blog based on the same subjects you discuss and would love to have you share some stories/information.
I know my audience would enjoy your work.
If you are even remotely interested, feel free to shoot me an email.
Very good article. I will be going through many of these issues as well..
https://seocontentify.59bloggers.com/27778589/Как-продвигать-сайт-с-xrumer-art
https://seoservify.ja-blog.com/27382114/Примеры-успешных-кейсов-студии-xrumer-art
Предлагаем вам выполнить консультацию (аудит) по подъему продаж а также доходы в вашем бизнесе. Формат аудита: личная встреча или онлайн конференция по скайпу. Делая верные, но простые усилия, доход от ВАШЕГО коммерциала получится увеличить в много раз. В нашем арсенале более 100 опробованных утилитарных схем повышения продаж и прибыли. В зависимости от вашего коммерциала подберем для вас максимально лучшие и будем постепенно претворять в жизнь.
– interestbook.ru
My developer is trying to persuade me to move to .net from PHP.
I have always disliked the idea because of the expenses.
But he’s tryiong none the less. I’ve been using Movable-type on various websites for about a year and am
worried about switching to another platform.
I have heard excellent things about blogengine.net.
Is there a way I can transfer all my wordpress content into it?
Any help would be really appreciated!
If you are going for finest contents like
myself, only visit this web site every day because it provides feature contents,
thanks
Asking questions are genuinely good thing if you are not understanding something entirely, except this article gives nice
understanding even.
Daddy Casino – Официальный телеграм канал со слотами от Дэдди Casino дэдди казино официальный сайт . Актуальное, рабочее зеркало официального сайта Дэдди Казино. Регистрируйся в Казино Daddy, получай бонус используя промокод, не забудь скачать
daddy casino играть
https://t.me/daddycasinorussia
I do consider all of the ideas you have presented in your post.
They are very convincing and will definitely work.
Still, the posts are very quick for newbies. Could you please lengthen them
a bit from next time? Thanks for the post.
What’s up every one, here every person is sharing these knowledge, thus it’s fastidious to read this web site, and I used to pay a quick visit this weblog daily.
musey-uglich.ru
It is not my first time to pay a visit this web page,
i am browsing this website dailly and obtain good data from here daily.
I absolutely love your blog and find most of your post’s
to be precisely what I’m looking for. can you offer guest writers
to write content for you? I wouldn’t mind creating
a post or elaborating on many of the subjects you write
concerning here. Again, awesome blog!
Thanks for sharing your thoughts about Betogel alternatif.
Regards
В современном мире, где диплом – это начало отличной карьеры в любом направлении, многие ищут максимально быстрый и простой путь получения качественного образования. Наличие официального документа об образовании переоценить попросту невозможно. Ведь диплом открывает дверь перед всеми, кто собирается вступить в сообщество профессионалов или продолжить обучение в любом ВУЗе.
Мы предлагаем максимально быстро получить этот важный документ. Вы можете приобрести диплом, и это становится отличным решением для человека, который не смог закончить образование, утратил документ или желает исправить плохие оценки. дипломы производятся с особой тщательностью, вниманием к мельчайшим элементам, чтобы в результате получился 100% оригинальный документ.
Превосходство этого решения состоит не только в том, что вы максимально быстро получите свой диплом. Весь процесс организован комфортно, с профессиональной поддержкой. Начав от выбора необходимого образца до консультации по заполнению персональной информации и доставки в любой регион России — все находится под полным контролем опытных специалистов.
В результате, для тех, кто ищет быстрый способ получения необходимого документа, наша услуга предлагает выгодное решение. Купить диплом – это значит избежать долгого процесса обучения и сразу перейти к личным целям: к поступлению в университет или к началу удачной карьеры.
seoyour.ru
Интернет-казино daddy казино играть — сайт или специальная программа, которые позволяют играть в азартные игры в интернете. Сайты интернет-казино могут специализироваться на различных азартных играх: игровых автоматах, рулетке, лотереях, викторинах, покере или других карточных и настольных играх. Википедия
daddy казино регистрация
https://t.me/daddycasinorussia
Currently it sounds like Expression Engine is the preferred blogging platform available right now.
(from what I’ve read) Is that what you’re using on your blog?
Have you ever thought about writing an ebook or guest authoring on other sites?
I have a blog centered on the same information you discuss and would love to have you share some stories/information. I know my subscribers would enjoy
your work. If you are even remotely interested, feel free
to shoot me an e-mail.
Телесуфлер — этто tvprompter.ru пишущий эти строки, кое помогает ораторам (а) также ведущим телепередач читать экспликация, никак не раскапывая позиции от камеры чи аудитории. Оно элементарно размещается унтер камерой да показывает текст один-другой содействием просвечивающего зеркала, создавая иллюзию ровного мнения в объектив. Текущий инструмент незаменим при проведении прямых эфиров также эпохально облегчает работу телеведущих.
http://tvprompter.ru/
Very energetic blog, I liked that a lot.
Will there be a part 2?
Hey, I think your site might be having browser compatibility issues. When I look at your website in Safari, it looks fine but when opening in Internet Explorer, it has some overlapping. I just wanted to give you a quick heads up! Other then that, very good blog!
http://www.musey-uglich.ru
I absolutely love your blog and find a lot of your post’s to be
precisely what I’m looking for. Does one offer guest writers to
write content for yourself? I wouldn’t mind creating a post or
elaborating on some of the subjects you write in relation to here.
Again, awesome site!
Wow, this piece of writing is fastidious, my younger
sister is analyzing these things, thus I am going to tell her.
geinoutime.com
적어도 그는 황태후가 화를 낼 수밖에 없다는 것을 알고 있었다.
Link exchange is nothing else but it is only placing the
other person’s webpage link on your page at appropriate place and other person will also do same in support of you.
슬롯 게임
가끔 해외에서 부자가 된다는 소문이 들리긴 하지만 이건 어디까지나 하늘의 얘기다.
Hey there I am so glad I found your site, I really found you by error,
while I was researching on Google for something else, Regardless I am here
now and would just like to say cheers for a marvelous post and a all round thrilling blog (I also love the theme/design),
I don’t have time to read it all at the moment but I have saved it
and also included your RSS feeds, so when I have time I will be back to read a lot
more, Please do keep up the awesome job.
Thanks for sharing your thoughts about login olxtoto.
Regards
you are truly a good webmaster. The web site loading speed is amazing.
It seems that you are doing any unique trick. Also, The contents are
masterwork. you’ve done a great job on this subject!
Great items from you, man. I have be mindful your stuff previous to and you are
just extremely fantastic. I actually like what you have received right here, really like what you’re saying and the way
by which you assert it. You’re making it entertaining and you continue to take care of to stay it sensible.
I cant wait to read much more from you. This is really a wonderful site.
В нашем мире, где диплом становится началом успешной карьеры в любой области, многие стараются найти максимально быстрый и простой путь получения образования. Важность наличия официального документа об образовании трудно переоценить. Ведь диплом открывает двери перед людьми, стремящимися вступить в профессиональное сообщество или учиться в высшем учебном заведении.
В данном контексте наша компания предлагает быстро получить этот необходимый документ. Вы имеете возможность купить диплом нового или старого образца, что становится отличным решением для всех, кто не смог завершить образование или потерял документ. Все дипломы выпускаются с особой аккуратностью, вниманием ко всем элементам. На выходе вы сможете получить продукт, 100% соответствующий оригиналу.
Преимущество данного решения состоит не только в том, что вы сможете оперативно получить свой диплом. Процесс организован комфортно, с нашей поддержкой. Начав от выбора подходящего образца до консультаций по заполнению персональных данных и доставки в любой регион страны — все под полным контролем качественных специалистов.
Таким образом, для всех, кто ищет быстрый способ получить требуемый документ, наша компания предлагает выгодное решение. Приобрести диплом – значит избежать долгого процесса обучения и сразу переходить к своим целям, будь то поступление в ВУЗ или начало карьеры.
seoyour.ru
wonderful points altogether, you simply gained a brand new reader.
What would you recommend in regards to your put up that you just made a few days
in the past? Any sure?
Great post. I used to be checking constantly this weblog and I’m impressed!
Extremely useful info specifically the closing part :
) I handle such information much. I used to be seeking this
certain info for a long time. Thank you and best of luck.
My family members always say that I am killing my time here at net, except I know I am getting knowledge every
day by reading such fastidious articles or reviews.
In fact when someone doesn’t know then its up to other people
that they will help, so here it happens.
When I originally commented I clicked the “Notify me when new comments are added” checkbox and now
each time a comment is added I get several e-mails with the same comment.
Is there any way you can remove me from that service?
Many thanks!
I’ve read a few just right stuff here. Certainly worth bookmarking for revisiting.
I surprise how much effort you put to create the sort of great informative web site.
It’s genuinely very complex in this busy life to listen news on Television,
therefore I just use web for that reason, and obtain the
most recent information.
В современном мире, где диплом является началом отличной карьеры в любой области, многие стараются найти максимально быстрый и простой путь получения качественного образования. Факт наличия официального документа об образовании переоценить попросту невозможно. Ведь именно он открывает двери перед людьми, стремящимися начать трудовую деятельность или учиться в высшем учебном заведении.
Мы предлагаем очень быстро получить этот необходимый документ. Вы сможете купить диплом, что становится выгодным решением для всех, кто не смог завершить образование, утратил документ или желает исправить плохие оценки. Все дипломы изготавливаются аккуратно, с максимальным вниманием к мельчайшим элементам. На выходе вы получите 100% оригинальный документ.
Плюсы этого подхода заключаются не только в том, что можно максимально быстро получить диплом. Процесс организовывается просто и легко, с нашей поддержкой. Начиная от выбора нужного образца документа до консультации по заполнению персональной информации и доставки по стране — все под абсолютным контролем квалифицированных мастеров.
Для тех, кто хочет найти быстрый и простой способ получить требуемый документ, наша услуга предлагает выгодное решение. Купить диплом – значит избежать долгого процесса обучения и сразу перейти к достижению собственных целей, будь то поступление в университет или старт карьеры.
http://www.seoyour.ru
Hi! I’m at work browsing your blog from my new iphone 3gs!
Just wanted to say I love reading your blog and look forward to
all your posts! Carry on the great work!
Hey There. I found your blog using msn. This is an extremely well written article.
I will make sure to bookmark it and come back to read more of your useful information. Thanks
for the post. I’ll definitely return.
Hi, its nice piece of writing on the topic of media print, we all be aware of media is a wonderful source of facts.
http://nebesis.ru/6-fevralya-znak-zodiaka-vodolej/
купить диплом моряка
http://labrusal.ru
купить диплом в миассе
Hey this is kind of of off topic but I was wanting to know if blogs use WYSIWYG editors or if you have to manually code with HTML. I’m starting a blog soon but have no coding expertise so I wanted to get guidance from someone with experience. Any help would be enormously appreciated!
купить диплом в кисловодске
http://nateconf.ru
http://i126.ru
http://cceis.ru
купить диплом в комсомольске-на-амуре
Just wish to say your article is as amazing. The clarity in your submit is just cool and that i can think you are
an expert in this subject. Fine along with your permission let me to grab your RSS feed
to keep up to date with impending post. Thank you 1,000,000 and
please carry on the enjoyable work.
Way cool! Some extremely valid points! I appreciate you writing this
article plus the rest of the website is extremely good.
Greate pieces. Keep posting such kind of info on your
blog. Im really impressed by your blog.
Hello there, You’ve done an excellent job.
I will definitely digg it and in my view suggest to my friends.
I am confident they’ll be benefited from this web site.
Great article. I am experiencing many of these issues as well..
It’s amazing to pay a visit this web page and reading the views of all friends about this article,
while I am also keen of getting knowledge.
Here is my page; dịch vụ vệ sinh công trình
Hey I know this is off topic but I was wondering if
you knew of any widgets I could add to my blog that automatically tweet my newest twitter updates.
I’ve been looking for a plug-in like this for quite some time and was hoping maybe
you would have some experience with something like this.
Please let me know if you run into anything. I truly enjoy reading your blog and I look forward to your new updates.
Hi, after reading this remarkable post i am as well glad to share my experience here with friends.
купить диплом в минусинске
http://neftgaztek.ru
http://transcollege.ru
http://copoko.ru
купить диплом нового образца
https://ultimatebabiesclub.com/2022/07/
https://f5fashion.vn/du-lich-hue-ghe-tham-nha-luu-niem-chu-tich-ho-chi-minh-o-lang-duong-no/
http://www.five-respect.co.jp/bbs/sunbbs.cgi?mode=form&no=126691467&page=
http://www.ferienwohnung-kettwig.de/index.php/component/phocagallery/1-appartement-bilder/index.php/component/phocagallery/1-appartement-bilder/gaestebuch.html?start=40
https://www.marinaioteatro.com/?page_id=12
We absolutely love your blog and find nearly all of your post’s to be exactly
I’m looking for. Do you offer guest writers to write content in your case?
I wouldn’t mind writing a post or elaborating on some of the subjects you write
about here. Again, awesome website!
Its such as you read my mind! You appear to grasp a lot
about this, such as you wrote the guide in it or something.
I think that you simply could do with some percent to power
the message house a little bit, but instead of that, this is excellent blog.
An excellent read. I will definitely be back.
I think the admin of this web site is in fact working hard in support
of his website, since here every information is quality
based data.
hi!,I really like your writing very a lot! share we keep up a correspondence more approximately
your article on AOL? I need an expert on this area to resolve
my problem. Maybe that’s you! Looking ahead to peer you.
What a data of un-ambiguity and preserveness of valuable know-how concerning unpredicted emotions.
купить диплом переводчика
http://uborka-kvartir-v-moskve.ru
http://r-diplom.ru
http://prohalal-faunia.ru
купить диплом логиста
I really like reading an article that will make men and women think.
Also, many thanks for permitting me to comment!
В современном мире, где диплом – это начало отличной карьеры в любом направлении, многие стараются найти максимально быстрый путь получения образования. Факт наличия документа об образовании трудно переоценить. Ведь диплом открывает двери перед всеми, кто желает начать трудовую деятельность или учиться в университете.
Предлагаем оперативно получить этот необходимый документ. Вы можете заказать диплом старого или нового образца, что является удачным решением для всех, кто не смог закончить обучение или потерял документ. диплом изготавливается аккуратно, с максимальным вниманием к мельчайшим нюансам. В итоге вы сможете получить полностью оригинальный документ.
Преимущества этого решения состоят не только в том, что вы сможете оперативно получить свой диплом. Процесс организован комфортно и легко, с профессиональной поддержкой. Начиная от выбора нужного образца до точного заполнения персональных данных и доставки в любое место России — все под полным контролем качественных мастеров.
Для тех, кто пытается найти быстрый и простой способ получения необходимого документа, наша услуга предлагает выгодное решение. Заказать диплом – это значит избежать длительного процесса обучения и сразу перейти к достижению своих целей, будь то поступление в ВУЗ или старт профессиональной карьеры.
http://281415.ru
http://n-seo.ru
http://to66minust.ru
http://diplomvash.ru
http://nti-nastavnik.ru
Howdy! This is kind of off topic but I need some guidance from an established blog.
Is it difficult to set up your own blog? I’m not very techincal but I can figure things out pretty quick.
I’m thinking about making my own but I’m
not sure where to begin. Do you have any ideas or suggestions?
Many thanks
Piece of writing writing is also a fun, if you be
familiar with after that you can write or else it is complicated to
write.
Its like you read my mind! You seem to know a lot about this,
like you wrote the book in it or something. I think that
you could do with a few pics to drive the message home a little bit, but instead of that, this is wonderful blog.
A fantastic read. I will definitely be back.
와일드 바운티 쇼다운
이렇게… 반년만에 고향으로 돌아갈 수 있다!
Hello, all is going fine here and ofcourse every one is sharing data, that’s really good,
keep up writing.
I do not even know how I finished up right here, but
I assumed this submit was great. I don’t recognize who you might be but
definitely you’re going to a well-known blogger in the event you are not already.
Cheers!
you are in point of fact a good webmaster. The site loading speed is incredible. It kind of feels that you’re doing any unique trick. Furthermore, The contents are masterwork. you’ve performed a fantastic task on this topic!
http://antennservice.ru/antenny-sputnikovye/aktuatory/vbox-ii-superjack-dp-6000-pozicioner-3-v-1-disegc-1-0-1-2-motor-59-pozicij/
купить диплом переводчика
http://energoprofi23.ru
купить диплом в кунгуре
Hello colleagues, pleasant article and nice urging commented at this place, I am genuinely enjoying by these.
http://s173028254.onlinehome.us/view_entry.php?id=800512&date=19760503
купить диплом в орле
http://stodorova.ru
купить диплом в кисловодске
Thanks for the auspicious writeup. It in fact was a enjoyment account it. Look advanced to more added agreeable from you! By the way, how could we communicate?
купить диплом в грозном
http://samogon-alko.ru
http://pojdelo-journal.ru
http://sartvo.ru
купить диплом в анжеро-судженске
I have read so many posts regarding the blogger lovers however this piece of writing is actually a good article, keep it up.
Greate article. Keep posting such kind of information on your site.
Im really impressed by your blog.
Hey there, You have done an incredible job. I will
certainly digg it and individually suggest to my friends.
I’m confident they’ll be benefited from this website.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/60297-ray-akuffo
https://chyoa.com/user/cfffg1959
http://www.babelcube.com/user/jessica-hunter-1
https://www.obesityhelp.com/members/bpedova1974/about_me/
https://www.quia.com/profiles/mafouks
Wow! This blog looks just like my old one! It’s on a entirely different subject but
it has pretty much the same page layout and design. Great choice of
colors!
I was excited to uncover this web site. I need to
to thank you for your time just for this fantastic read!!
I definitely appreciated every little bit of it and I have you book-marked to check out new stuff in your site.
I like the valuable info you provide in your articles. I will bookmark
your blog and check again here frequently. I’m quite sure I will learn many new stuff right
here! Best of luck for the next!
Pretty! This was an extremely wonderful post. Thank you for providing this info.
http://sapanet.ru/katalog-knig/advokatura/byt-advokatom-v-rossii-sotsiologicheskoe-issledovanie-professii1.html
купить диплом биолога
http://arxitekt.ru
можно ли купить диплом
В нашем обществе, где диплом – это начало удачной карьеры в любом направлении, многие стараются найти максимально быстрый путь получения качественного образования. Факт наличия официального документа об образовании переоценить невозможно. Ведь именно он открывает двери перед всеми, кто собирается начать профессиональную деятельность или учиться в каком-либо университете.
Наша компания предлагает максимально быстро получить любой необходимый документ. Вы имеете возможность приобрести диплом, что становится удачным решением для человека, который не смог завершить образование или потерял документ. Любой диплом изготавливается с особой аккуратностью, вниманием ко всем нюансам. На выходе вы получите продукт, 100% соответствующий оригиналу.
Плюсы этого решения состоят не только в том, что можно оперативно получить свой диплом. Процесс организован удобно и легко, с профессиональной поддержкой. Начав от выбора необходимого образца документа до точного заполнения личных данных и доставки в любой регион России — все под абсолютным контролем наших специалистов.
Всем, кто ищет оперативный способ получить необходимый документ, наша компания предлагает отличное решение. Купить диплом – значит избежать продолжительного процесса обучения и не теряя времени переходить к достижению своих целей, будь то поступление в университет или старт трудовой карьеры.
http://geniyz.ru
http://turgenev-2018.ru
http://samogon-alko.ru
http://kvn-irkutsk.ru
http://24-ur.ru
Thank you, I have recently been searching
for info approximately this subject for a long time
and yours is the best I’ve came upon so far. But, what in regards to the conclusion? Are you sure in regards to the source?
I have read so many posts about the blogger lovers but this piece of
writing is genuinely a pleasant article, keep it up.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://inferny1960.bandcamp.com/album/dna-chapter-4
http://www.babelcube.com/user/tracey-rivas
https://n3koo1990.diary.ru/
https://cannabis.net/user/151061
https://rentry.org/arw54k3k
I loved as much as you will receive carried out right here. The sketch is tasteful, your authored subject matter stylish. nonetheless, you command get got an edginess over that you wish be delivering the following. unwell unquestionably come further formerly again as exactly the same nearly a lot often inside case you shield this increase.
купить диплом в шахтах
http://1001diplom.ru
http://copoko.ru
http://opttextil.ru
купить диплом моряка
I’m really impressed together with your writing talents and
also with the layout to your blog. Is that this a paid topic or did you customize
it yourself? Either way keep up the excellent quality writing, it is uncommon to look
a nice weblog like this one today..
These are actually impressive ideas in concerning blogging.
You have touched some nice things here. Any way keep up wrinting.
Thanks for another magnificent article. Where else could anybody get
that kind of info in such an ideal method of writing? I have a presentation next week,
and I am at the look for such information.
This is my first time pay a visit at here and i am in fact impressed to read everthing at one place.
http://testkids.ru/zdorove/sindrom-dyryavoj-kishki.html
купить диплом в мурманске
http://pojdelo-journal.ru
купить диплом учителя физической культуры
This excellent website truly has all the information I wanted concerning this subject and didn’t know who to ask.
купить диплом в артеме
http://geafer.ru
http://ptncl.ru
http://galaxy-science.ru
купить диплом в воронеже
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/oni481954/profile
http://www.babelcube.com/user/angelica-luc
https://permacultureglobal.org/users/58547-robbie-clement
https://imageevent.com/liludala1972
https://rentry.org/vewxxo7y
Hello, constantly i used to check website posts here early in the daylight, since i love to gain knowledge of more and more.
http://ninokuni.ru/member.php?action=profile&uid=2098
http://verkehrsknoten.de/index.php/forum/gefahrgutrecht/85060-g228j#86015
http://yartube.ru/stati-v/456-nakrutka-vk-kak-mozhno-prodvinut-gruppu-vkontakte.html
http://led119.ru/forum/user/106177/
https://forum.qwas.ru/kupit-diplom-v-moskve-uslugi-ekspertov-t19723.html
The other day, while I was at work, my cousin stole my iPad and tested to see if it can survive a twenty five foot drop, just so she can be a youtube sensation. My apple ipad is now broken and she has 83 views. I know this is totally off topic but I had to share it with someone!
купить диплом математика
http://club-subaru.ru
http://naykadokcinema.ru
http://mediabat.ru
купить диплом в киселевске
Heya this is somewhat of off topic but I was wanting to know if blogs use WYSIWYG editors or if you have to manually code with HTML. I’m starting a blog soon but have no coding knowledge so I wanted to get guidance from someone with experience. Any help would be greatly appreciated!
https://salesianosayacucho.edu.pe/top-20-small-business-blogs-to-follow/
https://moskvic.actieforum.com/t3529-topic#6808
http://testowik.ru/index.php/forum/user/52046-emotusiro
http://feki-php.8u.cz/profile.php?lookup=11021
http://eurocodesguide.blogspot.com/2016/03/blog-post.html
Hi just wanted to give you a quick heads up and let you know a few of the pictures aren’t loading properly. I’m not sure why but I think its a linking issue. I’ve tried it in two different web browsers and both show the same outcome.
http://zanasite.free.fr/laripostedumardi/doku.php?id=landsdiploms
http://www.musichunt.pro/blogs/view.htm?id=79351
http://4asdaiprognoza.listbb.ru/viewtopic.php?f=2&t=539
https://together-19.com/read-blog/15811
http://www.ariawinebar.nyc/wv-contact.htmlВ
Its like you read my mind! You appear to know a lot about this, like you wrote the book in it or something.
I think that you could do with a few pics to drive the message home a bit, but other than that, this
is wonderful blog. A great read. I’ll definitely be back.
Please let me know if you’re looking for a author for your site. You have some really good posts and I think I would be a good asset. If you ever want to take some of the load off, I’d love to write some articles for your blog in exchange for a link back to mine. Please send me an email if interested. Regards!
купить диплом фельдшера
http://coaching-org.ru
http://megaskee.ru
http://fopko.ru
купить диплом в чапаевске
Hello! Someone in my Myspace group shared this website with us so I came to
look it over. I’m definitely loving the information. I’m bookmarking and will be tweeting this to my followers!
Great blog and brilliant style and design.
Pretty! This has been an incredibly wonderful article.
Thanks for supplying these details.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/kondratiy-197-4
https://rentry.org/4ehkvptx
https://www.obesityhelp.com/members/lllllllll1975/about_me/
http://www.nfomedia.com/profile?uid=rOjQfiB
https://imageevent.com/nurik1982
Very good information. Lucky me I found your blog by accident
(stumbleupon). I’ve book-marked it for later!
Подарок для конкурента
https://xrumer.ru/
Такой пакет берут для переспама анкор листа сайта или мощно усилить НЧ запросы
Подарок для конкурента
Heya i am for the primary time here. I came across this
board and I find It really useful & it helped me out a lot.
I’m hoping to offer something again and aid others such as you helped
me.
Right here is the perfect blog for anybody who wants to understand this topic. You realize so much its almost tough to argue with you (not that I personally would want to…HaHa). You certainly put a new spin on a topic that’s been discussed for many years. Great stuff, just wonderful!
купить диплом в старом осколе
http://school-internat5.ru
http://collegevalday.ru
http://unikum-student.ru
купить диплом для иностранцев
Hello! I know this is somewhat off topic but I was wondering which blog platform are you
using for this site? I’m getting sick and tired of WordPress because I’ve had issues with hackers and I’m looking at alternatives
for another platform. I would be awesome if you could
point me in the direction of a good platform.
It’s in point of fact a great and useful piece
of info. I’m glad that you shared this useful info with
us. Please stay us informed like this. Thank you for sharing.
Tremendous things here. I’m very happy to peer your article. Thanks a lot and I’m taking a look forward to contact you. Will you please drop me a e-mail?
http://xn--80ae1aidok.xn--p1ai/users/egajywam
taxizevs.ru/about/В
http://forumjustwoman.getbb.ru/viewtopic.php?f=43&t=900
http://g95334gq.beget.tech/2024/05/27/kupit-diplom-s-garantiey-podlinnosti.html
http://www.benedeek.com/blogs/79806/%D0%91%D1%8B%D1%81%D1%82%D1%80%D0%B0%D1%8F-%D0%B4%D0%BE%D1%80%D0%BE%D0%B3%D0%B0-%D0%BA-%D1%86%D0%B5%D0%BB%D0%B8-%D0%9A%D0%B0%D0%BA-%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC-%D0%B8-%D0%B4%D0%BE%D1%81%D1%82%D0%B8%D1%87%D1%8C-%D0%B6%D0%B5%D0%BB%D0%B0%D0%B5%D0%BC%D0%BE%D0%B3%D0%BEВ
I savor, cause I found exactly what I used to be looking for. You’ve ended my 4 day long hunt! God Bless you man. Have a nice day. Bye
купить диплом в братске
http://glavfinans.su
http://uspeh-kurs.ru
http://kdcmir.ru
купить диплом бакалавра
I’m really impressed with your writing skills as well as with the layout on your blog.
Is this a paid theme or did you customize it yourself? Either way keep up the excellent quality
writing, it’s rare to see a nice blog like this one today.
В нашем мире, где диплом – это начало удачной карьеры в любой сфере, многие стараются найти максимально быстрый путь получения образования. Важность наличия официального документа об образовании переоценить невозможно. Ведь диплом открывает двери перед людьми, стремящимися начать профессиональную деятельность или продолжить обучение в университете.
В данном контексте мы предлагаем оперативно получить этот важный документ. Вы сможете купить диплом, и это будет отличным решением для человека, который не смог закончить обучение, потерял документ или хочет исправить свои оценки. дипломы производятся с особой аккуратностью, вниманием к мельчайшим нюансам, чтобы в результате получился 100% оригинальный документ.
Плюсы этого решения состоят не только в том, что можно оперативно получить диплом. Процесс организован удобно, с нашей поддержкой. От выбора нужного образца до консультаций по заполнению личных данных и доставки по России — все под абсолютным контролем квалифицированных мастеров.
В результате, всем, кто хочет найти оперативный способ получения требуемого документа, наша компания предлагает отличное решение. Купить диплом – это значит избежать долгого обучения и сразу переходить к достижению собственных целей, будь то поступление в ВУЗ или старт успешной карьеры.
http://maksimdovzhenko.ru
http://turgenev-2018.ru
http://akadem-minstroy.ru
http://taxiasv.ru
http://interestbook.ru
Thanks on your marvelous posting! I seriously enjoyed
reading it, you can be a great author.I will make sure to bookmark your
blog and definitely will come back later on. I
want to encourage continue your great writing, have a nice afternoon!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/59438-carla-holmes
https://www.quia.com/profiles/jorgenorth
https://permacultureglobal.org/users/58783-andy-globa
https://thegapeon19771966.bandcamp.com/album/an-old-relation-pt-2
https://rentry.org/kcg259r6
It’s nearly impossible to find well-informed people for this subject, however, you seem like you know what you’re talking about!
Thanks
Hi there, I wish for to subscribe for this web site to take latest updates, so where can i do it please help.
https://poetzinc.com/read-blog/10756
https://www.fromrus.su/index.php?/topic/9671-%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC-%D0%BE-%D1%81%D1%80%D0%B5%D0%B4%D0%BD%D0%B5%D0%BC-%D1%81%D0%BF%D0%B5%D1%86%D0%B8%D0%B0%D0%BB%D1%8C%D0%BD%D0%BE%D0%BC-f659o/
http://www.castleofdracula.com.ru/forum/index.php
https://dolgoprudni.rusff.me/viewtopic.php?id=2078#p4730
https://huarenjiaohui.com/index.php?/topic/292-%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC-%D0%BE-%D1%81%D1%80%D0%B5%D0%B4%D0%BD%D0%B5%D0%BC-%D1%81%D0%BF%D0%B5%D1%86%D0%B8%D0%B0%D0%BB%D1%8C%D0%BD%D0%BE%D0%BC-x225g/
Hi there! Do you know if they make any plugins to safeguard against hackers? I’m kinda paranoid about losing everything I’ve worked hard on. Any suggestions?
http://aromatov.wooden-rock.ru/forum/topic.php?forum=1&topic=15533
http://sensemi.getbb.ru/viewtopic.php?f=6&t=501
https://wiki.hightgames.ru/index.php/пїЅпїЅпїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅпїЅ:_пїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅ_пїЅ_пїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅ
http://xn--d1aa3a4a.xn--p1ai/forum/topic.php?forum=3&topic=669
https://atlantcom.ru/forum/messages/forum1/topic89/message10633/?result=reply#message10633
Please let me know if you’re looking for a article writer for your blog. You have some really great posts and I think I would be a good asset. If you ever want to take some of the load off, I’d love to write some articles for your blog in exchange for a link back to mine. Please shoot me an e-mail if interested. Thank you!
https://ya.7bb.ru/viewtopic.php?id=11134#p32379
http://kamerton72.ru/node/220/done?sid=6830&token=5b2f5aff9d2e3b8070681e27a2f9f17e
https://roblox-studio-scripts.ru/profile/6664-michellhex/
https://milkyway.cs.rpi.edu/milkyway/show_user.php?userid=7072365
https://gmaii.ru/threads/1520/
My partner and I absolutely love your blog and find most of your post’s to be what precisely I’m looking for. Do you offer guest writers to write content available for you? I wouldn’t mind writing a post or elaborating on a number of the subjects you write about here. Again, awesome weblog!
купить диплом электромонтера
http://spravka-v-bassein.ru
http://stodorova.ru
http://lifelong-education.ru
купить диплом в вологде
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rayne12199719991968.diary.ru/
https://okwave.jp/profile/u3115695.html
https://launchpad.net/~asdef19661
https://rentry.org/3y5qpzgh
https://permacultureglobal.org/users/58885-elizabeth-madsen
What’s up Dear, are you genuinely visiting this website regularly, if so afterward you will definitely take good experience.
https://school-toksovo.ru/forum/messages/forum1/topic380/message559/?result=new#message559
http://myskupera.ru/forum/messages/forum1/topic12/message61255/?result=reply#message61255
http://forexrassia.ru/vasha-uspeshnaya-karera-nachinaetsya-s-diploma-pokupka-onlayn
эвакуатор-срочно-недорого.СЂС„/В
forexrassia.ru/page/4В
Definitely consider that which you stated. Your favorite reason appeared
to be on the net the simplest factor to consider of. I say to you,
I certainly get irked whilst people think about worries that they just don’t recognise about.
You controlled to hit the nail upon the highest and also defined out
the whole thing with no need side-effects , people can take a signal.
Will probably be again to get more. Thank you
Today, while I was at work, my cousin stole my apple ipad and tested to see if it can survive a 40 foot drop, just so she can be a youtube sensation. My apple ipad is now destroyed and she has 83 views. I know this is entirely off topic but I had to share it with someone!
купить диплом о высшем образовании
http://forum-ecology.ru
http://opendatachel.ru
http://students-programmers.ru
купить диплом в северодвинске
В нашем мире, где диплом – это начало успешной карьеры в любом направлении, многие ищут максимально быстрый путь получения качественного образования. Наличие официального документа об образовании переоценить невозможно. Ведь диплом открывает дверь перед людьми, стремящимися начать профессиональную деятельность или учиться в ВУЗе.
В данном контексте наша компания предлагает очень быстро получить этот важный документ. Вы сможете заказать диплом старого или нового образца, и это является выгодным решением для человека, который не смог закончить обучение или потерял документ. диплом изготавливается с особой тщательностью, вниманием ко всем элементам. На выходе вы сможете получить полностью оригинальный документ.
Плюсы такого решения состоят не только в том, что можно оперативно получить свой диплом. Весь процесс организован комфортно, с профессиональной поддержкой. От выбора требуемого образца документа до точного заполнения личных данных и доставки по стране — все будет находиться под полным контролем наших мастеров.
Для всех, кто хочет найти быстрый и простой способ получить требуемый документ, наша услуга предлагает отличное решение. Приобрести диплом – значит избежать долгого процесса обучения и не теряя времени перейти к достижению личных целей, будь то поступление в университет или начало карьеры.
http://galaxy-science.ru
http://dety-tymovsk.ru
http://fast-education.ru
http://primzona.ru
http://pg2017.ru
Attractive component to content. I simply stumbled upon your web site and in accession capital to claim that I get in fact loved account your weblog posts. Any way I will be subscribing in your augment or even I achievement you access persistently rapidly.
https://solo-it.ru/forum/?PAGE_NAME=profile_view&UID=3715
http://ya.10bb.ru/viewtopic.php?id=3432#p6251
https://inderculturaputumayo.gov.co/se-definieron-los-campeones-de-deportes-de-conjunto-de-los-juegos-intercolegiados-2023/
http://mayakminska.1stbb.ru/viewtopic.php?f=12&t=11944
https://goruss.ru/blogs/397/ упить-диплом-ваш-шаг-к-новой-профессиональной-жизни-без-лишних
What’s up, its good post about media print, we all be aware of media is a great source of facts.
http://maidanrb.blogspot.com/2014/09/blog-post_2.html
floriann.ru/buket-rozВ
http://samorezi.com/vse_novosti/novosti_iz_mira_homyachkov/
http://ulmo.ukrbb.net/index.php
http://diy-tour-kgd.blogspot.com/2011/11/blog-post.html
Hi everyone, it’s my first pay a quick visit
at this website, and article is actually fruitful in support of me,
keep up posting these content.
В нашем обществе, где диплом – это начало отличной карьеры в любом направлении, многие стараются найти максимально быстрый и простой путь получения образования. Наличие официального документа переоценить просто невозможно. Ведь диплом открывает двери перед всеми, кто стремится начать профессиональную деятельность или учиться в ВУЗе.
Предлагаем быстро получить этот необходимый документ. Вы имеете возможность купить диплом старого или нового образца, что становится удачным решением для человека, который не смог завершить образование или утратил документ. диплом изготавливается с особой тщательностью, вниманием ко всем деталям, чтобы в итоге получился документ, полностью соответствующий оригиналу.
Преимущество данного подхода состоит не только в том, что можно оперативно получить свой диплом. Процесс организовывается просто и легко, с нашей поддержкой. От выбора нужного образца до консультации по заполнению персональных данных и доставки по стране — все находится под абсолютным контролем наших мастеров.
Для всех, кто ищет быстрый способ получения необходимого документа, наша компания готова предложить выгодное решение. Купить диплом – это значит избежать длительного обучения и сразу перейти к важным целям: к поступлению в ВУЗ или к началу удачной карьеры.
http://akadem-minstroy.ru
http://students-programmers.ru
http://thejackwood.com
http://fgu-avto.ru
http://lawdiplom.ru
Hey there! I just wanted to ask if you ever have any
trouble with hackers? My last blog (wordpress) was hacked and
I ended up losing a few months of hard work due to no back up.
Do you have any solutions to stop hackers?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/uglycat1959/profile
https://haveagood.holiday/users/350123
https://okwave.jp/profile/u3115212.html
https://www.obesityhelp.com/members/xxwitcher1975/about_me/
https://imageevent.com/nerai1983
Wow that was odd. I just wrote an very long comment but after I clicked submit my comment didn’t appear. Grrrr… well I’m not writing all that over again. Regardless, just wanted to say superb blog!
http://www.ettoresilini.ch/2017/07/18/da-mio-giardino-con-furore-dictyophara-europea/
купить диплом в тобольске
http://opexobo.ru
купить диплом в димитровграде
I was recommended this blog by my cousin. I am not sure whether this post is written by him as
no one else know such detailed about my trouble. You’re incredible!
Thanks!
Marvelous, what a web site it is! This blog provides helpful data
to us, keep it up.
As the admin of this web page is working, no question very soon it will be renowned, due to its feature contents.
https://avt-avto.ru/categories/dlya-avtomoyki/ochistnye-sooruzheniya-dlya-avtomoek-/sistemy-ochistki-vody-na-4-moechnykh-posta/sistema_ochistki_vody_dlya_avtomoek_aros_4_dk_s_dozatorom_khim_reagenta_i_kartridzhnym_filtrom/
http://suhinfo.ru/index.php/пїЅпїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅ:_пїЅпїЅпїЅ_пїЅпїЅпїЅпїЅ_пїЅ_пїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅ_пїЅ_пїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅпїЅ!
http://avtoweek2016.ru/kak-legko-kupit-diplom-i-uspeshno-startovat-v-karere/
http://crazyhandmadic.blogspot.com/2013/05/1.html
http://klinlife.ru/forums/index.php?/topic/11580-kupit-diplom-podlinnie-dokumenti-po-dostupno/
Thank you for the auspicious writeup. It if truth be told used to be a amusement account it. Glance complicated to far delivered agreeable from you! By the way, how could we be in contact?
купить диплом бакалавра
http://kapremont-upgrade.ru
http://seozalex.ru
http://uinvestor.ru
купить аттестат за 9 классов
Pretty! This has been a really wonderful post. Many thanks for supplying these details.
https://idi.atu.edu.iq/?p=6050
https://rockufa.ru/forum/index.php?action=profile;u=176528
https://forum.l2gavno.ru/members/dustindSoame.6676/
https://www.ocf.berkeley.edu/~paultkim/coronavirus-continues-to-disrupt-the-e-reader-industry/
http://msfo-soft.ru/msfo/forum/messages/forum25/topic9298/message276187/?result=new#message276187
Hi there to all, the contents present at this website are really awesome
for people experience, well, keep up the nice work fellows.
I visited multiple web sites but the audio feature for audio songs
existing at this web page is in fact wonderful.
Hey there excellent website! Does running a blog like this require a
lot of work? I have absolutely no understanding of programming however I was hoping to start my own blog
soon. Anyhow, if you have any suggestions or tips for
new blog owners please share. I know this is off
topic but I simply needed to ask. Kudos!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196526
https://rentry.org/i8bbpzvu
https://imageevent.com/pivasixa1982
https://www.haikudeck.com/presentations/LVGqvXbB7y
https://rentry.org/if89mzwg
I loved as much as you’ll receive carried out right here. The sketch is attractive, your authored subject matter stylish. nonetheless, you command get bought an nervousness over that you wish be delivering the following. unwell unquestionably come further formerly again as exactly the same nearly very often inside case you shield this hike.
купить диплом моториста
http://cardioevent.ru
http://sartvo.ru
http://kolomna-junior.ru
купить диплом программиста
For newest information you have to go to see internet and on web I found this web page as a finest website for newest updates.
https://www.skillsmalaysia.gov.my/index.php/k2-full-width/item/7-quis-quam-elementum-ed-dignissim-ipsum-condi
https://blogs.rufox.ru/~runoklisfgy/42169.htm
http://www.josegregoriodelarivera.com/component/fst/?view=test&prodid=0
http://xn—-8sbkkh5cbijf2k.xn--p1ai/index.php?option=com_k2&view=itemlist&task=user&id=53351
http://anatoliyrud.ekafe.ru/viewforum.php?f=29
I think that everything posted made a bunch of sense.
But, what about this? what if you were to create a killer headline?
I ain’t suggesting your information isn’t good, but suppose you added a post
title that grabbed people’s attention? I mean Fornix (crus)/
Stria Terminalis – fxst | Whole Brain Protocol
for Tractography with Empirical MRI is a little boring. You ought to look at
Yahoo’s home page and watch how they write article titles to grab viewers to open the links.
You might try adding a video or a related picture or
two to get people interested about what you’ve written. Just
my opinion, it might make your posts a little livelier.
Fantastic data, Thanks a lot.
I used to be suggested this web site through my cousin.
I’m now not sure whether or not this post is written via
him as nobody else recognise such certain about my
difficulty. You’re incredible! Thank you!
I’m amazed, I have to admit. Seldom do I encounter a blog that’s both educative and entertaining, and let
me tell you, you have hit the nail on the head. The problem is something that not enough folks are speaking
intelligently about. I am very happy I came across this during my hunt for something regarding
this.
What i do not understood is in fact how you’re no longer actually much more well-favored than you might be now.
You are very intelligent. You recognize therefore significantly on the subject of this topic,
produced me in my opinion imagine it from so many varied angles.
Its like women and men are not fascinated until it is one thing to accomplish
with Lady gaga! Your personal stuffs nice. All the time handle
it up!
Hmm it looks like your blog ate my first comment (it was super long) so I guess I’ll
just sum it up what I had written and say, I’m thoroughly
enjoying your blog. I too am an aspiring blog blogger but I’m still new to everything.
Do you have any points for novice blog writers? I’d genuinely appreciate it.
В нашем обществе, где диплом становится началом удачной карьеры в любом направлении, многие ищут максимально простой путь получения качественного образования. Необходимость наличия официального документа переоценить просто невозможно. Ведь диплом открывает двери перед людьми, стремящимися вступить в профессиональное сообщество или учиться в любом институте.
В данном контексте наша компания предлагает быстро получить этот важный документ. Вы имеете возможность заказать диплом нового или старого образца, и это будет выгодным решением для человека, который не смог закончить обучение или потерял документ. Все дипломы выпускаются с особой аккуратностью, вниманием ко всем элементам. На выходе вы сможете получить полностью оригинальный документ.
Преимущество данного подхода состоит не только в том, что вы быстро получите диплом. Процесс организован удобно и легко, с профессиональной поддержкой. От выбора подходящего образца документа до грамотного заполнения персональных данных и доставки в любой регион России — все будет находиться под абсолютным контролем опытных мастеров.
Всем, кто хочет найти максимально быстрый способ получить необходимый документ, наша компания может предложить выгодное решение. Заказать диплом – это значит избежать длительного процесса обучения и сразу переходить к своим целям: к поступлению в ВУЗ или к началу успешной карьеры.
http://efawb.ru
http://ipuslu.ru
http://ddtzd.ru
http://borovskie-izvestiya.ru
http://nateconf.ru
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~remka19921
https://www.hentai-foundry.com/user/mamuka19721983/profile
https://www.haikudeck.com/presentations/djwCWlqJob
https://www.hentai-foundry.com/user/orionte1977/profile
https://rentry.org/ses76xkd
Can you tell us more about this? I’d love to find out some additional information.
купить свидетельство о рождении ссср
http://diplomgos.info/
http://ovolgodonske.ru
http://diplom61.ru
купить морской диплом
What’s Happening i am new to this, I stumbled upon this I have discovered It positively useful and it
has aided me out loads. I am hoping to give a contribution &
help different users like its aided me. Great job.
You’re so cool! I don’t think I have read anything like this before.
So wonderful to find someone with original thoughts on this subject matter.
Really.. thank you for starting this up. This site is one thing that is required on the internet, someone with a bit of originality!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOjQccG
https://haveagood.holiday/users/349672
http://www.nfomedia.com/profile?uid=rOiXhiJ
https://haveagood.holiday/users/351549
https://cannabis.net/user/150783
Hello! Do you use Twitter? I’d like to follow you if that would be okay. I’m undoubtedly enjoying your blog and look forward to new posts.
http://www.usman2.ru/dwsvid/195745-vzlom-moj-govoryaschij-tom.html
купить диплом в волгограде
http://efawb.ru
купить диплом в кропоткине
Доброго всем дня!
Закажите диплом ВУЗа по выгодной цене с доставкой в любой регион России без предоплаты и с полной уверенностью в его законности!
https://chrstms.ru/club/user/11503/blog/10563/
https://tatiananovo.com/hello-world/#comment-7319
http://www.starterkit.ru/html/index.php?name=account&op=info&uname=ocaluv
http://dnd.listbb.ru/viewtopic.php?f=3&t=411
https://forum.mbprinteddroids.com/showthread.php?tid=1235
Right here is the right website for anyone who wishes to find out about this topic. You realize a whole lot its almost tough to argue with you (not that I actually would want to…HaHa). You definitely put a new spin on a subject that has been discussed for decades. Excellent stuff, just wonderful!
купить диплом в курске
http://cleo-dance.ru
http://diplom61.ru
http://git-ugra.ru
купить диплом переводчика
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196804
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3197077
https://spirit019871998.diary.ru/
https://www.quia.com/profiles/donniemu
https://www.haikudeck.com/presentations/L1AwY7aKMF
В современном мире, где диплом является началом успешной карьеры в любой сфере, многие пытаются найти максимально быстрый путь получения образования. Необходимость наличия официального документа об образовании трудно переоценить. Ведь именно он открывает дверь перед всеми, кто хочет вступить в сообщество профессионалов или учиться в высшем учебном заведении.
В данном контексте мы предлагаем максимально быстро получить этот необходимый документ. Вы имеете возможность заказать диплом, и это становится удачным решением для человека, который не смог закончить обучение или потерял документ. Любой диплом изготавливается аккуратно, с особым вниманием ко всем деталям, чтобы в итоге получился продукт, максимально соответствующий оригиналу.
Преимущество этого решения заключается не только в том, что вы быстро получите свой диплом. Процесс организовывается удобно, с нашей поддержкой. От выбора нужного образца до консультаций по заполнению личной информации и доставки по России — все будет находиться под полным контролем наших мастеров.
Всем, кто хочет найти оперативный способ получить необходимый документ, наша компания предлагает отличное решение. Приобрести диплом – значит избежать продолжительного обучения и не теряя времени переходить к достижению своих целей: к поступлению в ВУЗ или к началу трудовой карьеры.
http://ot1do8.ru
http://eurasianlawstudents.ru
http://taxiasv.ru
http://referat-diplom.ru
http://family-spb.ru
geinoutime.com
이에 그는 곧바로 국방부로 달려가 국방부에 보고했다.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiYgdH
https://sunlu1953.bandcamp.com/album/hot-springs
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196698
http://www.babelcube.com/user/alan-barney
https://wertrew1970.bandcamp.com/album/great-everyday-sex
Hi there to all, how is all, I think every
one is getting more from this website, and your views are
pleasant in support of new people.
Can I just say what a relief to discover an individual who truly understands what they’re talking about on the net. You certainly know how to bring an issue to light and make it important. More people should check this out and understand this side of your story. I can’t believe you aren’t more popular given that you certainly possess the gift.
купить диплом в георгиевске
http://studyinkaluga.ru
http://seo1st.ru
http://kinelgorod.ru
купить диплом техникума
Thanks to my father who stated to me concerning this blog, this webpage is truly amazing.
В нашем обществе, где диплом – это начало удачной карьеры в любой отрасли, многие стараются найти максимально простой путь получения образования. Факт наличия официального документа переоценить невозможно. Ведь диплом открывает дверь перед всеми, кто желает начать профессиональную деятельность или учиться в высшем учебном заведении.
Мы предлагаем максимально быстро получить любой необходимый документ. Вы имеете возможность приобрести диплом нового или старого образца, что будет выгодным решением для всех, кто не смог закончить обучение или потерял документ. Любой диплом изготавливается аккуратно, с особым вниманием к мельчайшим элементам, чтобы в итоге получился продукт, максимально соответствующий оригиналу.
Преимущества данного подхода состоят не только в том, что можно быстро получить диплом. Весь процесс организован комфортно и легко, с профессиональной поддержкой. Начиная от выбора подходящего образца диплома до грамотного заполнения персональных данных и доставки по России — все под абсолютным контролем наших специалистов.
Для всех, кто ищет максимально быстрый способ получения необходимого документа, наша услуга предлагает отличное решение. Заказать диплом – это значит избежать длительного процесса обучения и не теряя времени переходить к достижению собственных целей, будь то поступление в университет или начало карьеры.
http://37vilok.ru
http://vapediscont.ru
http://books4study.info
http://avtoshkola142.ru
http://extra-art.ru
It’s really a cool and useful piece of info.
I’m glad that you simply shared this useful info with us. Please keep us
up to date like this. Thank you for sharing.
Good day I am so happy I found your blog page, I really found you by accident, while I was searching on Bing for something else, Regardless I am here now and would just like to say thank you for a fantastic post and a all round enjoyable blog (I also love the theme/design), I don’t have time to look over it all at the moment but I have book-marked it and also included your RSS feeds, so when I have time I will be back to read a lot more, Please do keep up the fantastic work.
http://socialle.com.br/blogs/35/We-purchase-a-diploma-on-the-Internet-Expert-Tips
купить диплом медицинского училища
http://referat-diplom.ru
купить диплом о высшем образовании
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195852
https://haveagood.holiday/users/351599
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3197533
https://permacultureglobal.org/users/60090-luis-navarro
https://www.hentai-foundry.com/user/lisher1975199819811981197919791969/profile
I loved as much as you’ll receive carried out right here. The sketch is attractive, your authored subject matter stylish. nonetheless, you command get got an nervousness over that you wish be delivering the following. unwell unquestionably come further formerly again since exactly the same nearly very often inside case you shield this hike.
https://reklama-net.ru/opros.php?opros=1
http://www.musichunt.pro/blogs/view.htm?id=79366
http://newsato.ru/kupit-diplom-garantiya-legalnosti
http://lider1c.ru/communication/forum/messages/forum5/message7103/5483-aurusdiploms?result=new#message7103
http://arbitrajniki.ru/forums/forum/forum/
After looking over a handful of the blog articles on your blog,
I honestly appreciate your way of blogging. I saved as a favorite it to my bookmark webpage list and will be checking back soon. Take a look at my website
as well and tell me your opinion.
Hi, I think your site might be having browser compatibility issues.
When I look at your blog in Opera, it looks fine but when opening in Internet Explorer, it
has some overlapping. I just wanted to give you a quick heads up!
Other then that, great blog!
Excellent blog you have here but I was curious if you knew of any forums that cover the same topics talked about in this article? I’d really love to be a part of online community where I can get responses from other experienced people that share the same interest. If you have any suggestions, please let me know. Thank you!
myworldavto.ru/page/8
http://www.lku2.com/blogs/4238/Where-to-buy-a-diploma-or-certificate-at-an-adequate
http://www.snipesocial.co.uk/blogs/370679/%D0%9E%D1%82%D0%BA%D1%80%D1%8B%D1%82%D0%B8%D0%B5-%D0%B2%D0%BE%D0%B7%D0%BC%D0%BE%D0%B6%D0%BD%D0%BE%D1%81%D1%82%D0%B5%D0%B9-%D1%86%D0%B5%D0%BD%D0%BD%D0%BE%D1%258?lang=tr_tr
pipeclub.net/index.php?showuser=110605&k=880ea6a14ea49e853634fbdc5015a024&setlanguage=1&langurlbits=showuser=110605&cal_id=&langid=2
goodgame.ru/user/1629660
This is really interesting, You are an overly skilled blogger. I have joined your rss feed and look forward to in quest of more of your excellent post. Additionally, I have shared your web site in my social networks
купить диплом в томске
https://mamuli.club/forum/topic/23701/
https://ekocom.ru/forum/?PAGE_NAME=profile_view&UID=12840
http://vseogirls.ru/kupit-diplom-vyisokie-standartyi-kachestva
http://avrorp.getbb.ru/viewtopic.php?f=183&t=571
http://bigbangclan.mybb.ru/viewtopic.php?id=351#p818
купить диплом в октябрьском
Первый партнер охраны труда
[url=https://www.ppot.ru/]Первый Партнер охраны труда[/url] – экспертный центр по аудиту и подготовке специалистов в области охраны труда и пожарной безопасности. Наша компания предоставляет широкий спектр услуг по обеспечению безопасности производственных процессов для индивидуальных предпринимателей и организаций различных секторов промышленности в Москве, Московской области и других регионах России.
Соблюдение правил безопасности труда – законодательное требование, за нарушение которого на организацию может быть наложен штраф или приостановлена деятельность предприятия на срок до 90 суток.
Мы помогаем решать вопросы обеспечения безопасности и организации труда на предприятии, разрабатываем и актуализируем документы по охране труда, проводим комплексное обследование охраны труда на соответствие государственным нормативным требованиям. Проводим обучение сотрудников по охране труда и пожарной безопасности, проверяем СУОТ и оцениваем профессиональные риски.
Hello, i think that i saw you visited my blog so i came to “return the favor”.I’m attempting to
find things to improve my website!I suppose its ok to use some
of your ideas!!
Admiring the time and effort you put into your blog and detailed information you provide. It’s great to come across a blog every once in a while that isn’t the same out of date rehashed material. Excellent read! I’ve bookmarked your site and I’m including your RSS feeds to my Google account.
http://define.listbb.ru/viewtopic.php?f=46&t=434
https://epistles.ru/blogs/905/ѕуть-к-успеху- упи-диплом-и-преуспей-в-карьере
http://maidanrb.blogspot.com/2014/09/blog-post_2.html
http://zimmu.com/api/manual/doku.php?id=landsdiploms
https://roblox-studio-scripts.ru/topic/18481-kupit-diplom-tehnikuma-nedorogo-e218t/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/evar1954/about_me/
https://www.hentai-foundry.com/user/marsingel1955/profile
http://www.babelcube.com/user/lori-aguilar
https://cannabis.net/user/152120
https://www.metal-archives.com/users/tsog1971
Thank you for sharing your thoughts. I truly appreciate your efforts and I am waiting for your further post thanks once again.
http://vipmails.0pk.me/viewtopic.php?id=5778#p12979
http://kotka.listbb.ru/viewtopic.php?f=2&t=399
https://rcdrift.ru/forum/member.php?u=21074
http://www.twentyfourpixel.de/beispiel-seite/
http://xn--b1aafrvecb0f6b.xn--p1ai/forum/messages/forum4/topic840/message264405/?result=reply#message264405
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/gloveren1976/profile
https://permacultureglobal.org/users/59225-ashley-johnson
https://imageevent.com/elerra1982
https://imageevent.com/pohhbsc19751971
https://anotepad.com/notes/rnp379dx
Hi there, of course this post is in fact pleasant and I have learned lot of things from it concerning blogging. thanks.
купить диплом в альметьевске
http://newsato.ru/kupit-diplom-otkroyte-dver-k-novyim-vozmozhnostyam
http://coloringcrew.com/iphone-ipad/?url=https://premialnie-diplomans.com/
http://gamesmaker.ru/forum/common/offtopics/#replyform
https://www.easyinsurancehub.co.uk/forum/thread-5625.html
https://www.khansaschool.com/blogs/49183/ упить-диплом-в-ћоскве- онфиденциально
купить диплом в ялте
It’s truly a nice and helpful piece of info. I am satisfied that
you simply shared this useful info with us. Please keep
us informed like this. Thank you for sharing.
It’s appropriate time to make a few plans for the long run and it is time
to be happy. I’ve learn this publish and if I may I want to recommend you some fascinating issues or tips.
Maybe you can write subsequent articles referring to this article.
I want to read more issues approximately it!
Hi there Dear, are you truly visiting this website regularly, if so then you will absolutely
obtain nice experience.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151890
https://devina1972.diary.ru/
https://chyoa.com/user/llama1987
https://www.quia.com/profiles/jiwelsh
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196105
What a stuff of un-ambiguity and preserveness of valuable familiarity on the topic of unexpected feelings.
http://grot.getbb.ru/viewtopic.php?f=6&t=964
https://www.shopwheel.ru/club/user/52/blog/
http://kinogonews.ru/poluchite-diplom-mechtyi-legko-i-dostupno
http://ninokuni.ru/index.php
https://www.ocf.berkeley.edu/~paultkim/macmillan-is-no-longer-embaroing-ebooks-to-public-libraries/
It is truly a great and useful piece of info. I’m happy that you simply shared this useful information with us.
Please keep us informed like this. Thanks for sharing.
Hello! I know this is somewhat off topic but I was wondering which blog platform
are you using for this site? I’m getting fed up of WordPress
because I’ve had problems with hackers and I’m looking at alternatives for another platform.
I would be fantastic if you could point me in the
direction of a good platform.
I got this web site from my pal who shared with me regarding this
website and now this time I am visiting this site and reading very informative content
here.
Good info. Lucky me I recently found your blog by accident (stumbleupon).
I have saved it for later!
купить диплом повара diplomvash.ru .
Рекомендуем купить грифы для гантелей на grify-dlya-gantely ruпо доступным ценамоптимальной длины. В изготовлении долговечных изделий применяются высококлассные марки металла. Комплектующие для гантелей производятся в трех востребованных диаметрах. Комплектующие предназначены для силовых занятий и выполнены с разметкой и насечками для надежного хвата. Продукты покрываются защитным слоем никеля. Российская компания создает широкий объем спортивного инвентаря для квартиры и фитнес клуба. Это многофункциональный инвентарь для силовых тренировок в любых условиях.
Привет, дорогой читатель!
Приобретите российский диплом по выгодной цене с гарантией прохождения проверок и доставкой в любой город РФ!
http://blogs.affordablehvacpa.com/a-guide-to-charging-heat-pumps-during-the-cold-season/#comment-409518
http://sfera100.ru/forum/user/1830/
http://army.clanfm.ru/viewtopic.php?f=2&t=35747
http://gadjetlife.ru/kupit-nastoyashhiy-diplom-o-vyisshem-obrazovanii
http://x-universe.ru/index.php?title=%D0%9D%D1%83%D0%B6%D0%B5%D0%BD_%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC%3F_%D0%9B%D1%83%D1%87%D1%88%D0%B8%D0%B5_%D1%86%D0%B5%D0%BD%D1%8B_%D1%82%D1%83%D1%82!
This text is worth everyone’s attention. Where can I
find out more?
My partner and I absolutely love your blog and find the majority of your post’s to be just what I’m looking for. Would you offer guest writers to write content for you personally? I wouldn’t mind producing a post or elaborating on a few of the subjects you write regarding here. Again, awesome website!
http://vsebanki-rf.ru/bank-otkryitie-vkladyi-fizicheskih-lits-v-rublyah-na-2016-god/
купить диплом в сочи
http://kdcmir.ru
купить диплом в саратове
Hi, i believe that i noticed you visited my weblog thus i came
to go back the prefer?.I’m attempting to in finding issues to enhance my
website!I guess its good enough to use some of your concepts!!
I am no longer sure where you are getting your info, but good topic.
I needs to spend some time learning more or figuring out more.
Thanks for magnificent information I used to be looking for this information for my mission.
Saved as a favorite, I like your site!
Thank you a bunch for sharing this with all people you really understand what you’re talking approximately! Bookmarked. Kindly additionally seek advice from my site =). We may have a hyperlink alternate arrangement between us
https://blog.kazakh-zerno.net/2024/05/31/kupit-diplom-v-moskve-bystroe-oformlenie.html
https://forum.dalmatin-club.ru/topic/26719-%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC-%D0%BE-%D1%81%D1%80%D0%B5%D0%B4%D0%BD%D0%B5%D0%BC-%D1%81%D0%BF%D0%B5%D1%86%D0%B8%D0%B0%D0%BB%D1%8C%D0%BD%D0%BE%D0%BC-%D0%BE%D0%B1%D1%80%D0%B0%D0%B7%D0%BE%D0%B2%D0%B0%D0%BD%D0%B8%D0%B8-%D1%86%D0%B5%D0%BD%D0%B0-j946l/
http://skvortsy.listbb.ru/viewtopic.php?f=31&t=1658
https://bravelight.net/profile.php?op=userinfo&username=minna_prout_301152&com=profile
http://allrealtor.ru/threads/665/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/uglycat1959/profile
https://permacultureglobal.org/users/58887-ahmed-glow
https://launchpad.net/~xuto19501
https://rentry.org/vqhuoqoz
https://www.obesityhelp.com/members/truejkeee19691953/about_me/
Fastidious response in return of this query
with genuine arguments and describing the whole thing regarding that.
I really like what you guys are up too. This kind of
clever work and reporting! Keep up the amazing works guys I’ve included you guys
to my personal blogroll.
Hello there! I simply wish to offer you a huge thumbs up for the excellent info you’ve got right
here on this post. I am coming back to your web site for more
soon.
Hi, after reading this remarkable paragraph i am
as well cheerful to share my know-how here with friends.
A person essentially help to make significantly posts I would state.
That is the first time I frequented your web page and so far?
I amazed with the research you made to create this particular submit amazing.
Wonderful activity!
I am not sure where you’re getting your info, but good topic.
I needs to spend some time learning much more or understanding more.
Thanks for great info I was looking for this info for my mission.
Simply want to say your article is as amazing. The clarity on your publish is simply cool and i could assume you are knowledgeable in this subject. Fine with your permission let me to grasp your feed to stay updated with approaching post. Thank you one million and please continue the gratifying work.
купить диплом в сыктывкаре
http://wiki.mdomtv.net/index.php?title=пїЅпїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅ:_пїЅпїЅпїЅ_пїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅ_пїЅ_пїЅпїЅпїЅпїЅпїЅ_пїЅпїЅпїЅпїЅпїЅпїЅпїЅ!
https://thefishbowled.com/userinfo.php?com=profile&from=space&username=eunice_rowley_301152
http://fremontrp.listbb.ru/viewtopic.php?f=18&t=351
http://bestcoolfun.ru/kupit-diplom-poshagovoe-rukovodstvo
http://finttech.ru/kupit-diplom-v-moskve-operativnoe-oformlenie
купить диплом в перми
Hello to all, how is the whole thing, I think every one is getting more from this site, and your views are pleasant for new people.
de-gustare.it/brasato-di-bue-grasso-birra-miele-e-cacao-amaro/
usaessayexperts.com/faq/
myanmarfootball.org/gallery/displayimage.php?album=lastup&cat=6&pos=0
bbs.17dra.com/home.php?mod=space&uid=225766
fabnews.ru/forum/showthread.php?p=76998
I’m curious to find out what blog system you happen to be utilizing?
I’m experiencing some small security issues with my latest blog and I would like to find something
more safe. Do you have any solutions?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/s9cndjaf
https://haveagood.holiday/users/352131
https://www.obesityhelp.com/members/farwa1978/about_me/
https://anotepad.com/notes/bj78htnb
https://permacultureglobal.org/users/58724-maria-morales
Your means of describing all in this paragraph is
genuinely nice, every one can simply understand it,
Thanks a lot.
?? Забирай 100,000 Рублей + 400 Фриспинов в виде Welcome бонуса, на официальном сайте 7К Казино ??
?????? Официальное зеркало ??????
7к казино скачать
НОВЫЕ ПРОМОКОДЫ ОТ НАШЕГО ТЕЛЕГРАМ КАНАЛА 7К КАЗИНО! МОЩНЫЙ СТАРТ НАШИМ ИГРОКАМ. ???? ЖЕЛАЕМ КРУТЫХ ЗАНОСОВ ??
Joker7070 – бонус 170%+ 70 FS в Joker Stoker
777WIN – бонус 100%+ 30 FS в Hot Triple Sevens
HOT100 – бонус 100 FS в Indiana’sQuest
?? Добавляйте промокод в разделе бонусов на официальном сайте 7К Казино.
?? Важно! Для новых игроков после регистрации 7K Casino приготовил Welcome бонус до 100,000 Рублей + 400 Фриспинов! Успейте активировать ???
?? Подробная информация на официальном сайте 7К в разделе бонусов! Каждый промокод можете использовать по очереди.
7k casino рабочее зеркало
https://t.me/s/casino_7k_official
Hmm is anyone else encountering problems with the images on this
blog loading? I’m trying to figure out if its a problem on my end or if it’s the blog.
Any feedback would be greatly appreciated.
Hello would you mind letting me know which
web host you’re utilizing? I’ve loaded your blog in 3 different web browsers and I must say this blog loads a
lot quicker then most. Can you recommend a good hosting provider at a
fair price? Thanks, I appreciate it!
Добрый день всем!
Приобретите российский диплом по выгодной цене с гарантией прохождения проверок и доставкой в любой город РФ!
http://www.sebastiafarre.com/ref-439/#comment-1357547
https://mediamemorial.ru/club/user/69594/forum/message/2855/12338/
http://www.uzaybilim.net/2014/01/gunberi-ve-gunote-perihel-ve-afel.html
http://trudobudny.blogspot.com/2012/02/microsoft-volume-licensing-service.html
http://worldpeacevolunteers.org/2020/03/18/world-peace-volunteers-campaign-for-peaceful-election-2008/#comment-540831
This site was… how do I say it? Relevant!! Finally I’ve found something that helped
me. Kudos!
Superb post however I was wondering if you could write a litte more on this
subject? I’d be very grateful if you could elaborate a
little bit further. Bless you!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/z77awmva
https://launchpad.net/~sdsw19501
https://haveagood.holiday/users/351379
https://cannabis.net/user/151171
https://www.hentai-foundry.com/user/bezzzumna1957/profile
This site was… how do I say it? Relevant!! Finally I’ve found something which helped me.
Thanks a lot!
Howdy! I know this is kinda off topic however I’d figured I’d ask. Would you be interested in exchanging links or maybe guest writing a blog article or vice-versa? My blog goes over a lot of the same subjects as yours and I believe we could greatly benefit from each other. If you happen to be interested feel free to send me an email. I look forward to hearing from you! Wonderful blog by the way!
http://shoelovershub.com/read-blog/504_we-buy-a-diploma-on-the-internet-expert-recommendations.html?mode=day
купить диплом мастера маникюра и педикюра
http://paradise-kursk.ru
купить диплом в ухте
Greetings I am so grateful I found your web site, I really found you by mistake, while I was researching on Google for something else, Anyhow I am here now and would just like to say cheers for a tremendous post and a all round interesting blog (I also love the theme/design), I don’t have time to browse it all at the minute but I have saved it and also added in your RSS feeds, so when I have time I will be back to read much more, Please do keep up the excellent jo.
ros-kruiz.ru/
jogibaba.com/blogs/32/Why-is-the-popularity-of-higher-education-rapidly-declining-now?lang=ru_ru
vsekruizi.ru/Special_offers/thailand_from_krasnodar/
http://www.bryners.ru/forum/memberlist.php?sk=c&sd=d&start=625
mcmon.ru/showthread.php?tid=35215
whoah this weblog is great i love studying your posts. Stay up the great work! You know, lots of individuals are hunting round for this information, you can aid them greatly.
купить диплом образцы
http://realtyintellect.ru/kupit-diplom-legalno-i-bezopasno
http://medik-look.ru/kupit-uchebnyie-svidetelstva
https://ivebo.co.uk/read-blog/28573
http://power.ekafe.ru/viewtopic.php?f=2&t=1643
http://allsportime.ru/individualnoe-obrazovanie-gde-kupit-diplom-sozdannyiy-spetsialno-dlya-tebya
купить диплом физика
I know this web page offers quality dependent content and other stuff,
is there any other web page which provides such things in quality?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/pohhbsc19751971
https://www.hentai-foundry.com/user/mormitor1963/profile
https://rentry.org/znb6dedw
http://www.nfomedia.com/profile?uid=rOjQZcG
https://www.hentai-foundry.com/user/moneybank1966/profile
Заявление Виктории Набойченко – бывшей супруги главного свидетеля обвинения Евгения Набойченко
https://www.youtube.com/watch?v=Q0smZ5_QOcU
Здравствуйте, меня зовут Виктория Набойченко.
Я являюсь бывшей супругой Набойченко Евгения Витальевича, с которым состояла в официальном браке с 2017 по 2022 год. Из средств массовой информации мне стало известно о фактах уголовного преследования лиц из окружения Романа Василенко, который является президентом международной бизнес-карты на территории России, Казахстана и Киргизии. На протяжении 20 лет Евгений Набойченко и Роман Василенко дружили, вместе путешествовали, проводили время, ездили на рыбалку. Роман очень доверял Евгению, делился с ним своими идеями, планами на коммерческую деятельность и реализацию новых проектов. Мне известно, что Роман предложил Евгению разработать фирменный логотип и сайт компании Life is Good, а впоследствии возглавить IT-отдел ЖК. Евгений с радостью согласился на предложение Романа и возглавил IT-отдел. Роман достойно оплачивал работу Евгения, гонорар которого вырос со 100,000 рублей до 500,000 рублей в месяц, что было связано с ростом и развитием кооператива Best. Роман всегда был щедр к Евгению и ценил его работу, за что ежегодно премировал.
Тот успех, которого достиг Роман за эти годы, очень задевал Евгения как мужчину. В ходе совместной жизни Евгений делился со мной, что мечтает однажды превзойти Романа в бизнесе, а его проект стать круче Илона Маска. Примерно с июня 2021 года Евгений стал нервным и злился на Романа, рассказывая мне, что Роман не хочет заплатить ему 170,000 евро. Я не вдавалась в подробности их взаимоотношений. Чуть позже я узнала о письмах Евгения к Роману, в которых он требовал от него указанную сумму и угрожал увечьями и убийством Роману и всем членам его семьи, в том числе и детям. Данные угрозы он отправлял голосовыми сообщениями и в электронных письмах. Мне известно, что Роман не реагировал на угрозы Евгения, что очень злило его, потому что он ждал от него указанную сумму. В моём присутствии Евгений постоянно по нескольку раз переслушивал свои голосовые сообщения, которые с угрозами отправлял в адрес Романа Василенко. Он получал от этого огромное удовольствие, с ухмылкой хвастаясь, что Роман обязательно заплатит.
[url=https://www.youtube.com/watch?v=Q0smZ5_QOcU]Best Way[/url]
17 февраля 2022 года в 7:00 по МСК в нашем доме в посёлке в Краснодарском крае произошел обыск. В доме находилась только я и дети, а Евгений в это время находился в Краснодаре на съёмной квартире, куда заблаговременно уехал, предупредив меня за 3 дня о предстоящем обыске в доме. Как будто его друзья из правоохранительных органов Петербурга предупредили его. На мои вопросы о том, что происходит, он отвечал, что это формальность и что мне не о чем беспокоиться. Позже мне стало известно, что подобные обыски проходили и у других сотрудников кооператива в Санкт-Петербурге, Краснодаре, Анапе и других городах. В обыске нашего дома принимали участие шесть оперативных работников ОББ города Санкт-Петербурга, фамилии которых указаны в протоколе обыска, копия которого имеется в моём распоряжении. В ходе обыска оперативные работники оскорбляли Евгения и Романа, называя их преступниками и мошенниками, а также иными оскорбительными словами. При обыске был изъят планшет Евгения, оперативные работники перевернули весь дом и напугали детей, а также пообещали, что задержат Евгения в тот же день.
17 февраля 2022 года Евгений был задержан сотрудниками оперативных служб в городе Краснодар и примерно через 3 дня был доставлен в Санкт-Петербург, где был допрошен в УП по рекомендации сотрудников полиции. Допрос происходил без участия его личного адвоката Ирины Черняховской, от услуг которой он отказался в тот же момент. Мне позвонила Ирина в бешенстве и недоумении, крича о том, что Евгений её опозорил перед сотрудниками полиции, отказавшись от её услуг в грубой форме. Впоследствии мне стало известно от Евгения, что он лично получил покровительство начальника ОББ города Санкт-Петербурга в своей безнаказанности по факту проводимого расследования. Об этом он похвастался мне в личном разговоре, будучи окрылённый поддержкой начальника ОББ, который с ним разговаривал на доверительной ноте.
Евгений мне рассказал, что получит от органов абсолютную неприкосновенность и поддержку за информацию, которая интересовала оперативных работников, за что Евгению также была предоставлена госохрана. В период с февраля по середину апреля Евгений находился в Питере под госзащитой и плотно взаимодействовал с правоохранительными органами. Он предоставлял им всю информацию, в том числе полный доступ к системе базы данных Best Way и Life is Good. Евгений инициативно предлагал органам заблокировать доступ к личным кабинетам пайщиков и членов ЖК БСТ, чем полностью парализовал хозяйственную деятельность кооператива, создавая напряжённость и негативные отзывы. Также Евгений активно призывал граждан к написанию заявлений в отношении Романа Василенко с целью возврата пайщиками своих денежных средств, делал он это через каналы Telegram и YouTube. Евгений был единственным, кто имел и имеет доступ к базам данных и ключам. Желая намеренно создать проблемы Роману, он отказывается восстанавливать работу системы Best Way при наличии такой возможности.
Миссия Евгения – это привлечение Романа Василенко к уголовной ответственности, что спровоцирует крах компании и полное прекращение её деятельности, а это в свою очередь 30,000 пайщиков, которые могут стать потерпевшими по уголовному делу. Всё это мне известно из личного общения с Евгением, когда я приезжала к нему в Санкт-Петербург на несколько дней. Мне также достоверно известно, что в настоящее время нет реальных заявлений от недовольных пайщиков, так как в Романа Василенко все верят. Самое главное, что у Евгения есть возможность в любой момент через свой личный ноутбук восстановить работу ЖК Best Way и вернуть доступ к личным кабинетам пайщиков. Однако он этого не делает, преследуя свои личные цели.
What i do not realize is in fact how you are now not actually a
lot more smartly-appreciated than you may be now. You are so
intelligent. You know thus significantly in relation to this topic, produced me for
my part believe it from a lot of various
angles. Its like women and men are not fascinated until
it is something to do with Woman gaga! Your
individual stuffs nice. At all times maintain it up!
Hi just wanted to give you a quick heads up and let you know a few of the pictures aren’t loading correctly. I’m not sure why but I think its a linking issue. I’ve tried it in two different web browsers and both show the same outcome.
купить диплом в балаково
https://re-port.ru/users/51485/
https://rabota.ys.tj/viewtopic.php?id=9345#p18404
https://faithstreamer.com/blogs/2366/пїЅпїЅпїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅпїЅ-пїЅ-пїЅпїЅпїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅ-пїЅпїЅпїЅпїЅпїЅпїЅ-пїЅпїЅпїЅпїЅпїЅпїЅ-пїЅ-пїЅпїЅпїЅпїЅпїЅпїЅпїЅ
http://worldavtonew.ru/unikalnoe-obrazovanie-priobreti-diplom-kotoryiy-vyidelyaet-tebya-sredi-drugih
http://scientistsufo.ru/diplom-po-vashim-zhelaniyam-vyibirayte-zakazyivayte-dostigayte-uspeha
купить диплом в энгельсе
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiYadH
https://www.haikudeck.com/presentations/PEEJcK9CNU
http://www.babelcube.com/user/john-year
https://aladik1965.diary.ru/
https://rentry.org/pxncfg3n
Great site you have here but I was curious about if you knew of any message boards that
cover the same topics discussed in this article?
I’d really like to be a part of community where I can get suggestions
from other experienced individuals that share the same interest.
If you have any suggestions, please let me know.
Kudos!
Pretty section of content. I just stumbled upon your web site and in accession capital to assert that I acquire in fact enjoyed account your blog posts. Any way I will be subscribing to your feeds and even I achievement you access consistently quickly.
foro.turismo.org/post281274.html
alpha.prime-pc.md/good_info/27721
shop2.rybolov.de/index.php?links_exchange=yes&page=260&show_all=yes
politictoday.ru/page/39
benhvienthammyasean.com/cau-chuyen-asean
I got this web page from my pal who shared with me regarding this site and now this time
I am visiting this web page and reading very informative
articles here.
What’s up to all, it’s in fact a good for me to pay a visit this
web site, it includes valuable Information.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/brent166b
https://www.hentai-foundry.com/user/gfsrrg1972/profile
https://www.haikudeck.com/presentations/qDk91Ftn1C
https://www.obesityhelp.com/members/tjfyk19921951/about_me/
https://rentry.org/yubsrc7m
Hi, Neat post. There is an issue together with your
website in web explorer, could test this? IE nonetheless is the market
chief and a big section of people will miss your magnificent writing due to this problem.
Exceptional post however I was wondering if you could write a litte more on this topic?
I’d be very thankful if you could elaborate a little bit further.
Kudos!
Nice weblog right here! Additionally your web site lots up fast!
What host are you using? Can I get your affiliate hyperlink in your host?
I desire my site loaded up as fast as yours lol
Nice post. I used to be checking constantly this blog
and I am inspired! Extremely useful information specifically the ultimate phase 🙂 I take care of
such information a lot. I used to be looking for this particular information for
a long time. Thanks and best of luck.
https://rentyachtsincyprus.com/en – hashish with a syringe 3 gram sleva. 35%
https://rentyachtsincyprus.com/en – powdered heroin – snorting 100 gram sale 22%
https://rentyachtsincyprus.com/en – ecstasy under the tongue – 2 gram sale 22%
https://rentyachtsincyprus.com/en – hachis con una jeringuilla 51 grama sleva. 38%
https://rentyachtsincyprus.com/en – heroina en polvo – esnifada 100 grama rebajas 42%
https://rentyachtsincyprus.com/en – extasis bajo la lengua – 2 gramas sleva 16%
https://rentyachtsincyprus.com/en – hasista ruiskulla 3 grams discounts. 23%
https://rentyachtsincyprus.com/en – heroiinijauhe – nuuskaaminen 30 gram discount 26%
https://rentyachtsincyprus.com/en – ekstaasi kielen alla – 2 gram sleva 21%
https://rentyachtsincyprus.com/en – haszysz za pomoca strzykawki 17 gram sale. 11%
https://rentyachtsincyprus.com/en – sproszkowana heroina – wciaganie 30 gram discount 47%
https://rentyachtsincyprus.com/en – ekstaza pod jezykiem – 11 gram sleva 14%
https://rentyachtsincyprus.com/en – du haschisch avec une seringue 3 grammes discounts. 23%
https://rentyachtsincyprus.com/en – heroine en poudre – sniffer 30 grammes discounts 14%
https://rentyachtsincyprus.com/en – ecstasy sous la langue – 30 grammes discount 43%
My spouse and I absolutely love your blog and find most
of your post’s to be just what I’m looking for.
can you offer guest writers to write content to suit your needs?
I wouldn’t mind publishing a post or elaborating on a number of the subjects you write related to here.
Again, awesome weblog!
I simply couldn’t leave your website before suggesting
that I actually loved the standard information an individual supply to your visitors?
Is gonna be again ceaselessly in order to inspect new posts
Howdy very nice web site!! Guy .. Beautiful .. Wonderful
.. I will bookmark your website and take the feeds also?
I’m glad to seek out a lot of useful information here in the post, we want develop extra techniques in this
regard, thank you for sharing. . . . . .
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3116935.html
https://anotepad.com/notes/ndixx639
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196848
https://chyoa.com/user/klassi1978
https://www.quia.com/profiles/mary440davis
What a data of un-ambiguity and preserveness of valuable know-how regarding unpredicted emotions.
https://baidubookmark.com/story16991160/%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B0%D1%82%D1%82%D0%B5%D1%81%D1%82%D0%B0%D1%82
http://booksshare.net/
https://drunkslut.com/
https://bookmarkinglog.com/story17080664/%D0%BA%D1%83%D0%BF%D0%B8%D1%82%D1%8C-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC-%D0%B2-%D0%9A%D0%B0%D0%B7%D0%B0%D0%BD%D0%B8
https://howtobuyessay.com/
I every time spent my half an hour to read this blog’s articles or reviews everyday along with a mug of coffee.
Hey there! This is my 1st comment here so I just wanted to give a quick shout out and tell you I truly enjoy reading through your posts. Can you recommend any other blogs/websites/forums that cover the same topics? Thanks a lot!
купить диплом в орле
https://hikariacademy.edu.vn/top-10-cau-hoi-va-tra-loi-tieng-nhat-p1/
https://spbgasu.topf.ru/viewtopic.php?id=250#p318
http://gamemotors.ru/kupit-diplom-rf-v-moskve-uskorennoe-oformlenie-i-garantii
http://bezgrani4nyemyryi.anihub.me/viewtopic.php?id=627#p11806
http://sampnewrp.listbb.ru/viewtopic.php?f=18&t=443&sid=b0c792af6aef133c75df7b4e2a3236fe
купить диплом в брянске
Hi there this is somewhat of off topic but I was wondering if
blogs use WYSIWYG editors or if you have to manually code
with HTML. I’m starting a blog soon but have no coding
skills so I wanted to get guidance from someone with experience.
Any help would be greatly appreciated!
купить диплом кандидата наук diplomvash.ru .
It’s really a nice and useful piece of info.
I’m glad that you just shared this useful information with us.
Please stay us up to date like this. Thanks for sharing.
Your style is unique compared to other folks I have read stuff from.
I appreciate you for posting when you’ve got the opportunity, Guess I will
just bookmark this page.
hello there and thank you for your information – I’ve definitely picked up anything
new from right here. I did however expertise several
technical points using this web site, since I experienced to reload the web site lots
of times previous to I could get it to load properly. I had been wondering if
your web host is OK? Not that I’m complaining, but slow loading instances times will
often affect your placement in google and can damage your high-quality score if ads and marketing with Adwords.
Well I’m adding this RSS to my e-mail and could look out for
a lot more of your respective exciting content. Make sure
you update this again soon.
Wow that was unusual. I just wrote an really long comment but after I clicked submit my comment didn’t show up. Grrrr… well I’m not writing all that over again. Regardless, just wanted to say great blog!
myteana.ru/forums/index.php?autocom=gallery&req=si&img=4490
http://www.forumklassika.ru/member.php?u=169637&tab=aboutme
tmwip-chelm.org.pl/profile.php?lookup=16720
art-gymnastics.ru/users/54?page=28
angelteam.uv.ro/profile.php?mode=viewprofile&u=207852
What’s Happening i’m new to this, I stumbled upon this I’ve found It absolutely helpful and it has aided me out loads.
I am hoping to contribute & help different users like its helped me.
Good job.
What a stuff of un-ambiguity and preserveness
of precious experience about unexpected emotions.
This post gives clear idea in favor of the new people of blogging,
that in fact how to do blogging and site-building.
If you desire to obtain much from this post then you
have to apply such strategies to your won webpage.
My brother recommended I might like this web site. He was
entirely right. This post actually made my day.
You cann’t imagine simply how much time I had spent for this information!
Thanks!
Hello there! Quick question that’s totally off topic.
Do you know how to make your site mobile friendly? My weblog looks weird
when viewing from my iphone4. I’m trying to find a theme or plugin that might be able to correct this
issue. If you have any recommendations, please share. Many thanks!
Do you have a spam problem on this site; I also am a blogger, and I was wondering your situation; we have developed some nice methods and we are looking to exchange strategies with other folks, be sure to shoot me an email if interested.
http://www.paladiny.ru/forummess.dwar.php?TopicID=26048&Offset=0
shoelovershub.com/read-blog/475_why-is-the-popularity-of-universities-declining-in-our-time.html?mode=day
http://www.camnangbenh.com/mu-mau/
brgdiamond.vn/en/services-residential/
onlineboxing.net/jforum/user/profile/287687.page
This site really has all the information and facts I needed concerning this subject and didn’t know who to ask.
If some one desires to be updated with newest technologies then he must be visit this site and be up to date all the time.
Article writing is also a fun, if you know after that you can write if not it is
difficult to write.
I know this site provides quality based content and other
information, is there any other web site which offers these things
in quality?
This is a topic which is near to my heart… Many thanks!
Exactly where are your contact details though?
Great article, totally what I needed.
It’s perfect time to make some plans for the future and it’s time to
be happy. I have read this post and if I could I wish to suggest you few
interesting things or suggestions. Maybe you could write next articles referring to this article.
I wish to read more things about it!
Quality articles or reviews is the crucial to be a focus for the
people to go to see the site, that’s what this site is providing.
https://Dr-nona.ru – Dr. Nona
I blog quite often and I truly appreciate your information. Your article has truly peaked my interest.
I will book mark your blog and keep checking for new details
about once a week. I subscribed to your RSS feed too.
Howdy I am so happy I found your weblog, I really found
you by mistake, while I was browsing on Google for something else, Nonetheless I am here now
and would just like to say kudos for a marvelous post and a all round exciting blog (I also love the theme/design), I don’t have time to browse it all at the moment but I have saved it and also added in your RSS feeds,
so when I have time I will be back to read much more, Please do keep
up the fantastic b.
오공 슬롯
화가 났지만 Hongzhi 황제는 여전히 상냥했습니다.
Wow, fantastic blog layout! How long have you been blogging for?
you make blogging look easy. The overall look of your website is
excellent, let alone the content!
It’s a pity you don’t have a donate button! I’d definitely donate
to this brilliant blog! I suppose for now i’ll settle for
book-marking and adding your RSS feed to my Google account.
I look forward to brand new updates and will share this site with my Facebook group.
Talk soon!
I’m truly enjoying the design and layout of your website.
It’s a very easy on the eyes which makes it much more pleasant for me to come here
and visit more often. Did you hire out a designer to create your theme?
Exceptional work!
Having read this I believed it was really enlightening.
I appreciate you taking the time and effort to put this short article together.
I once again find myself spending a lot of time
both reading and leaving comments. But so what,
it was still worth it!
Appreciate it about this subject. I learned a lot from the content. It’s informative and something I’ll revisit. Don’t forget to explore đăng ký 79king for more related content.
Nice response in return of this issue with firm arguments and
explaining everything regarding that.
Hey I am so grateful I found your weblog, I really found you by accident, while I was browsing on Digg for something else, Anyways I am here now and would just like to say cheers for a remarkable post and a all round entertaining blog (I also love the theme/design), I don’t have time to browse it all at the moment but I have book-marked it and also included your RSS feeds, so when I have time I will be back to read a great deal more, Please do keep up the awesome b.
smartwatchcolombia.com/producto/78/smartwatch-serie-8-ultra-11-2023–manilla-de-regalo/7arusak-diploms.com
blagmama.ru/forum/index.php?autocom=blog&blogid=4350&showentry=11569
wine.historic.ru/books/c0007_1.shtml
bizkerrefutboltaldea.com/bizkerre/Euskara/masculino/juvenil.htm
realtyintellect.ru/page/39
I believe that is among the so much significant info for me.
And i am glad reading your article. But want to commentary
on some basic things, The website taste is ideal,
the articles is really excellent : D. Excellent process, cheers
Thanks for the marvelous posting! I quite enjoyed
reading it, you could be a great author. I will be sure to bookmark your blog and will
eventually come back at some point. I want to encourage one to continue your great posts, have a nice
afternoon!
Thanks a bunch for sharing this with all of us you actually understand what
you are talking about! Bookmarked. Please also talk over with my
website =). We may have a link trade agreement among us
This article is captivating about this topic. I am quite interested in your perspective. It’s informative and definitely worth checking out. Don’t forget to explore sân chơi 79king for more related content.
Thank you about the information provided. I really enjoyed what you’ve written. It’s insightful and definitely worth checking out. Don’t forget to check out vào 79king không bị chặn for more related content.
You actually make it seem so easy with your presentation but I find this topic to be actually something which I think I would never understand.
It seems too complicated and very broad for me.
I’m looking forward for your next post, I’ll try to get
the hang of it!
Very good post. I am dealing with some of these issues as well..
Thanks a bunch about the insights shared. I truly appreciated what you’ve written. It’s insightful and definitely worth checking out. Don’t forget to explore vào 79king for more related content.
Excellent about what you’ve provided. I am highly interested in what you’ve written. It’s thought-provoking and something I’ll revisit. Don’t forget to explore vào 79king for more related content.
This was quite informative. More at ##anyKeyword##.
Thanks for the insightful write-up. More like this at review nhà cái .
This is the perfect site for everyone who hopes to understand this topic. You understand so much its almost hard to argue with you (not that I actually will need to…HaHa). You certainly put a brand new spin on a subject that’s been written about for years. Excellent stuff, just wonderful!
renebiemans.nl/users.php?m=details&id=31723
demo2.weboss.hk/1009/comment/html/?577.html
autogroupe.ru/page/10
gamemotors.ru/page/11
shop005.getmall.kr/board/board.php?pagetype=view&num=245&board=customerqna&block=0&gotopage=1&search=&s_check=
It’s a pity you don’t have a donate button! I’d certainly donate to this outstanding blog!
I suppose for now i’ll settle for book-marking and adding your RSS
feed to my Google account. I look forward to new updates and will talk about
this blog with my Facebook group. Talk soon!
This is very insightful. Check out top nhà cái uy tín nhất hiện nay for more.
This was quite useful. For more, visit Nhà cái uy tín số 1 .
With havin so much written content do you ever
run into any problems of plagorism or copyright violation? My
blog has a lot of completely unique content I’ve either
created myself or outsourced but it appears a lot of it is popping it
up all over the internet without my permission. Do you know any ways to help
reduce content from being stolen? I’d definitely appreciate
it.
I enjoyed this post. For additional info, visit review nhà cái .
Wonderful tips! Discover more at list nhà cái uy tín .
Awesome post.
Great job! Find more at Nhà cái uy tín số 1 .
Every weekend i used to go to see this web site, because i wish for enjoyment, since this this website conations in fact good funny information too.
This is very fascinating, You’re a very professional blogger.
I’ve joined your rss feed and look forward to in the
hunt for more of your wonderful post. Also, I have shared your site in my
social networks
Hi colleagues, its great piece of writing on the topic of educationand completely defined,
keep it up all the time.
This was quite helpful. For more, visit top nhà cái uy tín nhất hiện nay .
Attractive component of content. I simply stumbled upon your web
site and in accession capital to assert that I get actually enjoyed account your blog
posts. Any way I will be subscribing for your feeds or even I fulfillment you access
constantly rapidly.
Hello, I log on to your blogs like every week.
Your humoristic style is awesome, keep doing what you’re doing!
Greetings! This is my first visit to your blog! We are a group of volunteers
and starting a new initiative in a community in the same niche.
Your blog provided us valuable information to work on. You have done a extraordinary job!
Appreciate the great suggestions. For more, visit link chính thức của cwin .
Thanks for the helpful article. More like this at xem thêm tại go99 link chính thức .
I am curious to find out what blog system
you have been using? I’m having some minor security issues
with my latest blog and I’d like to find something more safe.
Do you have any suggestions?
This is the right site for everyone who wants to understand this topic.
You know so much its almost hard to argue with you (not that I actually would
want to…HaHa). You certainly put a brand new spin on a
subject that has been written about for decades. Excellent stuff, just excellent!
Appreciate the comprehensive advice. For more, visit trang tỷ lệ kèo .
Please let me know if you’re looking for a author for your blog.
You have some really great articles and I believe I would be a good asset.
If you ever want to take some of the load off, I’d really like to write some content for your blog in exchange for a link
back to mine. Please send me an email if interested. Many thanks!
Excellent post. I will be experiencing many of these issues as well..
This design is wicked! You definitely know how to keep a reader entertained.
Between your wit and your videos, I was almost moved to start my
own blog (well, almost…HaHa!) Fantastic job. I really enjoyed
what you had to say, and more than that, how you presented it.
Too cool!
This article will assist the internet people for setting
up new webpage or even a blog from start to end.
Howdy! I’m at work surfing around your blog from my
new iphone 4! Just wanted to say I love reading your blog and look forward to all your posts!
Keep up the outstanding work!
Right here is the right blog for anybody who wishes to understand this topic. You know a whole lot its almost hard to argue with you (not that I really will need to…HaHa). You certainly put a brand new spin on a subject that’s been discussed for years. Excellent stuff, just wonderful!
azat.on.kg/blogs/451/%D0%A1%D0%BA%D0%BE%D0%BB%D1%8C%D0%BA%D0%BE-%D1%81%D1%82%D0%BE%D0%B8%D1%82-%D1%81%D0%B5%D0%B9%D1%87%D0%B0%D1%81-%D0%BA%D0%B0%D1%87%D0%B5%D1%81%D1%82%D0%B2%D0%B5%D0%BD%D0%BD%D1%8B%D0%B9-%D0%B4%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC
krutovo.ru/index.php?name=Account&op=info&uname=ymonozav
эвакуатор-срочно-недорого.рф/
istoriya-kino.ru/kinematograf/alph0015.shtml
tuning-performance.ru/polirovka-kuzova-avtomobilya/
I like what you guys are usually up too. This kind of clever work and reporting!
Keep up the amazing works guys I’ve included
you guys to my personal blogroll.
В нашем мире, где диплом – это начало успешной карьеры в любой отрасли, многие ищут максимально простой путь получения качественного образования. Факт наличия официального документа сложно переоценить. Ведь именно диплом открывает двери перед любым человеком, который желает начать трудовую деятельность или продолжить обучение в любом ВУЗе.
В данном контексте наша компания предлагает быстро получить любой необходимый документ. Вы имеете возможность приобрести диплом, и это становится отличным решением для всех, кто не смог закончить обучение, утратил документ или желает исправить плохие оценки. диплом изготавливается с особой тщательностью, вниманием к мельчайшим элементам. В итоге вы сможете получить 100% оригинальный документ.
Преимущество этого решения заключается не только в том, что можно максимально быстро получить диплом. Весь процесс организовывается удобно, с нашей поддержкой. От выбора нужного образца до консультаций по заполнению персональных данных и доставки по стране — все будет находиться под абсолютным контролем опытных специалистов.
Всем, кто пытается найти оперативный способ получить необходимый документ, наша компания предлагает выгодное решение. Заказать диплом – значит избежать долгого обучения и сразу перейти к важным целям: к поступлению в университет или к началу успешной карьеры.
http://intsms.ru
http://schoolnumberone.ru
http://sibsocio.ru
http://kontur-plus.ru
http://park-robotov.ru
Such innovation! capcut pro
I’m not sure exactly why but this weblog is loading
incredibly slow for me. Is anyone else having this problem or is it a issue on my end?
I’ll check back later on and see if the problem still exists.
excellent points altogether, you simply received a emblem new reader.
What could you recommend in regards to your publish that you just
made some days ago? Any certain?
I’m not that much of a online reader to be honest but your blogs really nice, keep it up!
I’ll go ahead and bookmark your site to come back later.
All the best
I do accept as true with all of the ideas you’ve offered for your post.
They are very convincing and can certainly work.
Still, the posts are very quick for novices. May you please prolong them
a little from next time? Thank you for the post.
Hello, i read your blog occasionally and i
own a similar one and i was just curious if you get a lot of spam feedback?
If so how do you prevent it, any plugin or anything you can advise?
I get so much lately it’s driving me insane so any
help is very much appreciated.
Hey! This is my 1st comment here so I just wanted to give a
quick shout out and say I really enjoy reading your blog posts.
Can you recommend any other blogs/websites/forums that deal with the same subjects?
Thanks!
After I initially left a comment I appear to have clicked the -Notify me when new comments are added- checkbox and now each time a comment is added I receive 4 emails with the exact same comment.
Perhaps there is a means you are able to remove
me from that service? Kudos!
I love what you guys tend to be up too. This kind of clever
work and reporting! Keep up the terrific works guys I’ve incorporated you guys
to my personal blogroll.
Hi there, of course this piece of writing is really pleasant and I have learned lot
of things from it on the topic of blogging. thanks.
This information is invaluable. Where can I find out more?
http://www.camry-club.ru/member.php/61231-sonnick84?tab=activitystream&type=user&page=2
http://www.consult-exp.com/blogs/116058/%D0%94%D0%B8%D0%BF%D0%BB%D0%BE%D0%BC%D1%8B-%D0%BD%D0%B0-%D0%BB%D1%8E%D0%B1%D0%BE%D0%B9-%D0%B2%D0%BA%D1%83%D1%81-%D0%B8-%D1%86%D0%B2%D0%B5%D1%82-%D1%88%D0%B8%D1%80%D0%BE%D0%BA%D0%B8%D0%B9-%D0%B0%D1%81%D1%81%D0%BE%D1%80%D1%82%D0%B8%D0%BC%D0%B5%D0%BD%D1%82?lang=tr_tr
http://www.exoltech.com/wall/blogs/user/alanpoe?&page=1
forum.l2star.net/member.php?u=17186
programasites.com.br/blog/post/11666/e-mail-marketing-boas-praticas-e-temas-para-abordar
Hello there, just became alert to your blog through Google, and found that it is truly informative.
I am going to watch out for brussels. I will appreciate if you continue this
in future. Numerous people will be benefited from your writing.
Cheers!
Hey! Would you mind if I share your blog with my facebook
group? There’s a lot of folks that I think would really
enjoy your content. Please let me know. Thanks
Great goods from you, man. I have understand your stuff previous
to and you are just extremely wonderful.
I really like what you have acquired here, certainly like what you’re saying and
the way in which you say it. You make it enjoyable and you still care
for to keep it wise. I can’t wait to read far more from you.
This is actually a tremendous web site.
Wow that was odd. I just wrote an very long comment but after I clicked submit my comment didn’t show up. Grrrr… well I’m not writing all that over again. Anyways, just wanted to say excellent blog!
http://bike.by/forum/viewtopic.php?p=178124
купить диплом в томске
http://copoko.ru
купить диплом в каспийске
I blog often and I really thank you for your information. This
great article has really peaked my interest. I will book mark your website and
keep checking for new information about once a week.
I opted in for your RSS feed too.
Tuy nhiên, điều đó không có nghĩa là chúng tôi phải ưu ái cho các sản phẩm của Accesstrade, Adflex hay bất kỳ công ty nào khác. Cách chúng tôi đánh giá sản phẩm đều dựa trên tiêu chuẩn khách quan và chân thực nhất. Mặc dù đội ngũ biên tập của chúng tôi chèn liên kết vào các trang nhưng cách đánh giá
My brother recommended I might like this website. He was entirely right.
This post actually made my day. You can not imagine just how much
time I had spent for this information! Thanks!
Attractive section of content. I just stumbled upon your website and
in accession capital to assert that I acquire in fact enjoyed account your
blog posts. Any way I’ll be subscribing to your feeds and even I achievement you access consistently fast.
Hi there, just wanted to mention, I liked this post. It was inspiring.
Keep on posting!
I believe what you said made a bunch of sense. But, what about this? suppose you typed a catchier post title? I ain’t suggesting your content is not good, but suppose you added a post title that grabbed people’s attention? I mean %BLOG_TITLE% is a little boring. You should look at Yahoo’s home page and watch how they create news titles to get viewers to open the links. You might add a video or a pic or two to grab readers excited about what you’ve written. In my opinion, it might bring your posts a little bit more interesting.
http://mrkarpiuk.xaa.pl/boards/shopping/member.php?action=profile&uid=33570
https://tandaseru.id/2017/09/06/usai-divonis-bebas-ustadz-alfian-tanjung-kembali-ditangkap/
https://ravolutionfestival.com/
https://oldforum.citysakh.ru/?talkid=26125
http://collie.fatbb.ru/viewtopic.php?f=18&t=4165&start=0&view=print
Great post. I was checking constantly this blog and I am impressed!
Extremely useful info specially the remaining part 🙂 I care for such information a
lot. I was looking for this certain info for a very lengthy time.
Thanks and best of luck.
Greetings! Very helpful advice in this particular post!
It is the little changes that make the most important changes.
Many thanks for sharing!
I like what you guys are usually up too.
Such clever work and coverage! Keep up the great works guys I’ve included you guys to my own blogroll.
It’s awesome to go to see this website and reading the views of all
mates on the topic of this paragraph, while I am also zealous of getting experience.
Well done! Find more at https://79king6.co/.
Very well voiced of course. .
Thanks in support of sharing such a fastidious thought, piece of writing
is fastidious, thats why i have read it entirely
Thank you for every other wonderful post. Where else may just anybody get that type of information in such a perfect manner of writing? I have a presentation subsequent week, and I am at the look for such information.
http://tekst-pesen.ru/2018/04/04/
купить диплом провизора
http://primuothod.ru
купить диплом в магадане
Hello there, I believe your site could be having internet
browser compatibility issues. When I take a look at your web site in Safari, it
looks fine however when opening in Internet Explorer, it has some overlapping issues.
I just wanted to provide you with a quick heads up! Besides
that, wonderful website!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151970
https://rentry.org/qfrike64
https://rentry.org/4qps9srv
https://magfalcon1978.diary.ru/
https://okwave.jp/profile/u3115867.html
Undeniably believe that which you stated. Your favorite reason appeared to be
on the internet the easiest thing to be aware of. I
say to you, I definitely get annoyed while people think
about worries that they just do not know about. You managed to hit the nail upon the top as well as defined out the whole thing without having side-effects , people can take
a signal. Will probably be back to get more. Thanks
Excellent post. I used to be checking constantly this blog
and I’m impressed! Very useful information specially
the closing phase 🙂 I take care of such info a lot.
I was seeking this certain information for a long time.
Thanks and good luck.
Wow, superb blog layout! How long have you been blogging for?
you make blogging look easy. The overall look of your site is fantastic, as well as the content!
It’s really a cool and helpful piece of information. I’m happy that you just shared this helpful information with us.
Please stay us informed like this. Thanks for sharing.
Its like you read my mind! You appear to know a lot about this, like you wrote the book in it or something. I think that you could do with some pics to drive the message home a bit, but instead of that, this is excellent blog. A fantastic read. I’ll definitely be back.
http://cavale.enseeiht.fr/redmine/issues/215
купить диплом в петропавловске-камчатском
http://primuothod.ru
купить диплом в невинномысске
Great blog here! Also your website loads up very fast! What web
host are you using? Can I get your affiliate link
to your host? I wish my site loaded up as quickly as yours lol
Hello there, just became aware of your blog through Google, and found that it is really informative.
I’m gonna watch out for brussels. I’ll be grateful if you continue
this in future. Many people will be benefited from your writing.
Cheers!
I’m truly enjoying the design and layout of your website.
It’s a very easy on the eyes which makes it much more pleasant for me to come here and visit more often. Did you hire out a designer to create your theme?
Great work!
This site really has all of the info I wanted
concerning this subject and didn’t know who to ask.
срочный ремонт сотовых телефонов
Hi, after reading this remarkable post i am also cheerful to share my know-how here with mates.
If you are going for most excellent contents like I do, simply
visit this web site daily since it offers feature contents,
thanks
With havin so much content and articles do you ever run into any
problems of plagorism or copyright infringement?
My website has a lot of completely unique content
I’ve either created myself or outsourced but it seems a
lot of it is popping it up all over the web without my permission. Do you know any solutions to help reduce content from being stolen? I’d definitely
appreciate it.
Hi there, i read your blog occasionally and i own a similar one and i was just wondering
if you get a lot of spam comments? If so how
do you prevent it, any plugin or anything you can advise?
I get so much lately it’s driving me insane so any help is very much appreciated.
Thanks for finally talking about > Fornix (crus)/ Stria Terminalis
– fxst | Whole Brain Protocol for Tractography with Empirical
MRI < Liked it!
This is my first time visit at here and i am in fact happy to read all at one place.
Hi, Neat post. There’s a problem together with your website in web explorer, would
test this? IE still is the market leader and a good part of other people will pass over your fantastic writing due to this problem.
Hi there, just wanted to say, I loved this post. It was inspiring.
Keep on posting!
Hi, Neat post. There’s a problem along with your web site in internet explorer,
might test this? IE still is the marketplace leader and
a big component to other people will miss your fantastic
writing because of this problem.
hi!,I really like your writing so so much! share we keep up a correspondence
more approximately your article on AOL? I require a specialist on this area to
unravel my problem. Maybe that’s you! Taking a look ahead to
look you.
Hey there! I’ve been following your site for a long time now and finally got the
bravery to go ahead and give you a shout out from Houston Tx!
Just wanted to mention keep up the excellent job!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/350835
http://www.babelcube.com/user/cassandra-davis
https://permacultureglobal.org/users/59739-andrea-fleming
http://www.babelcube.com/user/lisa-johnson
https://www.hentai-foundry.com/user/slavascla1958/profile
It’s amazing for me to have a site, which is useful in favor of my knowledge.
thanks admin
Hi there Dear, are you truly visiting this web site daily, if so then you will definitely get pleasant experience.
http://profi-sk.ru/dizajn/pochemu-nalichie-diploma-vsyo-eshhe-igraet-sushhestvennuyu-rol.html
купить диплом в белорецке
http://diplomgos.info/
купить диплом для иностранцев
Superb blog! Do you have any recommendations for aspiring writers?
I’m planning to start my own website soon but I’m a little lost on everything.
Would you recommend starting with a free platform like
Wordpress or go for a paid option? There are
so many choices out there that I’m totally confused ..
Any tips? Cheers!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://smehno1951.diary.ru/
https://cannabis.net/user/151628
https://rentry.org/qopzmris
https://kaliter1995198419861951.diary.ru/
http://www.babelcube.com/user/mary-king
Howdy! Do you use Twitter? I’d like to follow you if that would be okay.
I’m absolutely enjoying your blog and look forward to new updates.
What’s up everybody, here every person is sharing these
knowledge, thus it’s fastidious to read this blog, and I used to visit
this web site daily.
Thanks for every other informative website. The place
else may I get that kind of info written in such
a perfect approach? I’ve a mission that I’m simply now running
on, and I have been at the look out for such information.
Good post. I learn something new and challenging on blogs I stumbleupon everyday.
It will always be interesting to read through articles from other
authors and practice something from their sites.
What you published was very logical. However, what about this? suppose you added a little information? I am not saying your content is not solid, however suppose you added a post title to possibly grab people’s attention? I mean %BLOG_TITLE% is a little boring. You ought to look at Yahoo’s front page and note how they create news titles to get people to click. You might add a related video or a related pic or two to grab readers interested about everything’ve written. In my opinion, it could bring your website a little livelier.
http://forum.kamorka.com/viewtopic.php?t=10021&start=105
купить диплом в озёрске
http://publicspaces17.ru
купить диплом биолога
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/apbush
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196847
https://cannabis.net/user/152072
https://okwave.jp/profile/u3117216.html
https://www.quia.com/profiles/natalie544williams
I am in fact thankful to the owner of this web page who has
shared this enormous post at here.
https://antimafia.se/ – hashish with a syringe 10 grams discounts. 11%
https://antimafia.se/ – powdered heroin – snorting 70 gram discount 22%
https://antimafia.se/ – ecstasy under the tongue – 1 gram discount 36%
https://antimafia.se/ – hachis con una jeringuilla 17 gramas descuento. 44%
https://antimafia.se/ – heroina en polvo – esnifada 30 gramas rebajas 25%
https://antimafia.se/ – extasis bajo la lengua – 1 grama rebajas 25%
https://antimafia.se/ – hasista ruiskulla 10 grams discount. 31%
https://antimafia.se/ – heroiinijauhe – nuuskaaminen 55 gram discount 11%
https://antimafia.se/ – ekstaasi kielen alla – 5 gram sale 18%
https://antimafia.se/ – haszysz za pomoca strzykawki 17 gram sleva. 47%
https://antimafia.se/ – sproszkowana heroina – wciaganie 70 gram discounts 27%
https://antimafia.se/ – ekstaza pod jezykiem – 5 gram discount 45%
https://antimafia.se/ – du haschisch avec une seringue 10 grammes discounts. 42%
https://antimafia.se/ – heroine en poudre – sniffer 30 gram sleva 15%
https://antimafia.se/ – ecstasy sous la langue – 15 grammes discount 13%
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/pcsvpguh
https://www.haikudeck.com/presentations/ZLXGyKsGLG
https://ellak.gr/user/opoo1962/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196317
https://cannabis.net/user/152196
Good day! Do you use Twitter? I’d like to follow you if that would be ok.
I’m definitely enjoying your blog and look forward to new updates.
Hi there fantastic blog! Does running a blog such as this require a large amount of
work? I’ve absolutely no expertise in coding but
I was hoping to start my own blog soon. Anyways, should you have any recommendations or techniques for new blog owners please share.
I know this is off topic however I simply had to ask.
Cheers!
Appreciating the persistence you put into your blog and in depth information you offer.
It’s great to come across a blog every once in a while that
isn’t the same unwanted rehashed information. Wonderful read!
I’ve saved your site and I’m adding your RSS feeds to my Google account.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/yhgjsn7j
https://gegeff1983.diary.ru/
https://tubeteencam.com/user/deadhonor19901973/profile
https://okwave.jp/profile/u3115931.html
https://imageevent.com/geoiter1976
Today, while I was at work, my sister stole my iphone and tested to see if it can survive a twenty five foot drop, just so she can be a youtube sensation. My apple ipad is now broken and she has 83 views. I know this is completely off topic but I had to share it with someone!
http://bbs2.xingxiancn.com/space-uid-393119.html
купить диплом в иркутске
http://seoklan.ru
купить диплом в ялте
Attractive section of content. I simply stumbled upon your site and
in accession capital to assert that I get in fact enjoyed account
your blog posts. Anyway I will be subscribing in your feeds
or even I success you get right of entry to persistently fast.
We absolutely love your blog and find most of your post’s
to be precisely what I’m looking for. Do you offer guest
writers to write content to suit your needs? I wouldn’t mind producing a post or elaborating on some of the subjects you write
in relation to here. Again, awesome weblog!
When someone writes an piece of writing he/she maintains the idea of a user in his/her mind that how a user can be
aware of it. Thus that’s why this article is outstdanding.
Thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3194619
https://anotepad.com/notes/dercfce9
http://www.nfomedia.com/profile?uid=rOiZacF
https://haveagood.holiday/users/351599
https://rentry.org/q76un4zp
Great article.
I was more than happy to discover this page.
I need to to thank you for your time for this fantastic read!!
I definitely really liked every part of it and i also have you saved to fav to
check out new information in your web site.
What’s up, yeah this paragraph is really good and I have learned lot of things from it concerning
blogging. thanks.
What’s up i am kavin, its my first occasion to commenting anywhere, when i read this paragraph i thought i could also make comment due to this good piece of writing.
http://ekzersiskaliningrad.ru/stili/modern.html
купить свидетельство о заключении брака
http://seo1st.ru
купить диплом в бугульме
It’s actually a great and helpful piece of information. I’m satisfied that
you shared this useful information with us. Please stay us up to date like this.
Thanks for sharing.
Hi, i read your blog occasionally and i own a similar one and i was just
curious if you get a lot of spam comments? If so how do you stop it, any plugin or anything you
can recommend? I get so much lately it’s driving me insane so any assistance is very much appreciated.
Greetings I am so grateful I found your web site,
I really found you by error, while I was searching on Yahoo
for something else, Regardless I am here now and would just like to say
thanks a lot for a tremendous post and a all round interesting blog (I
also love the theme/design), I don’t have
time to read it all at the minute but I have saved it and also added in your RSS
feeds, so when I have time I will be back to read a lot more, Please do keep up the
great work.
I appreciate about what you’ve presented. I am highly interested in your perspective. It’s thought-provoking and worth a second read. Don’t forget to check out top nhà cái for more related content.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiZYcB
https://anotepad.com/notes/akq4da9s
https://anotepad.com/notes/45emx3jn
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195321
https://www.hentai-foundry.com/user/kluker1999/profile
Hmm is anyone else experiencing problems with the images
on this blog loading? I’m trying to find out if its a problem on my end or if it’s the
blog. Any feed-back would be greatly appreciated.
https://fakeoff.org/ – hashish with a syringe 5 gram discounts. 43%
https://fakeoff.org/ – powdered heroin – snorting 70 gram discounts 11%
https://fakeoff.org/ – ecstasy under the tongue – 2 grams discounts 45%
https://fakeoff.org/ – hachis con una jeringuilla 51 gramas descuentos. 46%
https://fakeoff.org/ – heroina en polvo – esnifada 70 gramas descuento 35%
https://fakeoff.org/ – extasis bajo la lengua – 1 gramas rebajas 11%
https://fakeoff.org/ – hasista ruiskulla 3 grams discounts. 36%
https://fakeoff.org/ – heroiinijauhe – nuuskaaminen 70 grams sleva 34%
https://fakeoff.org/ – ekstaasi kielen alla – 3 grams discount 17%
https://fakeoff.org/ – haszysz za pomoca strzykawki 17 gramy discount. 27%
https://fakeoff.org/ – sproszkowana heroina – wciaganie 70 gram sale 38%
https://fakeoff.org/ – ekstaza pod jezykiem – 25 gram discount 26%
https://fakeoff.org/ – du haschisch avec une seringue 3 gram sleva. 27%
https://fakeoff.org/ – heroine en poudre – sniffer 30 grammes sale 18%
https://fakeoff.org/ – ecstasy sous la langue – 20 grammes sale 21%
Very shortly this web page will be famous among all blogging
viewers, due to it’s fastidious articles
My family members every time say that I am killing my time here at web, however I know I am getting know-how all the time by reading thes
pleasant articles.
Thông tin quan trọng về cổng game Sunwin đã được cập nhật đến với người chơi. Tham gia tại thương hiệu cá cược trực tuyến uy tín hàng đầu, bạn sẽ có được trải nghiệm đáng giá nhất. Dưới sự nâng đỡ và điều hành đến từ Suncity Group (Tập đoàn giải trí danh tiếng thế giới), Sunwin đã nhanh chóng hoàn t
Very good tips Thanks a lot.
Feel free to surf to my website: http://damoa2019.maru.net/bbs/board.php?bo_table=free&wr_id=11055
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/59350-ashley-tapia
https://imageevent.com/drakoc1990
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3194660
https://imageevent.com/pitbuldog199419641955
https://imageevent.com/jiofht1951
I just like the helpful info you provide in your articles.
I will bookmark your blog and take a look at again right here frequently.
I’m somewhat certain I’ll be informed many new stuff right here!
Best of luck for the following!
Hurrah! Finally I got a weblog from where I be able to truly obtain useful information concerning my study and knowledge.
Hey there! This is my 1st comment here so I just wanted to give a quick shout out and say I really enjoy reading your posts. Can you recommend any other blogs/websites/forums that go over the same subjects? Thanks a ton!
http://it-dix.ru/ob-avtore
купить диплом в сарове
http://git-ugra.ru
купить диплом в братске
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/reedab
https://permacultureglobal.org/users/60220-meghan-washington
https://okwave.jp/profile/u3116993.html
https://www.obesityhelp.com/members/xxwitcher1975/about_me/
https://leoguard1962.diary.ru/
I’ve learn several good stuff here. Certainly worth bookmarking for revisiting.
I wonder how much attempt you place to create such a excellent
informative website.
Hey! I know this is kinda off topic but I was wondering if you knew where I could get a captcha plugin for my comment form?
I’m using the same blog platform as yours and I’m having problems
finding one? Thanks a lot!
We’re a group of volunteers and starting a new scheme in our community.
Your web site provided us with valuable information to work on. You have
done a formidable job and our entire community will be grateful to you.
You’re so awesome! I do not think I’ve truly read through anything like that before.
So nice to discover someone with original thoughts on this subject
matter. Seriously.. thank you for starting this up. This web
site is one thing that is required on the internet, someone with a bit of originality!
https://dr-Nona.ru/ – доктор нонна
프라그마틱 무료 슬롯
당신은 너무 어리고 당신이하는 일이나 말에 대해 생각조차하지 않습니다.내 수고를 헛되이 낭비하는 것만으로도 정말 가슴이 아픕니다.
I used to be recommended this blog via my cousin. I am not
positive whether this publish is written by way of him as nobody else recognise such targeted approximately my difficulty.
You’re incredible! Thanks!
Hello Dear, are you genuinely visiting this web site daily, if so afterward you will without doubt obtain pleasant knowledge.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/152206
https://rentry.org/g6ub859w
https://okwave.jp/profile/u3115402.html
https://liza441992.bandcamp.com/album/surprise-1
http://www.babelcube.com/user/cynthia-fernandes
For hottest news you have to visit world wide web and on web I found this web page as a finest web site
for newest updates.
‘Sugary food is a drug for me’: A growing number of children are addicted to ultraprocessed foods
kraken14.at
Chicago native Jeffrey Odwazny says he has been addicted to ultraprocessed food since he was a child.
“I was driven to eat and eat and eat, and while I would overeat healthy food, what really got me were the candies, the cakes, the pies, the ice cream,” said the 54-year-old former warehouse supervisor
https://kraken-22.at
2krn.at
“I really gravitated towards the sugary ultraprocessed foods — it was like a physical drive, I had to have it,” he said. “My parents would find hefty bags full of candy wrappers hidden in my closet. I would steal things from stores as a kid and later as an adult.”
Some 12% of the nearly 73 million children and adolescents in the United States today struggle with a similar food addiction, according to research. To be diagnosed, children must meet Yale Food Addiction Scale criteria as stringent as any for alcohol use disorder or other addictions.
“Kids are losing control and eating to the point where they feel physically ill,” said Ashley Gearhardt, a professor of psychology at the University of Michigan in Ann Arbor who conducted the research and developed the Yale addiction scale.
“They have intense cravings and may be sneaking, stealing or hiding ultraprocessed foods,” Gearhardt said. “They may stop going out with friends or doing other activities they used to enjoy in order to stay at home and eat, or they feel too sluggish from overeating to participate in other activities.”
Her research also shows about 14% of adults are clinically addicted to food, predominantly ultraprocessed foods with higher levels of sugar, salt, fat and additives.
For comparison, 10.5% of Americans age 12 or older were diagnosed with alcohol use disorder in 2022, according to the National Survey on Drug Use and Health.
While many people addicted to food will say that their symptoms began to worsen significantly in adolescence, some recall a childhood focused on ultraprocessed food.
“By age 2 or 3, children are likely eating more ultraprocessed foods in any given day than a fruit or vegetable, especially if they’re poor and don’t have enough money in their family to have enough quality food to eat,” Gearhardt said. “Ultraprocessed foods are cheap and literally everywhere, so this is also a social justice issue.”
An addiction to ultraprocessed foods can highjack a young brain’s reward circuitry, putting the primitive “reptilian brain,” or amygdala, in charge — thus bypassing the prefrontal cortex where rational decision-making occurs, said Los Angeles registered dietitian nutritionist David Wiss, who specializes in treating food addiction.
판다스 포춘
그렇게 말하며 케이스 옆에 있던 찻잔을 집어서 문밖으로 내던졌다.
Общественное здание или его часть зетфликс онлайн с оборудованием для публичной демонстрации кинофильмов. Главное помещение кинотеатра – зрительный зал с экраном большого размера и системой воспроизведения звука. В современных кинотеатрах система звуковоспроизведения состоит из нескольких громкоговорителей, обеспечивающих объёмный звук. Первые стационарные кинотеатры назывались в России «электротеатрами»
зетфликс https://zetflixhd.life/
I don’t even know how I ended up here, but I thought
this post was great. I don’t know who you are but certainly you’re going to a famous blogger if you are not
already 😉 Cheers!
Thanks on your marvelous posting! I seriously enjoyed reading it,
you happen to be a great author.I will make certain to bookmark your blog and definitely will come back down the road.
I want to encourage yourself to continue your great job, have a nice evening!
Heya just wanted to give you a quick heads up and let you know a few of the
pictures aren’t loading correctly. I’m not sure why but I think its a linking issue.
I’ve tried it in two different internet browsers and both show the same outcome.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/crab7901959/about_me/
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195702
https://cannabis.net/user/151345
https://imageevent.com/lliakpuji1980
https://www.quia.com/profiles/wellingtonki
It is truly a nice and useful piece of info. I am glad that you simply shared this useful information with us.
Please keep us informed like this. Thank you for sharing.
Thank you for the auspicious writeup. It in fact was
a amusement account it. Look advanced to far added agreeable from you!
However, how could we communicate?
Hey there would you mind stating which blog platform you’re using?
I’m going to start my own blog in the near future but I’m having a hard time selecting between BlogEngine/Wordpress/B2evolution and Drupal.
The reason I ask is because your design seems different then most blogs and I’m looking
for something unique. P.S My apologies for being
off-topic but I had to ask!
Piece of writing writing is also a fun, if you be familiar with after that you can write otherwise it is complex
to write.
It’s a shame you don’t have a donate button! I’d most certainly donate
to this fantastic blog! I suppose for now i’ll settle for book-marking and adding your RSS feed to my Google
account. I look forward to fresh updates and will talk about this website with my Facebook group.
Talk soon!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/59275-brian-ashcraft
https://tubeteencam.com/user/prtyzn1988/profile
https://dmitriyua1987.bandcamp.com/album/wrong-turn
https://haveagood.holiday/users/352093
https://permacultureglobal.org/users/58920-april-cortez
I used to be suggested this website through my cousin. I am not positive whether or not this submit is written through him as no one else know
such detailed about my difficulty. You are wonderful!
Thanks!
Beneficial stuff With thanks.
Here is my site; http://Ict.wku.Ac.th/question/%ec%b6%9c%ec%9e%a5%eb%a7%88%ec%82%ac%ec%a7%80-all-day-and-you-will-realize-10-things-about-yourself-you-never-knew-6/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/simurgl1992
https://www.obesityhelp.com/members/liash1980/about_me/
https://www.obesityhelp.com/members/superb19841968/about_me/
https://haveagood.holiday/users/350446
https://rentry.org/ehfkz9ys
Your way of telling all in this article is really good, every one be able to effortlessly
be aware of it, Thanks a lot.
My brother suggested I would possibly like this website.
He used to be totally right. This put up truly made
my day. You cann’t consider simply how a lot time I had spent for this information! Thanks!
Hi there would you mind letting me know which webhost
you’re using? I’ve loaded your blog in 3 different internet browsers and
I must say this blog loads a lot quicker then most.
Can you suggest a good internet hosting provider at a
honest price? Cheers, I appreciate it!
Thank you for every other magnificent article.
Where else may just anybody get that kind of information in such a perfect approach of writing?
I have a presentation subsequent week, and I am at the look for
such information.
I am truly pleased to read this webpage posts which consists of
plenty of helpful data, thanks for providing these statistics.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/demager1977/profile
https://www.haikudeck.com/presentations/LM7WTMbUWp
https://www.haikudeck.com/presentations/aZNJCjUXCi
https://www.hentai-foundry.com/user/voroncov1997/profile
https://imageevent.com/confessor1998
Stunning quest there. What occurred after? Good luck!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/JEqDon4rvG
http://www.babelcube.com/user/brandon-capers
https://okwave.jp/profile/u3116767.html
https://tubeteencam.com/user/reala1970/profile
https://rentry.org/w35hpo77
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://klubochek1974.bandcamp.com/album/behavioral
https://rayne12199719991968.diary.ru/
https://okwave.jp/profile/u3117362.html
https://www.hentai-foundry.com/user/benladen21956/profile
https://www.quia.com/profiles/iabond
Hey there are using WordPress for your blog platform?
I’m new to the blog world but I’m trying to get started and create my own. Do you need any coding expertise to make your own blog?
Any help would be really appreciated!
Ahaa, its fastidious dialogue regarding this piece of writing here
at this weblog, I have read all that, so at this time me
also commenting here.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196526
https://imageevent.com/diableroo1953
http://www.babelcube.com/user/tracy-hope
https://okwave.jp/profile/u3116054.html
https://www.hentai-foundry.com/user/kluker1999/profile
Greetings! Very useful advice within this article!
It’s the little changes which will make the largest changes.
Thanks a lot for sharing!
Spot on with this write-up, I honestly believe this amazing
site needs a lot more attention. I’ll probably be back again to read
more, thanks for the advice!
Wow, that’s what I was exploring for, what a data! present here at this web site, thanks admin of this web
page.
You really make it appear really easy with your presentation but I in finding this matter to
be really one thing that I think I would by no means understand.
It sort of feels too complex and very huge for me. I am looking ahead
for your subsequent put up, I will try to get the dangle of it!
Hey! This post could not be written any better! Reading through this post reminds
me of my old room mate! He always kept chatting about this.
I will forward this article to him. Pretty sure he will have a
good read. Thanks for sharing!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/150814
https://chyoa.com/user/mopito1964
https://anotepad.com/notes/pcbbbsr2
http://www.babelcube.com/user/ashley-small
https://anotepad.com/notes/c7pr9kq4
Hello, i think that i saw you visited my web site so i came to “return the favor”.I’m attempting to
find things to improve my web site!I suppose its ok to use a few of your ideas!!
I am genuinely grateful to the owner of this web site who has shared this fantastic paragraph at at
this place.
Good post. I absolutely appreciate this site.
Keep it up!
The world’s most liveable cities for 2024
жесткое русское порно
It’s considered among the most beautiful cities in the world to visit, and it seems that Vienna may also be an unbeatable place to live.
The Austrian city has been crowned the most liveable city in the world yet again in the annual list from the Economist Intelligence Unit (EIU), which was released today.
The EIU, a sister organization to The Economist, ranked 173 cities across the globe on a number of significant factors, including health care, culture and environment, stability, infrastructure and education.
Vienna topped the list for the third consecutive year, receiving “perfect” scores in four out of five of the categories — the city was marked lower for culture and environment due to an apparent lack of significant sporting events.
Just behind the Austrian capital, Denmark’s Copenhagen retained its second place position, while Switzerland’s Zurich moved up from sixth place to third on the list.
Australia’s Melbourne fell from third to fourth place, while Canadian city Calgary tied for fifth place with Swiss city Geneva.
Canada’s Vancouver and Australia’s Sydney were in joint seventh place, and Japan’s Osaka and New Zealand’s Auckland rounded out the top 10 in joint ninth place.
«Король» обвинителей «Лайф-из-Гуд» Набойченко – кто он на самом деле?
Кооператив Бествей
Евгений Набойченко – бывший IT-директор компании «Лайф-из-Гуд», а также сисадмин российского сегмента платежной системы международной инвестиционной компании «Гермес». Он – ключевой свидетель обвинения в так называемом деле «Лайф-из-Гуд» – «Гермес» – кооператива «Бест Вей» и основателя «Лайф-из-Гуд»/«Бест Вей» Романа Василенко, которое рассматривается сейчас судьей Екатериной Богдановой в Приморском районном суде Санкт-Петербурга.
С выполненной им блокировки российского сегмента платежной системы компании «Гермес», а также с его (нетрезвых) сообщений в социальных сетях (сейчас все они им стерты) началась эскалация этого уголовного дела. Но, скорее всего, дело им же самим и его подельниками из питерской полиции и создано.
Вор и дебошир
Наши источники делятся информацией о хищении активов с кошельков клиентов «Гермеса», к которым Евгений Набойченко имел доступ.
Набойченко не гнушался воровать даже у близких. Однажды остановившись у своей любовницы Светланы Н., Евгений избил ее и украл деньги с ее криптовалютного кошелька, пока она пыталась прийти в себя после побоев.
Он бил не только любовницу, но жену и детей, с которыми жил в Краснодарском крае. А после того как его супруга Виктория подала на развод, разгромил, ободрал и ограбил общий дом – унес все, что можно было унести. Надо ли говорить, что Набойченко который год не платит супруге алименты.
Вымогатель
Евгений Набойченко, по словам его бывшей жены, в один прекрасный день решил заработать и с помощью вымогательства. Он шантажировал Романа Василенко – требовал с него деньги: 170 тыс. евро. При этом, по утверждению Виктории Набойченко, угрожал увечьями и самому Роману Василенко, и его супруге, и детям. Похвалялся перед (тогда еще) женой своими матерными сообщениями с угрозами, которые он посылал Василенко и его близким.
Кроме того, вымогал деньги у клиентов «Гермеса»: свидетельства такого рода нам предоставлены.
Алкоголик и наркоман
Деньги нужны Набойченко в том числе для того, чтобы «питать» свою наркотическую зависимость – с некоторых пор он предпочитает шлифовать водку галлюциногенными грибами, об увлечении которыми с упоением рассказывал даже в своих соцсетях.
Он не всегда был алкоголиком и наркоманом – устроившись 10 лет назад в IT-службу, не пил и не употреблял. Но большие деньги, красивая жизнь, которая вдруг стала доступна для простого астраханского парня из низов, развратили его.
Клеветник
Он допустил целый ряд клеветнических высказываний в адрес Романа Василенко – за которые против него заведено уголовное дело. Но расследуется оно ни шатко ни валко, так как расследование тормозит начальник УЭБиПК ГУ МВД России по Санкт-Петербургу и Ленинградской области.
Завербованный органами
Виктория Набойченко сообщает, что за три дня до обыска в их тогда еще совместном доме Евгений уехал и ей сказал, что придут с обыском – но это пустая формальность. Он уже был в сговоре с полицией, так как за три дня был предупрежден об этом обыске.
15 февраля 2022 года Евгений Набойченко был задержан, но отправился не в СИЗО – куда отправили других сотрудников «Лайф-из-Гуд», не был помещен в изолятор или КПЗ, а жил на полицейской квартире.
Он отказался от услуг адвокатов и на этой квартире, по свидетельству одного из полицейских, «бухал с операми». А также каждый день водил оперов по барам и ресторанам – угощал их за свой счет.
В разговоре с женой он очень гордился покровительством начальника УЭБиПК питерского главка МВД.
Наши источники считают, что он украл деньги клиентов «Гермеса», разделив их со своими друзьями из полиции. Вся его дальнейшая жизнь и привычки указывают на то, что он использует ворованные деньги.
Трус и лжец
Набойченко сообщил в социальных сетях, что к нему пришло с адреса fall@mybizon.online письмо, в котором содержится угроза в адрес получателя, призывающая его не являться в суд для дачи показаний. В письме указано, что если он поедет в суд, то не доедет и будет трупом.
Создатель преступного сообщества
Нет сомнений, что Набойченко устроил историю с обрушением платежной системы, чтобы скрыть свое воровство активов клиентов. Также чтобы скрыть свое воровство, он обвинил во всем Василенко и технических специалистов компании «Лайф-из-Гуд», которые находятся сейчас на скамье подсудимых. Оклеветал всех, чтобы отвести подозрения от себя и своих подельников из питерской полиции.
Итак, кризис в «Гермесе» устроил Набойченко. Платежную систему разгромил Набойченко. Активы украл Набойченко, поделившись с питерскими полицейскими.
А теперь сидит и бездельничает, прожигая сворованные деньги. И боится идти в суд – чтобы не превратиться в суде из свидетеля в обвиняемого.
Скорее всего, в суд Набойченко не пускают сами полицейские – опасаясь его неадекватности и перекрестного допроса, на котором вскроется лживость его показаний. То есть силовики заметают следы своего преступления, совершенного с помощью марионетки Набойченко.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151857
https://haveagood.holiday/users/350984
https://tubeteencam.com/user/esindorf1957/profile
http://www.nfomedia.com/profile?uid=rOiXhiF
http://www.nfomedia.com/profile?uid=rOjQgcH
The world’s most liveable cities for 2024
смотреть гей порно
It’s considered among the most beautiful cities in the world to visit, and it seems that Vienna may also be an unbeatable place to live.
The Austrian city has been crowned the most liveable city in the world yet again in the annual list from the Economist Intelligence Unit (EIU), which was released today.
The EIU, a sister organization to The Economist, ranked 173 cities across the globe on a number of significant factors, including health care, culture and environment, stability, infrastructure and education.
Vienna topped the list for the third consecutive year, receiving “perfect” scores in four out of five of the categories — the city was marked lower for culture and environment due to an apparent lack of significant sporting events.
Just behind the Austrian capital, Denmark’s Copenhagen retained its second place position, while Switzerland’s Zurich moved up from sixth place to third on the list.
Australia’s Melbourne fell from third to fourth place, while Canadian city Calgary tied for fifth place with Swiss city Geneva.
Canada’s Vancouver and Australia’s Sydney were in joint seventh place, and Japan’s Osaka and New Zealand’s Auckland rounded out the top 10 in joint ninth place.
«Король» обвинителей «Лайф-из-Гуд» Набойченко – кто он на самом деле?
Лайф из Гуд последние новости
Евгений Набойченко – бывший IT-директор компании «Лайф-из-Гуд», а также сисадмин российского сегмента платежной системы международной инвестиционной компании «Гермес». Он – ключевой свидетель обвинения в так называемом деле «Лайф-из-Гуд» – «Гермес» – кооператива «Бест Вей» и основателя «Лайф-из-Гуд»/«Бест Вей» Романа Василенко, которое рассматривается сейчас судьей Екатериной Богдановой в Приморском районном суде Санкт-Петербурга.
С выполненной им блокировки российского сегмента платежной системы компании «Гермес», а также с его (нетрезвых) сообщений в социальных сетях (сейчас все они им стерты) началась эскалация этого уголовного дела. Но, скорее всего, дело им же самим и его подельниками из питерской полиции и создано.
Вор и дебошир
Наши источники делятся информацией о хищении активов с кошельков клиентов «Гермеса», к которым Евгений Набойченко имел доступ.
Набойченко не гнушался воровать даже у близких. Однажды остановившись у своей любовницы Светланы Н., Евгений избил ее и украл деньги с ее криптовалютного кошелька, пока она пыталась прийти в себя после побоев.
Он бил не только любовницу, но жену и детей, с которыми жил в Краснодарском крае. А после того как его супруга Виктория подала на развод, разгромил, ободрал и ограбил общий дом – унес все, что можно было унести. Надо ли говорить, что Набойченко который год не платит супруге алименты.
Вымогатель
Евгений Набойченко, по словам его бывшей жены, в один прекрасный день решил заработать и с помощью вымогательства. Он шантажировал Романа Василенко – требовал с него деньги: 170 тыс. евро. При этом, по утверждению Виктории Набойченко, угрожал увечьями и самому Роману Василенко, и его супруге, и детям. Похвалялся перед (тогда еще) женой своими матерными сообщениями с угрозами, которые он посылал Василенко и его близким.
Кроме того, вымогал деньги у клиентов «Гермеса»: свидетельства такого рода нам предоставлены.
Алкоголик и наркоман
Деньги нужны Набойченко в том числе для того, чтобы «питать» свою наркотическую зависимость – с некоторых пор он предпочитает шлифовать водку галлюциногенными грибами, об увлечении которыми с упоением рассказывал даже в своих соцсетях.
Он не всегда был алкоголиком и наркоманом – устроившись 10 лет назад в IT-службу, не пил и не употреблял. Но большие деньги, красивая жизнь, которая вдруг стала доступна для простого астраханского парня из низов, развратили его.
Клеветник
Он допустил целый ряд клеветнических высказываний в адрес Романа Василенко – за которые против него заведено уголовное дело. Но расследуется оно ни шатко ни валко, так как расследование тормозит начальник УЭБиПК ГУ МВД России по Санкт-Петербургу и Ленинградской области.
Завербованный органами
Виктория Набойченко сообщает, что за три дня до обыска в их тогда еще совместном доме Евгений уехал и ей сказал, что придут с обыском – но это пустая формальность. Он уже был в сговоре с полицией, так как за три дня был предупрежден об этом обыске.
15 февраля 2022 года Евгений Набойченко был задержан, но отправился не в СИЗО – куда отправили других сотрудников «Лайф-из-Гуд», не был помещен в изолятор или КПЗ, а жил на полицейской квартире.
Он отказался от услуг адвокатов и на этой квартире, по свидетельству одного из полицейских, «бухал с операми». А также каждый день водил оперов по барам и ресторанам – угощал их за свой счет.
В разговоре с женой он очень гордился покровительством начальника УЭБиПК питерского главка МВД.
Наши источники считают, что он украл деньги клиентов «Гермеса», разделив их со своими друзьями из полиции. Вся его дальнейшая жизнь и привычки указывают на то, что он использует ворованные деньги.
Трус и лжец
Набойченко сообщил в социальных сетях, что к нему пришло с адреса fall@mybizon.online письмо, в котором содержится угроза в адрес получателя, призывающая его не являться в суд для дачи показаний. В письме указано, что если он поедет в суд, то не доедет и будет трупом.
Создатель преступного сообщества
Нет сомнений, что Набойченко устроил историю с обрушением платежной системы, чтобы скрыть свое воровство активов клиентов. Также чтобы скрыть свое воровство, он обвинил во всем Василенко и технических специалистов компании «Лайф-из-Гуд», которые находятся сейчас на скамье подсудимых. Оклеветал всех, чтобы отвести подозрения от себя и своих подельников из питерской полиции.
Итак, кризис в «Гермесе» устроил Набойченко. Платежную систему разгромил Набойченко. Активы украл Набойченко, поделившись с питерскими полицейскими.
А теперь сидит и бездельничает, прожигая сворованные деньги. И боится идти в суд – чтобы не превратиться в суде из свидетеля в обвиняемого.
Скорее всего, в суд Набойченко не пускают сами полицейские – опасаясь его неадекватности и перекрестного допроса, на котором вскроется лживость его показаний. То есть силовики заметают следы своего преступления, совершенного с помощью марионетки Набойченко.
Hi! I know this is kinda off topic but I’d figured I’d ask.
Would you be interested in exchanging links or maybe guest writing a blog post or vice-versa?
My blog goes over a lot of the same topics as yours and
I think we could greatly benefit from each other.
If you are interested feel free to send me an email.
I look forward to hearing from you! Wonderful blog by the way!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/egaqp4ap
https://cannabis.net/user/151833
https://rentry.org/q92xam8m
http://www.babelcube.com/user/saad-campbell
https://rentry.org/oshvuy3y
Наш сервисный центр предлагает профессиональный сервисный центр по ремонту духовых шкафов с гарантией всех типов и брендов. Мы знаем, насколько важны для вас ваши духовки, и стремимся предоставить услуги первоклассного уровня. Наши опытные мастера оперативно и тщательно выполняют работу, используя только сертифицированные компоненты, что обеспечивает надежность и долговечность выполненных работ.
Наиболее распространенные поломки, с которыми сталкиваются владельцы духовых шкафов, включают неисправности термостата, поломку таймера, поврежденную дверцу, сбои контроллера, ошибки вентиляционной системы и повреждения электроники. Для устранения этих поломок наши профессиональные техники проводят ремонт нагревательных элементов, термостатов, таймеров, дверец, контроллеров, вентиляторов и электроники. Обратившись к нам, вы обеспечиваете себе надежный и долговечный мастерская по ремонту духовых шкафов.
Подробная информация доступна на сайте: https://remont-duhovyh-shkafov-ace.ru
Hey There. I found your blog the usage of msn. This is an extremely smartly written article.
I’ll make sure to bookmark it and come back to read more of your helpful information. Thanks for
the post. I’ll definitely return.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/paxanchik1975/profile
https://chyoa.com/user/elktra1959
https://permacultureglobal.org/users/58799-hoi-stakelbeck
https://nikixd19741969.bandcamp.com/album/lady-of-the-forest
http://www.nfomedia.com/profile?uid=rOiYajD
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/p577wdxm
https://milin1966.bandcamp.com/album/daddy-dom-and-me
https://www.obesityhelp.com/members/iseda1950/about_me/
https://tubeteencam.com/user/hfff19821950/profile
https://www.hentai-foundry.com/user/l1s14k1996/profile
Below, we record a variety of the prime websites within the GB with the best football odds. By the way, they also provide bonuses for brand new and common players — so ensure you control those. To discover more worth in sports betting, gamers want to match the chances of various betting sites. Odds
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/withmoon19961953
http://www.nfomedia.com/profile?uid=rOiYYeC
https://cannabis.net/user/152154
https://www.obesityhelp.com/members/nomiz21950/about_me/
https://permacultureglobal.org/users/60303-florence-gonzalez
Hey there! Would you mind if I share your blog with my zynga group?
There’s a lot of folks that I think would really appreciate your content.
Please let me know. Thanks
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3115316.html
https://www.hentai-foundry.com/user/almaz0001985/profile
https://tubeteencam.com/user/qaradaqir19941973/profile
https://tubeteencam.com/user/drstrange1983/profile
https://haveagood.holiday/users/351132
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiYgkJ
https://www.haikudeck.com/presentations/nnr2XLBkLP
https://www.quia.com/profiles/alex541nicholson
https://haveagood.holiday/users/350764
http://www.babelcube.com/user/michelle-waters
Наш сервисный центр предлагает высококачественный мастер по ремонту индукционной плиты адреса любых брендов и моделей. Мы знаем, насколько важны для вас ваши индукционные варочные панели, и готовы предложить сервис первоклассного уровня. Наши профессиональные техники проводят ремонтные работы с высокой скоростью и точностью, используя только сертифицированные компоненты, что гарантирует длительную работу выполненных работ.
Наиболее частые неисправности, с которыми сталкиваются пользователи индукционных варочных панелей, включают проблемы с нагревом, неисправности сенсорных панелей, ошибки ПО, неработающие разъемы и поломки корпуса. Для устранения этих проблем наши квалифицированные специалисты выполняют ремонт нагревательных элементов, сенсорных панелей, ПО, разъемов и механических компонентов. Обратившись к нам, вы гарантируете себе качественный и надежный мастерская по ремонту индукционных плит.
Подробная информация представлена на нашем сайте: https://remont-indukcionnyh-plit-now.ru
Hello there! This article couldn’t be written any better!
Reading through this article reminds me of my previous roommate!
He constantly kept preaching about this. I’ll send this post to
him. Fairly certain he will have a good read. Thanks for sharing!
Наша мастерская предлагает профессиональный официальный ремонт айпада на выезде всех типов и брендов. Мы понимаем, насколько необходимы вам ваши планшеты Apple, и обеспечиваем ремонт наилучшего качества. Наши квалифицированные специалисты работают быстро и аккуратно, используя только качественные детали, что обеспечивает надежность и долговечность наших услуг.
Наиболее общие проблемы, с которыми сталкиваются владельцы iPad, включают поврежденный экран, проблемы с батареей, ошибки ПО, неработающие разъемы и повреждения корпуса. Для устранения этих поломок наши профессиональные техники оказывают ремонт экранов, батарей, ПО, разъемов и механических компонентов. Обращаясь в наш сервисный центр, вы обеспечиваете себе качественный и надежный центр ремонта айпада адреса.
Подробная информация размещена на сайте: https://remont-ipad-pro.ru
Hello to all, the contents present at this site are truly remarkable for people knowledge, well, keep
up the nice work fellows.
Hey There. I found your weblog the use of msn. This is an extremely smartly written article.
I will be sure to bookmark it and come back to learn extra of your helpful info.
Thank you for the post. I’ll certainly comeback.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/emhgsr9h
http://www.nfomedia.com/profile?uid=rOiZddG
http://www.nfomedia.com/profile?uid=rOiYcgE
https://imageevent.com/renyu1960
http://www.nfomedia.com/profile?uid=rOjQgcH
Наш сервисный центр предлагает профессиональный вызвать мастера по ремонту кофемашин в москве всех типов и брендов. Мы знаем, насколько необходимы вам ваши кофемашины, и стремимся предоставить услуги наилучшего качества. Наши опытные мастера работают быстро и аккуратно, используя только оригинальные запчасти, что гарантирует надежность и долговечность выполненных работ.
Наиболее распространенные поломки, с которыми сталкиваются пользователи кофеварок, включают неисправности нагревательных элементов, неисправности насоса, неисправности программного обеспечения, неработающие разъемы и механические повреждения. Для устранения этих неисправностей наши квалифицированные специалисты выполняют ремонт нагревательных элементов, насосов, ПО, разъемов и механических компонентов. Доверив ремонт нам, вы обеспечиваете себе качественный и надежный вызвать мастера по ремонту кофемашин на выезде.
Подробная информация доступна на сайте: https://remont-kofemashin-top.ru
Do you have a spam problem on this blog; I also am a blogger, and I was wondering your situation; we
have developed some nice practices and we are looking to exchange solutions with
others, please shoot me an email if interested.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://lordtoor1989.diary.ru/
https://tubeteencam.com/user/weter1981/profile
https://okwave.jp/profile/u3118337.html
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195343
https://tubeteencam.com/user/voody7871971/profile
I do not know whether it’s just me or if everybody else encountering issues with your site.
It looks like some of the written text within your content are running off the
screen. Can someone else please comment and let me know if this is happening to them
as well? This may be a issue with my browser because I’ve had this happen previously.
Cheers
Pretty! This was a really wonderful post. Thanks for
supplying these details.
Следственная группа по делу «Бест Вей»: следователи или оборотни в погонах?
ОПГ Колокольцева
Следственная группа по делу кооператива «Бест Вей» во главе со следователями ГСУ ГУ МВД по Санкт-Петербургу и Ленобласти майором юстиции Екатериной Сапетовой и подполковником юстиции Константином Иудичевым играет собственную партию либо для того, чтобы выслужиться перед непосредственным начальством, либо из-за прямой заинтересованности в уничтожении кооператива со стороны конкурентов и/или банков.
Следствие по делу кооператива «Бест Вей» вступило в активную фазу ровно год назад. За это время арестованы четверо технических сотрудников, и уже год они томятся в СИЗО, пятеро граждан объявлены в розыск.
Офис кооператива буквально разгромлен двумя обысками, документация изъята, и следствие запрещает ее копировать, счета и недвижимость арестованы. Под арестом — активы на более чем 15 млрд рублей, хотя ущерб для так называемых потерпевших, пул которых следователи год собирали под разными предлогами, не превышает 115 млн рублей. Это спустя почти полтора года с момента возбуждения уголовного дела, не считая периода длительного проведения оперативно-разыскных мероприятий. Гора родила мышь.
При этом значительная часть этих потерпевших, например, бывшие пайщики, которые внесли невозвратный вступительный взнос, но затем не смогли копить на квартиру и сами вышли из кооператива, а теперь по наущению следствия подали заявление о том, что кооператив им не вернул то, что и не должен был вернуть. Потерпевших 100 с лишним человек, притом что в кооперативе — более 19 тыс. пайщиков, то есть капля в море, да и эти 100 не имеют юридически обоснованных претензий к кооперативу. Среди представленных следствием потерпевших есть и конкуренты — бывшие пайщики, которые еще несколько лет назад вышли из «Бест Вей» и создали «альтернативный» кооператив «Вера».
Следователи обосновывают арест счетов тем, что «деньги могут украсть», хотя они же сами как минимум участвуют во вполне криминальном использовании чужих денег, уже год не давая пайщикам ни приобрести квартиру за счет средств, которые они скопили на счете в кооперативе, ни забрать деньги, при этом предоставляя возможность банкам. Следователи уже год дают возможность банкам пользоваться счетами почти на 4 млрд рублей, притом постоянно пополняющимися, так как большинство пайщиков продолжает вносить паевые и членские взносы.
Парадокс в том, что новая история обиженных дольщиков (в данном случае пайщиков) всероссийского масштаба возникла не из-за того, что у организации не оказалось средств для выполнения обязательств, а из-за того, что следственная группа лишила ее возможности пользования средствами и прямо блокирует выполнение обязательств перед пайщиками.
Адвокатам кооператива несколько раз удавалось добиться снятия арестов с активов кооператива, однако следователи и банки просто игнорируют решения судов. Потом, пользуясь высоким статусом следователей главка, заходят с новым ходатайством об аресте через кабинет судьи и без участия представителей кооператива, и активы снова арестовываются — до следующей успешной апелляции адвокатов, после которой в дело вступают банки, по договоренности со следствием блокирующие счета в рамках процедур комплаенс, игнорируя решения судов.
Должностное преступление
Действия следственной группы не могут квалифицироваться иначе, как должностное преступление, так как в них прослеживается заинтересованность в уничтожении кооператива. Фактически они сами организуют преступление, которое якобы расследуют, целенаправленно разрушают кооператив, а не спасают пайщиков, не нуждающихся в их спасении и 15 февраля проводящих всероссийскую акцию в защиту своего кооператива от следственной группы.
Адвокаты подавали в Следственный комитет заявление о преступлении, но оно было положено под сукно.
К службе не годны
Отдельная тема — вопиющая профнепригодность следственной группы. Иудичев в своих ответах на запросы ничтоже сумняшеся пишет, что Центральный банк признал кооператив финансовой пирамидой, хотя ЦБ, во-первых, не суд, чтобы делать такие «признания», во-вторых, такого признания просто не было: утверждение не соответствует действительности.
ЦБ внес кооператив в так называемый предупредительный список, являющийся инструментом информирования потребителей финансовых услуг об организациях, работа с которыми чревата для граждан рисками, и решение о включении в этот список состоялось после возбуждения уголовного дела, скорее всего, как раз на основе информации о его возбуждении.
Любимая тема следственной группы — якобы незаконность переименования кооператива из жилищного в потребительский: они не могут даже прочитать Жилищный кодекс, где черным по белому написано, что жилищный кооператив — один из видов жилищного. И подобных примеров некомпетентности и выдумок в деятельности следователей масса.
В итоге мы имеем шитое белыми нитками дело, наносящее колоссальный ущерб тысячам пайщиков кооператива, среди которых преобладают малообеспеченные люди.
Возникают вопросы: не пора ли министру Колокольцеву разобраться со своими сотрудниками, не пора ли главе Следкома Бастрыкину обратить внимание на должностные преступления в «соседнем» ведомстве?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://violitta1988.diary.ru/
https://www.haikudeck.com/presentations/TH7kPcfJPs
https://spaiider1979.diary.ru/
https://tubeteencam.com/user/viax1950/profile
https://permacultureglobal.org/users/58793-scott-reed
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/b262beach
https://permacultureglobal.org/users/60196-laura-shockey
http://www.babelcube.com/user/shannon-bailey
https://launchpad.net/~derkesth19601
https://chyoa.com/user/nuh651950
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiZZkF
https://imageevent.com/goliat1964
https://www.quia.com/profiles/anwoolford
https://rentry.org/79q4vndv
https://www.obesityhelp.com/members/abhsfgvay19761950/about_me/
My brother suggested I might like this website.
He was entirely right. This post actually made my day. You cann’t imagine simply how much time
I had spent for this info! Thanks!
Howdy! I could have sworn I’ve been to this blog before but after going through some of the
articles I realized it’s new to me. Anyhow, I’m definitely happy I stumbled
upon it and I’ll be book-marking it and checking back regularly!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/posh1969/profile
https://www.quia.com/profiles/summert593
https://anotepad.com/notes/6e9jh734
https://rentry.org/fve7kg8t
https://krikxe1958.diary.ru/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://anotepad.com/notes/ykrcam6h
https://haveagood.holiday/users/352266
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195416
https://cannabis.net/user/151231
https://ellak.gr/user/purple401999/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3194585
https://www.quia.com/profiles/dianasanchez560
https://ckaip1996.diary.ru/
https://rentry.org/yd5sron5
https://www.quia.com/profiles/sguidry134
bookmarked!!, I like your blog!
Excellent goods from you, man. I have understand your stuff previous to and you’re just too fantastic.
I actually like what you’ve acquired here, really like what
you are saying and the way in which you say it.
You make it enjoyable and you still take care of to keep it
sensible. I can not wait to read far more from you.
This is actually a wonderful website.
Hi there! This is my first visit to your blog! We are a team of volunteers and starting
a new initiative in a community in the same niche.
Your blog provided us valuable information to work
on. You have done a outstanding job!
I blog often and I seriously appreciate your content. The article has
really peaked my interest. I am going to take a note of your website and keep checking
for new details about once per week. I subscribed to your RSS feed as well.
Разбираю ситуацию вокруг “кооператива” Life is Good
порно жесток
Я решил написать этот пост, чтоб пройтись по доводам тех, кто привлекает новых “пайщиков” в структуры г-на Василенко (на деле – оплачивающих существование пирамиды Life is Good). Понятно, что, в этом сообществе крутятся сотни тех, кто занимается привлечением и теперь ЦБ с прокуратурой их оставят не только без заработка, но, возможно, придется отвечать за свои действия в суде. А деньги уже, наверняка, потрачены. Также есть люди, которые уже живут в квартире от LIG или выплатили некую сумму, они верят до конца и отказываются принимать ситуацию, в которой надо признать, что они могут остаться и без денег, и без квартиры. В любом случае, даже если кто-то живет в квартире от LIG, она ему не принадлежит и, в отличие от банковского залога, урегулировать вопрос не получится (их просто выселят через суд при банкротстве LIG, хотя ситуация скандальная и возможно государство что-то предпримет, чтоб сгладить проблему).Теперь по сути, что такое LIG.
Компания LIG – «Лайф-из-гуд» заявляет два основных направления деятельности: инвестиционные счета «Виста» в компании Hermes Management Ltd. и жилищный кооператив «Бест-вей» — альтернатива ипотеке.
Если верить консультантам «Лайф-из-гуд», доходность счетов «Виста» в компании «Гермес-менеджмент» последние годы держится в районе 20% годовых в долларах. Это довольно много (Уоррен Баффет позавидовал бы такой доходности), при этом непонятно, за счет чего достигается такая доходность.
Деятельность «Гермес-менеджмента» непрозрачна, на сайте не видно полноценных отчетов, нет сведений о структуре компании и ее руководстве.
«Гермес-менеджмент» привлекает российских клиентов через компанию «Лайф-из-гуд», но у него нет лицензии Центрального банка России на брокерскую деятельность или доверительное управление.
«Гермес-менеджмент» зарегистрирован в Белизе, офшорной зоне в Центральной Америке. Там же расположен офис компании. Еще в документах упоминается компания Hermes M в Шотландии. В случае проблем придется разбираться с зарубежной компанией, что делает задачу бесперспективной. Более того, учитывая реалии мошеннических схем, бывшие пайщики столкнутся еще и с лже-юристами, обещающими вернуть им вложенные деньги и конечно, останутся в крайне бедственном положении, с кредитами, без жилья и перспектив (у кого-то могут опуститься руки со всеми вытекающими).
Компания платит за привлечение новых клиентов, открывающих инвестиционные счета «Виста» (по сути – ничем не обеспеченные цифры на сайте. Заведение средств происходит через перевод средств на счета частных лиц(!), выполняющих роль консультантов LIG (а это сразу нарушает законы РФ и привлекает внимание регулятора), вывод – через перепродажу воздуха других буратинам. После внесения LIG в черный список, чаты структуры полны объявлений о продажи токенов Виста, но покупателей почти нет)) и вступающих в жилищный кооператив «Бест-вей». Система комиссионных выглядит такой разветвленной и продуманной, что становится понятно, что именно в ней суть компании, а не в инвестициях или альтернативе ипотеке.
Сам по себе сетевой маркетинг — вполне законный способ привлечения новых клиентов. Но когда он выглядит как основной вид деятельности, а клиентам при этом рассказывают про инвестиции, пассивный доход и успех в жизни, это наводит на очевидные выводы.
Суть предложения «Гермеса» такая: инвесторы открывают счет и перечисляют компании доллары или евро, а взамен получают пассивный доход. Минимальная сумма инвестиционного контракта — 10 тысяч долларов или евро. При этом первый взнос может быть 1130 долларов или 1100 евро, а оставшуюся сумму можно довнести позже.
Чтобы открыть счет «Виста», надо заполнить регистрационную форму на сайте «Лайф-из-гуд». При регистрации сайт просит поставить галочку «Я ознакомился и принимаю условия указанные в публичной оферте» (пунктуация сохранена).
По ссылке открывается не оферта, а договор об оказании услуг. В договоре не указаны название и реквизиты компании, с которой заключается договор. Кроме того, речь в нем идет не об управлении активами, а только об «образовательных и консультационных услугах об инвестиционных механизмах и способах управления капиталом». Это не похоже на обещанный пассивный доход от слова совсем. Таким образом, зазывалы в пирамиду пытаются уйти от ответственности.
жесткое гей порно
Чтобы открыть счет и пользоваться им, придется заплатить различные комиссии. Все они прописаны в договоре:
Комиссия за открытие счета: 100 евро или 130 долларов.
Ажио, или входная комиссия: 7% от суммы лицевого счета. На сайте есть пример расчета: при вложении 100 000 $ ажио составит 7000 $.
Комиссия за управление счетом: 1% в год от размера счета, взимается каждый месяц по 1/12.
Комиссия с прибыли: 20% от ежемесячной прибыли, «полученной от инвестиционных продуктов сторонних партнерских компаний или физических лиц, в результате рекомендаций и деятельности Исполнителя».
Итак, LIG состоит из двух основных компаний: Гермес, которая предлагает участникам инвестиционный доход (с идеей полученную прибыль направить на квартиру) и Бест-Вэй, являющаяся ЖНК. Что это такое?
Это Жилищный накопительный кооператив (общепринятое сокращение — ЖНК) — вид потребительского кооператива, сочетающий в себе качества финансового и жилищно-строительного кооператива. ЖНК работают в соответствии с № 215-ФЗ «О жилищных накопительных кооперативах». Целью деятельности жилищных накопительных кооперативов является удовлетворения потребностей членов кооператива в жилых помещениях путём объединения членами кооператива паевых взносов. Жилищный накопительный кооператив имеет право:
– привлекать и использовать денежные средства граждан на приобретение жилых помещений;
– вкладывать имеющиеся у него денежные средства в строительство жилых помещений (в том числе в многоквартирных домах), а также участвовать в строительстве жилых помещений в качестве застройщика или участника долевого строительства;
– приобретать жилые помещения;
– привлекать заёмные денежные средства, общая величина не должна превышать сорок процентов стоимости имущества кооператива.
Членами жилищного накопительного кооператива могут быть только граждане РФ. Жилищный накопительный кооператив создаётся по инициативе не менее чем пятидесяти человек и не более чем пяти тысяч человек. Контроль за деятельностью кооператива по привлечению и использованию денежных средств граждан на приобретение жилых помещений, а также за соблюдением кооперативом требований действующего законодательства Российской Федерации осуществляет Центральный банк Российской Федерации.
Уже отсюда понятно, что заявления заинтересованных лиц из структур LIG о том, что ЦБ не в праве вмешиваться в деятельность собственно LIG лишены оснований. Может и вмешался.
Если говорить о цифрах, то на 13 декабря 2021 года в LIG было свыше 19 000 пайщиков изо всех уголков РФ. LIG выкупил и передал в безвозмездное пользование пайщикам примерно 2 500 объектов недвижимости, из них по 247 пайщики полностью выплатили пай, оформив недвижимость в собственность. Интересная статистика – 19 000 вступили, заплатив членский взнос в 160 000 рублей, но лишь 8 часть из них накопила необходимые 35% для получения недвижимости при том, что фактически недвижимость их собственностью не является. Заявления о том, что получившие недвижимость прописаны в квартирах – это пережитки советского прошлого и, вероятно, сознательное введение остальных в заблуждение. В современной РФ не существует института прописки, есть регистрация, постоянная и временная. Есть найм жилья и есть собственность. Допускаю, что какая-то часть людей все еще проживает в государственных квартирах и не оформила их в собственность, но это – давно не норма, как это было в СССР (где собственность как раз была редким явлением).
По сути кооператив — посредник между покупателем и продавцом жилья, а это увеличивает риски. Приходится полагаться на то, что кооператив соблюдает закон и что он не закроется до того, как вы оформите квартиру на себя.
Даже если все по закону, риск потерять деньги остается. Если кооператив по каким-то причинам перестанет работать, его собственность распродадут. Сначала он рассчитается с кредиторами и только затем вернет деньги пайщикам — пропорционально их паевым взносам. Хватит ли у кооператива денег, чтобы вернуть все паевые взносы, неизвестно.
Разбираю ситуацию вокруг “кооператива” Life is Good
русское гей порно
Я решил написать этот пост, чтоб пройтись по доводам тех, кто привлекает новых “пайщиков” в структуры г-на Василенко (на деле – оплачивающих существование пирамиды Life is Good). Понятно, что, в этом сообществе крутятся сотни тех, кто занимается привлечением и теперь ЦБ с прокуратурой их оставят не только без заработка, но, возможно, придется отвечать за свои действия в суде. А деньги уже, наверняка, потрачены. Также есть люди, которые уже живут в квартире от LIG или выплатили некую сумму, они верят до конца и отказываются принимать ситуацию, в которой надо признать, что они могут остаться и без денег, и без квартиры. В любом случае, даже если кто-то живет в квартире от LIG, она ему не принадлежит и, в отличие от банковского залога, урегулировать вопрос не получится (их просто выселят через суд при банкротстве LIG, хотя ситуация скандальная и возможно государство что-то предпримет, чтоб сгладить проблему).Теперь по сути, что такое LIG.
Компания LIG – «Лайф-из-гуд» заявляет два основных направления деятельности: инвестиционные счета «Виста» в компании Hermes Management Ltd. и жилищный кооператив «Бест-вей» — альтернатива ипотеке.
Если верить консультантам «Лайф-из-гуд», доходность счетов «Виста» в компании «Гермес-менеджмент» последние годы держится в районе 20% годовых в долларах. Это довольно много (Уоррен Баффет позавидовал бы такой доходности), при этом непонятно, за счет чего достигается такая доходность.
Деятельность «Гермес-менеджмента» непрозрачна, на сайте не видно полноценных отчетов, нет сведений о структуре компании и ее руководстве.
«Гермес-менеджмент» привлекает российских клиентов через компанию «Лайф-из-гуд», но у него нет лицензии Центрального банка России на брокерскую деятельность или доверительное управление.
«Гермес-менеджмент» зарегистрирован в Белизе, офшорной зоне в Центральной Америке. Там же расположен офис компании. Еще в документах упоминается компания Hermes M в Шотландии. В случае проблем придется разбираться с зарубежной компанией, что делает задачу бесперспективной. Более того, учитывая реалии мошеннических схем, бывшие пайщики столкнутся еще и с лже-юристами, обещающими вернуть им вложенные деньги и конечно, останутся в крайне бедственном положении, с кредитами, без жилья и перспектив (у кого-то могут опуститься руки со всеми вытекающими).
Компания платит за привлечение новых клиентов, открывающих инвестиционные счета «Виста» (по сути – ничем не обеспеченные цифры на сайте. Заведение средств происходит через перевод средств на счета частных лиц(!), выполняющих роль консультантов LIG (а это сразу нарушает законы РФ и привлекает внимание регулятора), вывод – через перепродажу воздуха других буратинам. После внесения LIG в черный список, чаты структуры полны объявлений о продажи токенов Виста, но покупателей почти нет)) и вступающих в жилищный кооператив «Бест-вей». Система комиссионных выглядит такой разветвленной и продуманной, что становится понятно, что именно в ней суть компании, а не в инвестициях или альтернативе ипотеке.
Сам по себе сетевой маркетинг — вполне законный способ привлечения новых клиентов. Но когда он выглядит как основной вид деятельности, а клиентам при этом рассказывают про инвестиции, пассивный доход и успех в жизни, это наводит на очевидные выводы.
Суть предложения «Гермеса» такая: инвесторы открывают счет и перечисляют компании доллары или евро, а взамен получают пассивный доход. Минимальная сумма инвестиционного контракта — 10 тысяч долларов или евро. При этом первый взнос может быть 1130 долларов или 1100 евро, а оставшуюся сумму можно довнести позже.
Чтобы открыть счет «Виста», надо заполнить регистрационную форму на сайте «Лайф-из-гуд». При регистрации сайт просит поставить галочку «Я ознакомился и принимаю условия указанные в публичной оферте» (пунктуация сохранена).
По ссылке открывается не оферта, а договор об оказании услуг. В договоре не указаны название и реквизиты компании, с которой заключается договор. Кроме того, речь в нем идет не об управлении активами, а только об «образовательных и консультационных услугах об инвестиционных механизмах и способах управления капиталом». Это не похоже на обещанный пассивный доход от слова совсем. Таким образом, зазывалы в пирамиду пытаются уйти от ответственности.
гей секс порно
Чтобы открыть счет и пользоваться им, придется заплатить различные комиссии. Все они прописаны в договоре:
Комиссия за открытие счета: 100 евро или 130 долларов.
Ажио, или входная комиссия: 7% от суммы лицевого счета. На сайте есть пример расчета: при вложении 100 000 $ ажио составит 7000 $.
Комиссия за управление счетом: 1% в год от размера счета, взимается каждый месяц по 1/12.
Комиссия с прибыли: 20% от ежемесячной прибыли, «полученной от инвестиционных продуктов сторонних партнерских компаний или физических лиц, в результате рекомендаций и деятельности Исполнителя».
Итак, LIG состоит из двух основных компаний: Гермес, которая предлагает участникам инвестиционный доход (с идеей полученную прибыль направить на квартиру) и Бест-Вэй, являющаяся ЖНК. Что это такое?
Это Жилищный накопительный кооператив (общепринятое сокращение — ЖНК) — вид потребительского кооператива, сочетающий в себе качества финансового и жилищно-строительного кооператива. ЖНК работают в соответствии с № 215-ФЗ «О жилищных накопительных кооперативах». Целью деятельности жилищных накопительных кооперативов является удовлетворения потребностей членов кооператива в жилых помещениях путём объединения членами кооператива паевых взносов. Жилищный накопительный кооператив имеет право:
– привлекать и использовать денежные средства граждан на приобретение жилых помещений;
– вкладывать имеющиеся у него денежные средства в строительство жилых помещений (в том числе в многоквартирных домах), а также участвовать в строительстве жилых помещений в качестве застройщика или участника долевого строительства;
– приобретать жилые помещения;
– привлекать заёмные денежные средства, общая величина не должна превышать сорок процентов стоимости имущества кооператива.
Членами жилищного накопительного кооператива могут быть только граждане РФ. Жилищный накопительный кооператив создаётся по инициативе не менее чем пятидесяти человек и не более чем пяти тысяч человек. Контроль за деятельностью кооператива по привлечению и использованию денежных средств граждан на приобретение жилых помещений, а также за соблюдением кооперативом требований действующего законодательства Российской Федерации осуществляет Центральный банк Российской Федерации.
Уже отсюда понятно, что заявления заинтересованных лиц из структур LIG о том, что ЦБ не в праве вмешиваться в деятельность собственно LIG лишены оснований. Может и вмешался.
Если говорить о цифрах, то на 13 декабря 2021 года в LIG было свыше 19 000 пайщиков изо всех уголков РФ. LIG выкупил и передал в безвозмездное пользование пайщикам примерно 2 500 объектов недвижимости, из них по 247 пайщики полностью выплатили пай, оформив недвижимость в собственность. Интересная статистика – 19 000 вступили, заплатив членский взнос в 160 000 рублей, но лишь 8 часть из них накопила необходимые 35% для получения недвижимости при том, что фактически недвижимость их собственностью не является. Заявления о том, что получившие недвижимость прописаны в квартирах – это пережитки советского прошлого и, вероятно, сознательное введение остальных в заблуждение. В современной РФ не существует института прописки, есть регистрация, постоянная и временная. Есть найм жилья и есть собственность. Допускаю, что какая-то часть людей все еще проживает в государственных квартирах и не оформила их в собственность, но это – давно не норма, как это было в СССР (где собственность как раз была редким явлением).
По сути кооператив — посредник между покупателем и продавцом жилья, а это увеличивает риски. Приходится полагаться на то, что кооператив соблюдает закон и что он не закроется до того, как вы оформите квартиру на себя.
Даже если все по закону, риск потерять деньги остается. Если кооператив по каким-то причинам перестанет работать, его собственность распродадут. Сначала он рассчитается с кредиторами и только затем вернет деньги пайщикам — пропорционально их паевым взносам. Хватит ли у кооператива денег, чтобы вернуть все паевые взносы, неизвестно.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3115404.html
https://anotepad.com/notes/4ieq8tcj
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3194993
https://www.quia.com/profiles/midavenport338
https://rentry.org/iwgohbm2
Actually when someone doesn’t know then its up to other
viewers that they will assist, so here it occurs.
Hi! I just would like to offer you a huge thumbs up for your
excellent info you have right here on this post.
I am coming back to your web site for more soon.
Thanks for finally writing about > Fornix (crus)/ Stria Terminalis –
fxst | Whole Brain Protocol for Tractography with Empirical MRI < Liked it!
This is the right webpage for everyone who wishes to understand this topic.
You understand so much its almost hard to argue with you (not that I personally will need to…HaHa).
You definitely put a new spin on a subject that’s been written about for many years.
Great stuff, just wonderful!
new capcut mod apk,capcut mod apk free download,capcut apk cracked download,capcut pro apk download 2021,capcut download mod,capcut premium free download,capcut pro apk 2020, icapcut.com capcut download pro,capcut 4k resolution download,capcut photo music video editor mod apk,download capcut pro mod apk no watermark,
유니콘 슬롯
Zhu Houzhao는 Zhang Yong을 가리 켰습니다. “아버지, 그 사람입니다.”
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195636
https://tubeteencam.com/user/l7pu3pak1973/profile
https://rentry.org/f75atroz
https://rentry.org/9hxo224q
https://tubeteencam.com/user/tahalkora1973/profile
Hey I know this is off topic but I was wondering if you knew of any widgets I could
add to my blog that automatically tweet my newest twitter updates.
I’ve been looking for a plug-in like this for quite some time and
was hoping maybe you would have some experience with something like this.
Please let me know if you run into anything. I truly enjoy reading
your blog and I look forward to your new updates.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/yubsrc7m
https://permacultureglobal.org/users/60065-stephanie-webber
https://www.hentai-foundry.com/user/kirill331986/profile
https://imageevent.com/geoiter1976
https://www.quia.com/profiles/reedab
magnificent issues altogether, you simply received a emblem
new reader. What may you suggest in regards to your put up that you simply made some days ago?
Any positive?
You are so cool! I do not think I’ve read through anything like this before.
So nice to find somebody with a few unique thoughts on this issue.
Seriously.. thank you for starting this up. This website is something that is needed on the web, someone with some originality!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/sasuke1001955/profile
http://www.nfomedia.com/profile?uid=rOiZcgI
https://haveagood.holiday/users/351707
https://haveagood.holiday/users/350287
https://imageevent.com/kiska20091986
My brother recommended I would possibly like this web site.
He was entirely right. This post truly made my day.
You can not consider just how a lot time I had spent for this information! Thanks!
I read this piece of writing completely on the topic of the comparison of most
up-to-date and earlier technologies, it’s awesome article.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/jeremy-monroe
https://rentry.org/7zsn9ozc
https://n3koo1990.diary.ru/
https://www.quia.com/profiles/na150harris
https://rentry.org/kdepfno7
Write more, thats all I have to say. Literally,
it seems as though you relied on the video to make your point.
You obviously know what youre talking about, why waste your
intelligence on just posting videos to your blog when you could be giving us something informative to read?
I was suggested this blog via my cousin. I’m now not sure whether or not this post is written by way of him as no one
else know such designated about my trouble. You are
wonderful! Thank you!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/moonnomad1986
http://www.babelcube.com/user/michael-ortiz-1
https://imageevent.com/kaitel1952
https://anotepad.com/notes/pbxbihx8
https://chyoa.com/user/calips19751960
ОБЩЕСТВО «Транс Инвест» – единая экономика да мультимодальные грузоперевозки по России c 2012 лета
https://ticargo.ru/
Объединение узлов от обществе БИОН. Проводим объединение аграрных участков. ЯЗЫК нас хоть наложить запрет рециклирование грунтов а также земляных мест, что-что также энергообъединение площадей в течение СНТ.
https://bion-online.ru/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/ooppe1966/about_me/
https://anotepad.com/notes/hemrkgq6
https://cannabis.net/user/151143
http://www.babelcube.com/user/lisa-young
https://tubeteencam.com/user/volf6661981/profile
I will right away grab your rss as I can not in finding your email subscription link or newsletter service.
Do you have any? Kindly permit me understand in order that I may subscribe.
Thanks.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151714
https://haveagood.holiday/users/351352
https://imageevent.com/igorjan1963
https://cannabis.net/user/151926
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195136
KMSpico: What is it?
vk cc 56f2xq
Operating systems and Office suites are among the primary Microsoft software items that still need to be paid for. Some consumers may find alternate activation methods due to the perceived expensive cost of these items. There may be restrictions, unforeseen interruptions, and persistent activation prompts if these items are installed without being properly activated.
Our KMSpico app was created as a solution to this issue. By using this program, customers may access all of the functionality of Microsoft products and simplify the activation procedure.
KMSPico is a universal activator designed to optimize the process of generating and registering license codes for Windows and Office. Functionally, it is similar to key generators, but with the additional possibility of automatic integration of codes directly into the system. It is worth paying attention to the versatility of the tool, which distinguishes it from similar activators.
The above discussion primarily focused on the core KMS activator, the Pico app. Understanding what the program is, we can briefly mention KMSAuto, a tool with a simpler interface.
By using the KMSPico tool, you can setup Windows&Office for lifetime activation. This is an essential tool for anybody looking to reveal improved features and go beyond limitations. Although it is possible to buy a Windows or Office key.
KMSPico 11 (last update 2024) allows you to activate Windows 11/10/8 (8.1)/7 and Office 2021/2019/2016/2013/2010 for FREE.
KMSpico Download | Official KMS Website New July 2024
windows loader скачать
Are you looking for the best tool to activate your Windows & Office? Then you should download and install KMSpico, as it is one of the best tools everyone should have. In this article, I will tell you everything about this fantastic tool, even though I will also tell you if this is safe to use.
In this case, don’t forget to read this article until the end, so you don’t miss any critical information. This guide is for both beginners and experts as I came up with some of the rumours spreading throughout the internet.
Perhaps before we move towards downloading or installing a section, we must first understand this tool. You should check out the guide below on this tool and how it works; if you already know about it, you can move to another section.
What is KMSPico?
KMPico is a tool that is used to activate or get a license for Microsft Windows as well as for MS Office. It was developed by one of the most famous developers named, Team Daz. However, it is entirely free to use. There is no need to purchase it or spend money downloading it. This works on the principle of Microsft’s feature named Key Management Server, a.k.a KMS (KMSPico named derived from it).
The feature is used for vast companies with many machines in their place. In this way, it is hard to buy a Windows License for each device,, which is why KMS introduced. Now a company has to buy a KMS server for them and use it when they can get a license for all their machines.
However, this tool also works on it, and similarly, it creates a server on your machine and makes it look like a part of that server. One thing different is that this tool only keeps the product activated for 180 days. This is why it keeps running on your machine, renews the license keys after 180 days, and makes it a permanent activation.
KMSAuto Net
Microsoft Toolkit
Windows Loader
Windows 10 Activator
Features
We already know what this tool means, so let’s talk about some of the features you are getting along with KMSPico. Reading this will surely help you understand whether you are downloading the correct file.
Ok, so here are some of the features that KMSPico provides:
Activate Windows & Office
We have already talked about this earlier, as using this tool, you will get the installation key for both Microsoft Products. Whether it is Windows or Office, you can get a license in no time; however, this supports various versions.
Supports Multi-Arch
Since this supports both products, it doesn’t mean you have to download separate versions for each arch. One version is enough, and you can get the license for both x32-bit or even the x64-bit.
It Is Free To Use
Undoubtedly, everything developed by Team Daz costs nothing to us. Similarly, using this tool won’t cost you either, as it is entirely free. Other than this, it doesn’t come with any ads, so using it won’t be any trouble.
Permanent License
Due to the KMS server, this tool installs on our PC, we will get the license key for the rest of our lives. This is because the license automatically renews after a few days. To keep it permanent, you must connect your machine to the internet once 180 days.
Virus Free
Now comes the main feature of this tool that makes it famous among others. KMSPico is 100% pure and clean from such viruses or trojans. The Virus Total scans it before uploading to ensure it doesn’t harm our visitors.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151628
https://rentry.org/yrnx34oh
https://rentry.org/audk39es
https://haveagood.holiday/users/351634
https://rentry.org/2tf7xfqn
https://gadalika.ru/ –
Всем здравия, кто посетил мою страничку. Меня зовут Анна Тимофеева. И я хочу поделиться с вами своими способностями. Как и все потомственные целители, я унаследовала свой Дар по крови от пробабушки, известной в России целительницы. Ее звали Анной, в неё честь и я ношу это имя. Корни нашего рода уходят глубоко во времена, когда целительство было обычным явлением. Прабабушка была дочерью священника. С юности она…
I’m impressed, I have to admit. Rarely do I come across a blog
that’s equally educative and amusing, and let me tell you, you’ve hit
the nail on the head. The issue is something that
too few men and women are speaking intelligently about.
I am very happy that I found this in my search for something concerning this.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3115569.html
https://permacultureglobal.org/users/59507-maria-jensen
https://cannabis.net/user/151763
https://rentry.org/arw54k3k
https://okwave.jp/profile/u3118227.html
https://gadalke.ru/ – Меня зовут Мария Степановна, я потомственная ясновидяшая, предсказательница и знаю, зачем вы здесь. Уверена, вы получите ответ и помощь на моем официальном сайте. Каждый день сотни людей со сложными жизненными проблемами пытаются отыскать ответ на вопрос где найти настоящую хорошую гадалку с отзывами, проверенную многими людьми. Гадалку, которая реально помогает. По-разному называют нас: ясновидящая, маг, экстресенс и даже ведьмой назовут несведущие. Я не обижаюсь. Ведь все одно это: Дар, данный Богом. Шарлатанов, выдающих себя за гадалок теперь много стало. Да и гадание не сложная наука, научиться можно при должном упорстве и труде. Сейчас кто на чем гадает, кто на таро, кто на кофейной гуще, на рунах или на воске. Гадают, да не помогают. Потому как не по крови дано, а через ум вошло. А знать, ни порчу снять, ни мужа вернуть не сумеют — а я помогу!
Heya! I just wanted to ask if you ever have any trouble with hackers?
My last blog (wordpress) was hacked and I ended
up losing many months of hard work due to no back up. Do you have any methods
to stop hackers?
Interesting blog! Is your theme custom made or did you download it from somewhere?
A design like yours with a few simple adjustements
would really make my blog shine. Please let me know where you got your theme.
Cheers
В чёрный список пирамид и лохотронов 31 августа 2022 года внесены следующие организации, обладающие признаками финансовой пирамиды или признаками мошенничества.
мальчик гей
Etihad Rail (etihadrail-ae.com)
Club Unite To Live, Life Is Good, Unite To Live (uniteto.live). Новый домен пирамиды находящегося в розыске Романа Василенко.
TopBoom, «платформа хедж-фонда, специализирующегося на сделках с обратным исходом спортивного события» (topboom.vip, t.me/TopBoom001)
Change Team, мошенники указывают реквизиты чужого юрлица ООО «АТТИС ГРУПП» (ОРГН 1207700314128, ИНН 9702022065), телеграм-канал «Change Team: Главный Канал» (change-team.ru). Фальшивый обменник раньше назывался C Exchange.
Ещё лохотроны:
Scam! Чёрный список пирамид и лохотронов и отзывы
В чёрный список пирамид и лохотронов включены организации, имеющие признаки финансовой пирамиды по классификации Центрального банка; организации, обладающие признаками матричной пирамиды (массовый привод друзей, «столы»); сборы денег с пенсионеров и прочих незащищённых слоёв населения; хайпы; организации, имеющие негативные отзывы; структуры, оказывающие финансовые услуги без лицензии и прочие лохотроны за исключением псевдоброкеров, кооперативов (с ними тоже не рекомендуем связываться) и некоторых других категорий.
В чёрный список пирамид и лохотронов 31 августа 2022 года внесены следующие организации, обладающие признаками финансовой пирамиды или признаками мошенничества.
[url=https://rating-market.com/de/rejting-cfd-brokerov/unite-to-live-otzyvy-o-brokere-v-2022-godu?ysclid=ly1suv5j4q366928553]раз анальный секс[/url]
Etihad Rail (etihadrail-ae.com)
Club Unite To Live, Life Is Good, Unite To Live (uniteto.live). Новый домен пирамиды находящегося в розыске Романа Василенко.
TopBoom, «платформа хедж-фонда, специализирующегося на сделках с обратным исходом спортивного события» (topboom.vip, t.me/TopBoom001)
Change Team, мошенники указывают реквизиты чужого юрлица ООО «АТТИС ГРУПП» (ОРГН 1207700314128, ИНН 9702022065), телеграм-канал «Change Team: Главный Канал» (change-team.ru). Фальшивый обменник раньше назывался C Exchange.
Ещё лохотроны:
Scam! Чёрный список пирамид и лохотронов и отзывы
В чёрный список пирамид и лохотронов включены организации, имеющие признаки финансовой пирамиды по классификации Центрального банка; организации, обладающие признаками матричной пирамиды (массовый привод друзей, «столы»); сборы денег с пенсионеров и прочих незащищённых слоёв населения; хайпы; организации, имеющие негативные отзывы; структуры, оказывающие финансовые услуги без лицензии и прочие лохотроны за исключением псевдоброкеров, кооперативов (с ними тоже не рекомендуем связываться) и некоторых других категорий.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/sadsadq1954/profile
https://avallon1991.bandcamp.com/album/a-bewitched-housewife-part-2
https://www.metal-archives.com/users/rtytp198019971963
https://rentry.org/uvhuzzkm
https://wwoww1973.bandcamp.com/album/the-agreement-1
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/ba3ndeed
https://permacultureglobal.org/users/59507-maria-jensen
https://chyoa.com/user/fauler1994
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3197148
https://okwave.jp/profile/u3115867.html
Keep this going please, great job!
Hi everyone, it’s my first visit at this web page, and article is
really fruitful in favor of me, keep up posting these posts.
mehndi design for 9 years old girl
mehndi design 9t9 arts
1st anniversary mehndi design
mehndi designs of 90
12345 mehndi design
It’s hard to find educated people about this topic, but you seem like you know what you’re talking
about! Thanks
I appreciate, result in I found exactly what I was looking for.
You’ve ended my 4 day lengthy hunt! God Bless you man. Have a nice day.
Bye
Awesome postings, Thanks!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/kcavuzbArG
https://okwave.jp/profile/u3118207.html
https://cannabis.net/user/151283
https://haveagood.holiday/users/350781
https://cannabis.net/user/152196
You explained it very well.
Feel free to surf to my web page: http://www.dbulut.com/question/you-can-thank-us-later-4-reasons-to-stop-%ec%b6%9c%ec%9e%a5%ec%95%88%eb%a7%88ing-now-9/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/artemkagg1992/about_me/
https://tubeteencam.com/user/lahardie1961/profile
https://www.haikudeck.com/presentations/ckjOEhDMxC
https://permacultureglobal.org/users/60046-bruce-millsaps
https://permacultureglobal.org/users/59507-maria-jensen
Have you ever thought about creating an e-book or guest authoring on other websites?
I have a blog based on the same topics you discuss and would really like
to have you share some stories/information. I know my viewers would appreciate your
work. If you are even remotely interested, feel free to send
me an e-mail.
Fantastic website. Lots of useful information here. I am
sending it to some buddies ans also sharing in delicious.
And naturally, thank you to your effort!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3115275.html
https://www.obesityhelp.com/members/miiz1968/about_me/
https://rentry.org/282zse8r
https://chyoa.com/user/artshyz1974
https://imageevent.com/toffo121978
Hurrah, that’s what I was looking for, what a information! existing here at this webpage, thanks
admin of this website.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/anarta1974
https://limwej1998.bandcamp.com/album/first-meeting-with-micheal
https://imageevent.com/lirina1981
https://haveagood.holiday/users/350068
https://tubeteencam.com/user/dikiizan1954/profile
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/fQN6BWOjWy
https://chyoa.com/user/diavol721960
https://permacultureglobal.org/users/60295-laura-collins
https://tubeteencam.com/user/koza8719541999/profile
https://www.quia.com/profiles/brent183se
Good day! Do you know if they make any plugins to help with
SEO? I’m trying to get my blog to rank for some targeted keywords but I’m not seeing very good results.
If you know of any please share. Kudos!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/otvarka1957/profile
https://www.obesityhelp.com/members/saiiiia1972/about_me/
https://torny1955.diary.ru/
https://www.hentai-foundry.com/user/ruyzaki1962/profile
https://imageevent.com/macgregor1964
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://darkgrey1971.bandcamp.com/album/the-sexy-stranger
https://cannabis.net/user/150807
https://okwave.jp/profile/u3116960.html
https://rentry.org/xnh977wx
https://www.hentai-foundry.com/user/wilhelmin1958/profile
obviously like your web-site however you need to take a look at the spelling on quite a few of
your posts. Many of them are rife with spelling problems and I in finding it very troublesome to inform
the reality then again I will definitely come again again.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/illiad1989
https://imageevent.com/nordmc1963
https://chyoa.com/user/sorvento1977
https://permacultureglobal.org/users/59796-cindy-logan
https://permacultureglobal.org/users/60237-carrie-smith
Hi my family member! I want to say that this post is awesome, great written and
include approximately all important infos. I would like to peer more posts like this .
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://okwave.jp/profile/u3115904.html
https://shikarnay1974.diary.ru/
http://www.nfomedia.com/profile?uid=rOiYahC
https://haveagood.holiday/users/350875
https://www.haikudeck.com/presentations/rRY6RPewZc
Об агентстве недвижимости СЦН
Агентство недвижимости Саровский Центр Недвижимости (СЦН) занимает лидирующие позиции среди профессиональных компаний в городе Саров. Мы оказываем квалифицированную поддержку клиентам, желающим приобрести, реализовать или обменять квартиру, или другие объекты недвижимости, а также осуществить разнообразные сделки в этой сфере. Наше агентство заключает официальные договоры, и оказываем услуги по прейскуранту.
купить квартиру в сарове
Опытные риэлторы в нашей команде обладают значительным опытом в области недвижимости и помогут вам найти идеальное жилье на вторичном рынке Сарова, а также подобрать вторичку и новостройки по всей России. Мы работаем с крупным всероссийским агентством недвижимости, что позволяет нам оказывать высококачественные услуги на рынке недвижимости как в пределах России, так и за рубежом.
Наше агентство предлагает выгодные условия по страхованию ипотеки, а также специальные скидки при оформлении ипотеки для партнеров банка. Мы установили партнерские отношения с известными крупными страховыми компаниями и банками, что обеспечивает нашим клиентам надежность и прозрачность процесса сделки. Юрист с двадцатилетним стажем, Антон Марсов, обеспечивает правовую защиту интересов наших клиентов на каждом этапе взаимодействия.
Мы также предлагаем услуги по ремонту и отделке квартир, а наш дизайнер интерьеров поможет воплотить в жизнь ваши дизайнерские идеи. Если вы ищете квартиру в Сарове для инвестиций, мы найдем для вас оптимальное решение. Мы также готовы рассмотреть сотрудничество с инвесторами для совместных проектов в сфере недвижимости. Наша профессиональная бригада ремонтников гарантирует высокое качество выполнения работ.
Не стесняйтесь обращаться к нам, если вам необходима помощь в приобретении или реализации недвижимости в Сарове. СЦН – ваш партнер по всем вопросам недвижимости в городе и за его пределами.
квартиры в сарове
https://сцн.рф
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/pwuzotfx
https://tubeteencam.com/user/esenya1967/profile
https://anotepad.com/notes/4ddeb5sk
http://www.babelcube.com/user/alisha-delgado
https://ddreterte1999.diary.ru/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/nice20001999/about_me/
https://anotepad.com/notes/mt7mb7yp
https://imageevent.com/maximkals1961
https://okwave.jp/profile/u3115324.html
https://haveagood.holiday/users/351461
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://chyoa.com/user/ahojki1954
http://www.nfomedia.com/profile?uid=rOiYceG
https://permacultureglobal.org/users/59809-johnny-edwards
http://www.nfomedia.com/profile?uid=rOiZcbE
https://chyoa.com/user/aleha281988
Звон Колокольцева
Бест Вей
Поздравляем вас, гражданин министр, соврамши!
Выступая прошлой осенью в Совете Федерации, министр внутренних дел Владимир Колокольцев рассказывал, о так называемом уголовном деле «Лайф-из-Гуд» – «Гермес» – «Бест Вей», обещал миллиарды рублей ущерба и десятки тысяч потерпевших. Пресс-служба МВД под руководством его боевой подруги Ирины Волк заявила о том, что вскрыта деятельность крупнейшей в истории России финансовой пирамиды.
Однако в уголовном деле, расследованном или, вернее сказать, изготовленном ГСУ питерского главка МВД, которое в феврале начал рассматривать Приморский районный суд Санкт-Петербурга, 282 млн рублей ущерба и 221 лицо, признанное следствием потерпевшим: никаких миллиардов и десятков тысяч потерпевших.
Министр и его Волк публично солгали – на основе информации, переданной замначальника ГСУ руководителем следственной группы полковником юстиции А.Н. Винокуровым, фактически даже руководившей СГ его заместительницей майором, а затем подполковником юстиции Е.А. Сапетовой по материалам, состряпанным опером УЭБиПК питерского главка МВД майором полиции А.Ю. Машевским, при попустительстве (или соучастии?) начальника ГСУ Негрозова и замначальника Следственного департамента федерального министерства Вохмянина. Полковник и подполковник с примерно пятого-шестого уровня иерархии МВД виляет генералом полиции Российской Федерации, постоянным членом Совета безопасности России – это позор для государства.
Так называемые потерпевшие и реально пострадавшие
Большинство «потерпевших» на суде заявляют суммы около 1–2 млн рублей, при этом в ходе судебного следствия выясняется, что они, как правило, получали немалый доход, причем, они еще и налоговые/валютные преступники, так как этот доход не декларировали. Среди «потерпевших» есть граждане, заявляющие смехотворные суммы в 50–70 тыс., то есть количество потерпевших специально накручивалось следствием. И даже с этим накручиванием удалось набрать так мало – учитывая, что у «Гермеса», по данным самого же следствия, более 200 тыс. клиентов в России, а в кооперативе «Бест Вей» – около 20 тыс. пайщиков.
То есть большинство и клиентов «Гермеса», и пайщиков кооператива не считают себя потерпевшими от деятельности этих организаций.
Судя по тысячам обращений во все инстанции, нескольким волнам митингов, прокатившихся по России, они считают себя потерпевшими от деятельности органов внутренних дел.
Ведь именно завербованный питерским УЭБиПК сисадмин российской платежной системы «Гермеса» Набойченко заблокирован и разгромил в феврале 2022 года эту платежную систему, повесив на сайте дисклеймер: «Обращайтесь в правоохранительные органы», что на месяцы прекратило вывод средств. Именно действия правоохранительных органов в отношении компании до и после затруднили вывод средств. Дело в том, что для вывода средств многими использовался механизм p2p, позволяющий не платить комиссию, то есть для вывода средств нужно, чтобы кто-то вносил средства (что, понятно, резко сократилось из-за уголовного дела) и происходил обмен. Однако этот способ не единственный, вывод средств так или иначе осуществляется.
Тысячи пайщиков кооператива тем более не считают себя потерпевшими от его деятельности – потому что именно правоохранительные органы воспрепятствовали приобретению недвижимости с помощью кооператива, а она из-за более чем двухлетнего ареста его счетов, на которых около 4 млрд рублей, не может быть приобретена по прежней цене.
Именно правоохранительные органы прямо запрещают выплаты пайщикам кооператива, решившим забрать свой пай, – даже по исполнительным листам судов. И клиентам «Гермеса», и пайщикам кооператива правоохранительными органами нанесен колоссальный ущерб – и материальный, и моральный, который они намерены взыскать с государства.
«Следователи»-преступники должны сидеть в тюрьме
Весьма скромный результат следствия МВД был достигнут откровенно преступным путем.
1. Некоторые из преступлений следствия были фактически признаны судами. 1 декабря прошлого года Приморский районный суд города Санкт-Петербурга признал незаконным, нарушающим УПК фактический отказ кооперативу в ознакомлении с материалами уголовного дела.При рассмотрении дела в суде выяснилось, что следственная группа ГСУ питерского главка МВД, формально руководимая замначальника ГСУ полковником юстиции А.Н. Винокуровым, а фактически – подполковником юстиции Е.А. Сапетовой, подделала документы. Автор подделки – Сапетова – еще в феврале была уволена из ГСУ «по собственному желанию».
Уличенная адвокатами кооператива в нарушении УПК, следственная группа составила письмо об удовлетворении ходатайства задним числом и попыталась представить дело так, что кооператив не получил письмо по своей вине. Ложь была выявлена в том числе и с помощью системы электронного документооборота питерского главка МВД.
2. Подделка документов была вынужденным преступлением для сокрытия более серьезного: незаконного содержания под стражей. Следственная группа грубо нарушила права гражданских истцов и ответчиков, потому что без этого нарушения она не успела за 30 суток до истечения предельного срока содержания четверых обвиняемых под стражей начать ознакомление обвиняемых с материалами дела –а это было единственное основание продления им срока содержания под стражей свыше предельного.
Следственная группа из-за спешки даже толком не смогла завершить следственные действия, незаконно вела параллельное расследование по «резервному» делу, но позднее, отбросив стыд, из-за отсутствия материала для составления «нужного» обвинительного заключения, незаконно продолжила расследование «основного» дела, в том числе проводила следственные действия, которые, согласно УПК, невозможны после начала ознакомления обвиняемых с материалами дела. Все эти ухищрения были необходимы для того, чтобы ни в коем случае не выпускать обвиняемых и продолжать держать их в заложниках.
3. Де-факто происходит уголовное наказание неосужденных людей – четверо подсудимых уже второй год сидят в тюрьме. При этом наказываются явно ни в чем неповинные люди – технические сотрудники «Лайф-из-Гуд»: даже если предположить, что действительно работала пирамида – что опровергается показаниями свидетелей самого обвинения, которые сообщают суду, что компания «Гермес» хорошо работала, они были довольны получаемым доходом, и проблемы начались после того, как российская платежная система компании была обрушена завербованным полицией петербургским сисадмином компании Набойченко. Все подсудимые – технические сотрудники компании «Лайф-из-Гуд» и ни к каким управленческим решениям отношения никогда не имели. Их взяли в заложники для того, чтобы они дали показания на руководство компании.
4. Еще одно преступление – заведомо подложное постановление руководителя следственной группы А.Н. Винокурова о привлечении кооператива «Бест Вей» в качестве гражданского ответчика на 16 млрд рублей, тогда как сумма ущерба в уголовном деле –282 млн, и в деле нет ни одного искового заявления – даже на 100 рублей.
5. Следствие стимулировало двух особенно активных так называемых потерпевших написать заявления о моральном ущербе на миллиард (!) рублей каждое – исключительно для ложного обоснования ареста активов кооператива, но понятно, что это ничтожные документы, так как моральный ущерб во всех случаях, не связанных с причинением смерти, присуждается российскими судами в размере не более десятков тысяч рублей.
6. Ни один из «потерпевших» не доказал обоснованность своих претензий в гражданском суде. При этом арестованы активы кооператива почти на 4 млрд рублей и активы частных лиц на такую же сумму. Это не что иное, как попытка захвата активов при участии органов внутренних дел некой заинтересованной группой клиентов «Гермеса» – необязательно из числа «потерпевших», то есть коррупционное преступление, которое упорно игнорирует ГУСБ МВД.
Механизм для такого захвата есть – это передача средств под управление Федерального общественно-государственного фонда по защите прав вкладчиков и акционеров. Понятно, почему не ограничиваются активами подсудимых и обвиняемых: этого недостаточно для удовлетворения аппетитов тех, кто стоит за заказным уголовным делом. И понятно, почему одно юридическое лицо – кооператив «Бест Вей» – незаконно пытаются привлечь к ответственности за другое – компанию «Гермес»: активы «Гермеса» – за рубежом.
7. Следствие, а теперь и прокуратура совершают еще одно преступление: незаконно удерживает средства пайщиков кооператива, отказываясь их вернуть, то есть совершают хищение.
Министр, управляемый мафией
Итак, Колокольцев публично солгал Совету Федерации, выдумав пирамиду, выдумав многомиллиардный ущерб и десятки тысяч пострадавших, он – некомпетентный руководитель, абсолютно ведомый своей камарильей, стремящейся создавать громкие пиар-истории на пустом месте и с коррупционной выгодой для себя. Он вызвал социальный протест десятков тысяч пайщиков кооператива «Бест Вей» и клиентов компании «Гермес» и членов их семей – в большинстве своем представителей социально незащищенных слоев населения, на годы лишенных возможности пользоваться своими деньгами и приобрести недвижимость, на которую они собрали средства, – в том числе десятков участников СВО. Министр вредит в тылу тем, кто защищает страну на фронте.
Глава МВД находится под влиянием или даже контролем давно ставшей притчей во языцех питерской полицейской мафии. Ликвидация этой мафии, как и ликвидация преступной, коррупционной, вредящей социально-политической стабильности системы управления МВД, со стороны Колокольцева является важнейшей государственной задачей.
Звон Колокольцева
Бест Вей
Поздравляем вас, гражданин министр, соврамши!
Выступая прошлой осенью в Совете Федерации, министр внутренних дел Владимир Колокольцев рассказывал, о так называемом уголовном деле «Лайф-из-Гуд» – «Гермес» – «Бест Вей», обещал миллиарды рублей ущерба и десятки тысяч потерпевших. Пресс-служба МВД под руководством его боевой подруги Ирины Волк заявила о том, что вскрыта деятельность крупнейшей в истории России финансовой пирамиды.
Однако в уголовном деле, расследованном или, вернее сказать, изготовленном ГСУ питерского главка МВД, которое в феврале начал рассматривать Приморский районный суд Санкт-Петербурга, 282 млн рублей ущерба и 221 лицо, признанное следствием потерпевшим: никаких миллиардов и десятков тысяч потерпевших.
Министр и его Волк публично солгали – на основе информации, переданной замначальника ГСУ руководителем следственной группы полковником юстиции А.Н. Винокуровым, фактически даже руководившей СГ его заместительницей майором, а затем подполковником юстиции Е.А. Сапетовой по материалам, состряпанным опером УЭБиПК питерского главка МВД майором полиции А.Ю. Машевским, при попустительстве (или соучастии?) начальника ГСУ Негрозова и замначальника Следственного департамента федерального министерства Вохмянина. Полковник и подполковник с примерно пятого-шестого уровня иерархии МВД виляет генералом полиции Российской Федерации, постоянным членом Совета безопасности России – это позор для государства.
Так называемые потерпевшие и реально пострадавшие
Большинство «потерпевших» на суде заявляют суммы около 1–2 млн рублей, при этом в ходе судебного следствия выясняется, что они, как правило, получали немалый доход, причем, они еще и налоговые/валютные преступники, так как этот доход не декларировали. Среди «потерпевших» есть граждане, заявляющие смехотворные суммы в 50–70 тыс., то есть количество потерпевших специально накручивалось следствием. И даже с этим накручиванием удалось набрать так мало – учитывая, что у «Гермеса», по данным самого же следствия, более 200 тыс. клиентов в России, а в кооперативе «Бест Вей» – около 20 тыс. пайщиков.
То есть большинство и клиентов «Гермеса», и пайщиков кооператива не считают себя потерпевшими от деятельности этих организаций.
Судя по тысячам обращений во все инстанции, нескольким волнам митингов, прокатившихся по России, они считают себя потерпевшими от деятельности органов внутренних дел.
Ведь именно завербованный питерским УЭБиПК сисадмин российской платежной системы «Гермеса» Набойченко заблокирован и разгромил в феврале 2022 года эту платежную систему, повесив на сайте дисклеймер: «Обращайтесь в правоохранительные органы», что на месяцы прекратило вывод средств. Именно действия правоохранительных органов в отношении компании до и после затруднили вывод средств. Дело в том, что для вывода средств многими использовался механизм p2p, позволяющий не платить комиссию, то есть для вывода средств нужно, чтобы кто-то вносил средства (что, понятно, резко сократилось из-за уголовного дела) и происходил обмен. Однако этот способ не единственный, вывод средств так или иначе осуществляется.
Тысячи пайщиков кооператива тем более не считают себя потерпевшими от его деятельности – потому что именно правоохранительные органы воспрепятствовали приобретению недвижимости с помощью кооператива, а она из-за более чем двухлетнего ареста его счетов, на которых около 4 млрд рублей, не может быть приобретена по прежней цене.
Именно правоохранительные органы прямо запрещают выплаты пайщикам кооператива, решившим забрать свой пай, – даже по исполнительным листам судов. И клиентам «Гермеса», и пайщикам кооператива правоохранительными органами нанесен колоссальный ущерб – и материальный, и моральный, который они намерены взыскать с государства.
«Следователи»-преступники должны сидеть в тюрьме
Весьма скромный результат следствия МВД был достигнут откровенно преступным путем.
1. Некоторые из преступлений следствия были фактически признаны судами. 1 декабря прошлого года Приморский районный суд города Санкт-Петербурга признал незаконным, нарушающим УПК фактический отказ кооперативу в ознакомлении с материалами уголовного дела.При рассмотрении дела в суде выяснилось, что следственная группа ГСУ питерского главка МВД, формально руководимая замначальника ГСУ полковником юстиции А.Н. Винокуровым, а фактически – подполковником юстиции Е.А. Сапетовой, подделала документы. Автор подделки – Сапетова – еще в феврале была уволена из ГСУ «по собственному желанию».
Уличенная адвокатами кооператива в нарушении УПК, следственная группа составила письмо об удовлетворении ходатайства задним числом и попыталась представить дело так, что кооператив не получил письмо по своей вине. Ложь была выявлена в том числе и с помощью системы электронного документооборота питерского главка МВД.
2. Подделка документов была вынужденным преступлением для сокрытия более серьезного: незаконного содержания под стражей. Следственная группа грубо нарушила права гражданских истцов и ответчиков, потому что без этого нарушения она не успела за 30 суток до истечения предельного срока содержания четверых обвиняемых под стражей начать ознакомление обвиняемых с материалами дела –а это было единственное основание продления им срока содержания под стражей свыше предельного.
Следственная группа из-за спешки даже толком не смогла завершить следственные действия, незаконно вела параллельное расследование по «резервному» делу, но позднее, отбросив стыд, из-за отсутствия материала для составления «нужного» обвинительного заключения, незаконно продолжила расследование «основного» дела, в том числе проводила следственные действия, которые, согласно УПК, невозможны после начала ознакомления обвиняемых с материалами дела. Все эти ухищрения были необходимы для того, чтобы ни в коем случае не выпускать обвиняемых и продолжать держать их в заложниках.
3. Де-факто происходит уголовное наказание неосужденных людей – четверо подсудимых уже второй год сидят в тюрьме. При этом наказываются явно ни в чем неповинные люди – технические сотрудники «Лайф-из-Гуд»: даже если предположить, что действительно работала пирамида – что опровергается показаниями свидетелей самого обвинения, которые сообщают суду, что компания «Гермес» хорошо работала, они были довольны получаемым доходом, и проблемы начались после того, как российская платежная система компании была обрушена завербованным полицией петербургским сисадмином компании Набойченко. Все подсудимые – технические сотрудники компании «Лайф-из-Гуд» и ни к каким управленческим решениям отношения никогда не имели. Их взяли в заложники для того, чтобы они дали показания на руководство компании.
4. Еще одно преступление – заведомо подложное постановление руководителя следственной группы А.Н. Винокурова о привлечении кооператива «Бест Вей» в качестве гражданского ответчика на 16 млрд рублей, тогда как сумма ущерба в уголовном деле –282 млн, и в деле нет ни одного искового заявления – даже на 100 рублей.
5. Следствие стимулировало двух особенно активных так называемых потерпевших написать заявления о моральном ущербе на миллиард (!) рублей каждое – исключительно для ложного обоснования ареста активов кооператива, но понятно, что это ничтожные документы, так как моральный ущерб во всех случаях, не связанных с причинением смерти, присуждается российскими судами в размере не более десятков тысяч рублей.
6. Ни один из «потерпевших» не доказал обоснованность своих претензий в гражданском суде. При этом арестованы активы кооператива почти на 4 млрд рублей и активы частных лиц на такую же сумму. Это не что иное, как попытка захвата активов при участии органов внутренних дел некой заинтересованной группой клиентов «Гермеса» – необязательно из числа «потерпевших», то есть коррупционное преступление, которое упорно игнорирует ГУСБ МВД.
Механизм для такого захвата есть – это передача средств под управление Федерального общественно-государственного фонда по защите прав вкладчиков и акционеров. Понятно, почему не ограничиваются активами подсудимых и обвиняемых: этого недостаточно для удовлетворения аппетитов тех, кто стоит за заказным уголовным делом. И понятно, почему одно юридическое лицо – кооператив «Бест Вей» – незаконно пытаются привлечь к ответственности за другое – компанию «Гермес»: активы «Гермеса» – за рубежом.
7. Следствие, а теперь и прокуратура совершают еще одно преступление: незаконно удерживает средства пайщиков кооператива, отказываясь их вернуть, то есть совершают хищение.
Министр, управляемый мафией
Итак, Колокольцев публично солгал Совету Федерации, выдумав пирамиду, выдумав многомиллиардный ущерб и десятки тысяч пострадавших, он – некомпетентный руководитель, абсолютно ведомый своей камарильей, стремящейся создавать громкие пиар-истории на пустом месте и с коррупционной выгодой для себя. Он вызвал социальный протест десятков тысяч пайщиков кооператива «Бест Вей» и клиентов компании «Гермес» и членов их семей – в большинстве своем представителей социально незащищенных слоев населения, на годы лишенных возможности пользоваться своими деньгами и приобрести недвижимость, на которую они собрали средства, – в том числе десятков участников СВО. Министр вредит в тылу тем, кто защищает страну на фронте.
Глава МВД находится под влиянием или даже контролем давно ставшей притчей во языцех питерской полицейской мафии. Ликвидация этой мафии, как и ликвидация преступной, коррупционной, вредящей социально-политической стабильности системы управления МВД, со стороны Колокольцева является важнейшей государственной задачей.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/if89mzwg
https://okwave.jp/profile/u3115275.html
https://tubeteencam.com/user/niakria1983/profile
https://www.haikudeck.com/presentations/aZNJCjUXCi
https://www.obesityhelp.com/members/reddiska1953/about_me/
Hmm is anyone else having problems with the pictures on this blog loading?
I’m trying to determine if its a problem on my end or if it’s the blog.
Any responses would be greatly appreciated.
This article is very good, hopefully it will increase your ranking on Google
GITAR100
Hey! I just wanted to ask if you ever have any issues
with hackers? My last blog (wordpress) was hacked and I ended up
losing months of hard work due to no backup. Do you have any methods to
protect against hackers?
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.obesityhelp.com/members/pain471986/about_me/
https://www.hentai-foundry.com/user/fioloo21987/profile
http://www.babelcube.com/user/saad-campbell
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3194685
https://permacultureglobal.org/users/59018-kimberly-brewer
I do accept as true with all the concepts you’ve introduced for
your post. They’re really convincing and can certainly
work. Still, the posts are too brief for beginners.
Could you please prolong them a bit from subsequent time?
Thank you for the post.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/archangel1990
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196496
https://rentry.org/88a4in2r
https://doyo19871955.diary.ru/
https://okwave.jp/profile/u3116858.html
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.metal-archives.com/users/copcar219991970
https://okwave.jp/profile/u3115970.html
https://www.hentai-foundry.com/user/nobynagi1995/profile
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195120
http://www.nfomedia.com/profile?uid=rOjQfiB
I am now not positive where you are getting your information, but great topic.
I needs to spend some time studying more or understanding more.
Thank you for magnificent information I used to be in search of this information for my mission.
Ahaa, its fastidious dialogue concerning this post at this place at
this webpage, I have read all that, so now me also commenting here.
Why a rare image of one of Malaysia’s last tigers is giving conservationists hope
MEGA onion
Emmanuel Rondeau has photographed tigers across Asia for the past decade, from the remotest recesses of Siberia to the pristine valleys of Bhutan. But when he set out to photograph the tigers in the ancient rainforests of Malaysia, he had his doubts.
“We were really not sure that this was going to work,” says the French wildlife photographer. That’s because the country has just 150 tigers left, hidden across tens of thousands of square kilometers of dense rainforest.
https://me3ga-gl.com
m3ga.gl
“Tiger numbers in Malaysia have been going down, down, down, at an alarming rate,” says Rondeau. In the 1950s, Malaysia had around 3,000 tigers, but a combination of habitat loss, a decline in prey, and poaching decimated the population. By 2010, there were just 500 left, according to WWF, and the number has continued to fall.
The Malayan tiger is a subspecies native to Peninsular Malaysia, and it’s the smallest of the tiger subspecies in Southeast Asia.
“We are in this moment where, if things suddenly go bad, in five years the Malayan tiger could be a figure of the past, and it goes into the history books,” Rondeau adds.
Determined not to let that happen, Rondeau joined forces with WWF-Malaysia last year to profile the elusive big cat and put a face to the nation’s conservation work.
It took 12 weeks of preparations, eight cameras, 300 pounds of equipment, five months of patient photography and countless miles trekked through the 117,500-hectare Royal Belum State Park… but finally, in November, Rondeau got the shot that he hopes can inspire the next generation of conservationists.
https://mega555darknet5.com
мега сайт
“This image is the last image of the Malayan tiger — or it’s the first image of the return of the Malayan tiger,” he says.
Explore Iceland in 3 days with a curated itinerary.
Discover Reykjavik’s charm, delve into wilderness adventures, savor Icelandic cuisine, and
immerse in cultural treasures. Learn weather conditions, driving
laws, and full packing lists. Book your Icelandic adventure now!
This is the perfect web site for everyone who hopes to find out about this topic. You realize a whole lot its almost tough to argue with you (not that I actually would want to…HaHa). You certainly put a fresh spin on a topic that has been written about for years. Great stuff, just wonderful.
Tributes flood in for BBC sport commentator whose wife and daughters were killed in suspected crossbow attack
[url=https://freedmanclub.com/life-is-good-pochemu-proizoshjol-tehnicheskij-skam-proekta/]гей член[/url]
Figures from across the United Kingdom have offered their condolences to a BBC sport commentator, after his wife and two daughters were killed by an alleged crossbow attacker, in deaths that again drew attention to the epidemic of violence against women.
Carol Hunt, 61, wife of BBC horse racing commentator, John Hunt, and their daughters, Hannah Hunt, 28, and Louise Hunt, 25, died from injuries sustained in an attack in Bushey, just northwest of London, on Tuesday, according to police and Britain’s public broadcaster.
A 26-year-old suspect wanted in connection with the killings, and named as Kyle Clifford, was found by British police in Enfield, north London, on Wednesday, following a sprawling manhunt. There were no previous reports to the force over Clifford, who is in serious condition in hospital and is yet to speak with officers, Hertfordshire Police said in a statement.
A crossbow was recovered as part of the investigation, which police believe was used in a “targeted incident.”
Epidemic of violence against women
The killings of the three women rocked Britain, where mass murders are infrequent but violence against women and girls has been officially labeled as a national threat.
A woman is killed by a man every three days in the UK and one in four women will experience domestic violence in her lifetime, the United Nations’ special rapporteur on violence against women and girls, Reem Alsalem, said earlier this year.
Charities and human rights organizations have subsequently reiterated urgent demands to tackle femicide in the UK.
Tributes flood in for BBC sport commentator whose wife and daughters were killed in suspected crossbow attack
[url=https://newscryptocoin.com/news/chto-delat-postradavshim-ot-deyatelnosti-life-is-good-bestway-i-hermes-kotorye-osnoval-beglyj-roman-vasilenko/]смотреть порно жесток[/url]
Figures from across the United Kingdom have offered their condolences to a BBC sport commentator, after his wife and two daughters were killed by an alleged crossbow attacker, in deaths that again drew attention to the epidemic of violence against women.
Carol Hunt, 61, wife of BBC horse racing commentator, John Hunt, and their daughters, Hannah Hunt, 28, and Louise Hunt, 25, died from injuries sustained in an attack in Bushey, just northwest of London, on Tuesday, according to police and Britain’s public broadcaster.
A 26-year-old suspect wanted in connection with the killings, and named as Kyle Clifford, was found by British police in Enfield, north London, on Wednesday, following a sprawling manhunt. There were no previous reports to the force over Clifford, who is in serious condition in hospital and is yet to speak with officers, Hertfordshire Police said in a statement.
A crossbow was recovered as part of the investigation, which police believe was used in a “targeted incident.”
Epidemic of violence against women
The killings of the three women rocked Britain, where mass murders are infrequent but violence against women and girls has been officially labeled as a national threat.
A woman is killed by a man every three days in the UK and one in four women will experience domestic violence in her lifetime, the United Nations’ special rapporteur on violence against women and girls, Reem Alsalem, said earlier this year.
Charities and human rights organizations have subsequently reiterated urgent demands to tackle femicide in the UK.
Воины-пайщики оказались без защиты в тылу
Пайщики «Бест Вей» — участники СВО готовы защищать свой кооператив
[url=https://svpressa.ru/society/article/421904]Пайщики кооператива Бест Вей[/url]
Многие пайщики кооператива «Бест Вей» и те, в чьих интересах приобретаются квартиры в кооперативе, — участники СВО. И они сами, и их родственники возмущены событиями, происходящими вокруг кооператива. Ведь защищая интересы страны на фронте, в тылу они не защищены от того, что не могут приобрести недвижимость из-за того, что счета кооператива с почти 4 млрд рублей уже более двух лет находятся под арестом — притом, что сумма ущерба по уголовному делу, рассматриваемому Приморским районным судом Санкт-Петербурга, согласно обвинительному заключению, составляет 282 млн рублей.
«Надеемся, новое руководство Министерства обороны поможет урегулировать ситуацию»
Наталья Пригаро, мать пайщика кооператива — участника СВО из Нефтеюганска Даниила Пригаро:
— Сын расплатился за однокомнатную квартиру, приобретенную с помощью кооператива, еще два года назад, подал пакет документов на оформление квартиры в собственность в Росреестр в начале сентября 2022 года — и все это время не может зарегистрировать собственность на нее из-за того, что в Росреестре она значится как арестованная. В конце сентября 2022 года он был мобилизован. Сын брал кредит, чтобы расплатиться за квартиру с кооперативом. Планировал продать эту квартиру и взять квартиру побольше в ипотеку. Однако неоднократно накладывался арест на недвижимость кооператива, несмотря на то что сейчас ареста нет — последний арест, накладывавшийся осенью 2023 года, 19 февраля истек, ходатайства о новом аресте дважды отклонены судом, по Росреестру арест с квартиры до сих пор не снят. В результате кредит приходится платить, квартира в собственность до сих пор не оформлена.
У сына нет никакой физической и финансовой возможности выяснять отношения с Росреестром — в административном или судебном порядке, он приезжает в отпуска на неделю-полторы, ему не до сутяжничества. Я нахожусь в Анапе, приобрела квартиру с помощью кооператива — сын не имеет возможности сделать на меня генеральную доверенность, а я не могу из Анапы приехать в Нефтеюганск и решать вопросы в Росреестре — кроме поезда, сейчас, сами понимаете, ничего не ходит (конец цитаты).
«На безобразие, которое происходит с кооперативом, мы пишем жалобы во все инстанции, — говорит она. — Надеемся, новое руководство Министерства обороны поможет урегулировать ситуацию».
«Надежда, что все разрешится, остается — не может же все быть коррумпировано»
Елена Бойко — пайщик кооператива из Новосибирска и мать солдата, воющего в СВО, для которого она собиралась приобрести одну из двух квартир в кооперативе. «Я стала пайщиком кооператива в 2021 году. В январе 2021 года подала заявление на покупку двух квартир с помощью кооператива по накопительной программе. У меня двое детей — сын и дочь, им нужно жилье. В 2020 году взяли в ипотеку загородный дом — причем оформили на сына как единственного, кому ее могли одобрить, а в 2021 году он ушел в армию как призывник, а потом подписал контракт. Его относительно недавно отправили „за ленточку“. Думали, что поживем в ипотечном доме, и тут как раз подойдет очередь приобретать квартиры с помощью кооператива. Мы останемся жить в частном доме, а дети въедут в кооперативные квартиры».
«Ипотека — это разорительно, — подчеркивает Елена, — мы взяли 2,5 млн, а за 30 лет отдадим 6 млн. Когда появился кооператив, для нас это была возможность предоставить каждому из детей жилье: за 10 лет или раньше они расплатятся с кооперативом, причем с минимальной по сравнению ипотекой переплатой, и у них будет собственное жилье. Из-за репрессий в отношении кооператива мы можем быть этой возможности лишены».
«Я всем сердцем переживаю за ситуацию с кооперативом, заявление о выходе из кооператива пока не пишу — надеюсь, что его деятельность „разморозится“, — говорит Елена. — Обращаемся во все инстанции, в прокуратуру — приходят отписки. Смысл происходящего я понимаю хорошо — когда-то занималась кредитным потребительским кооперативом: нас тогда закрыл Центральный банк. Нашел формальный повод. Но настоящая причина была в том, что мы мешали банкам обирать клиентов. Но надежда на то, что все разрешится, остается — не может же быть все коррумпировано».
«Больше всего в нашей ситуации возмущает то, что наши воины стоят на стороне государства на фронте, а здесь, в тылу, их интересы нарушаются государственными структурами. Такого быть не должно!»
«Для многих кооператив — единственная возможность приобрести квартиру»
Лариса Зискинд — пайщица «Бест Вей», переехавшая с помощью кооператива с семьей — с дочерью, зятем и маленьким внуком - с Дальнего Востока в Белгородскую область. Ее зять, живущий в кооперативной квартире, находится на фронте СВО. «С помощью кооператива я приобрела квартиру в двухэтажном доме в селе Пушкарное Белгородской области — мы с дочерью переехали сюда. Мы уже два года расплачиваемся за него с кооперативом. Не до конца сделали ремонт, кухни фактически нет: моем посуду в ванной. Финансовых средств не хватает, так как зять уже год на СВО, пока ни разу не был в отпуске, и перевести деньги с его карты в ВТБ сейчас непросто, потому что карта — в расположении, до ближайшего банка ехать далеко, а он сам — на линии фронта. Мы фактически живем с выплат на ребенка, на мне кредиты — у меня полпенсии уходит на их оплату. Квартира по Росреестру находится под арестом, хотя арест по квартирам кооператива арест истек — а я боюсь ехать в Белгород, чтобы решать по ней вопрос. Я с ребенком на улицу гулять не выхожу в нынешней военной обстановке — сирены воздушной тревоги постоянные. Побыстрее бы наши войска создали санитарную зону в Харьковской области!»
По поводу покупки квартиры Лариса сообщает, что «ипотека точно никак не светила. Кооператив — это 14 тыс. в месяц и всего лишь 10 лет. В кооперативе даже студентка может приобрести квартиру, почти не переплачивая. При покупке через банк придется заплатить еще как минимум за одну квартиру».
«Среди пайщиков кооператива очень много пенсионеров, социально незащищенных людей, это была их единственная возможность приобрести квартиру, — говорит Лариса. — Из-за репрессий против кооператива мы можем лишиться единственного жилья. Мы куда только не писали в защиту кооператива. Но, видимо, слишком большой куш. Кооперативу не дают даже платить налоги. Надеемся, что отношение к кооперативу в результате справедливого разбирательства изменится в лучшую сторону».
«Мы очень устали. Но боремся»
Людмила Гарина — пайщица кооператива из Саратова, зять которой, проживающий вместе с ней в кооперативной квартире, участвует в СВО. «Я проживаю в кооперативной квартире почти три года вместе с дочерью и внуком, вносим ежемесячные платежи через госуслуги».
«Возможности взять ипотеку не было, — говорит Людмила. — Фактически у нас один работающий человек — дочь, зять мобилизован, внук — студент, внучке — четыре года. Мы с мужем пенсионеры».
«У нас с кооперативом проблем не было никаких, — говорит она, — но сейчас уверенности в завтрашнем дне нет. Пишем во все возможные инстанции, в том числе в Администрацию президента, ходили на прием к Володину. Пока ничего не помогает. Вся надежда на то, что суд разберется».
«Хотелось, чтобы вопрос о возобновлении работы кооператива быстрее решился, — говорит она, — тогда стало бы проще. Когда кооператив заработает, будет совершенно иная ситуация. Мы очень устали. Но боремся».
«20 тыс. пайщиков кооператива не так просто остановить»
Алла Яровая — пайщик кооператива из Пятигорска и представитель пайщицы Людмилы Дворцовой, сын которой, Игорь Дворцов, находился на СВО, а теперь служит внутри страны. «Людмила приобрела с помощью кооператива в Пятигорске квартиру для своего сына. Вступила в кооператив она в 2017 году, сначала находилась в накопительной программе — накапливала на первоначальный паевый взнос, накопила, потом стояла в очереди на приобретение жилье, приобрела с помощью кооператива однокомнатную квартиру за 1450 тыс. рублей. Потом приобрела квартиру и постепенно полностью за нее расплатилась. Работала для этого на двух работах, в том числе санитаркой в инфекционном отделении».
Но оформить квартиру в собственность не удалось, так как она оказалась под арестом, поясняет Алла. И хотя арест 19 февраля истек и больше не накладывался, суд первой инстанции в Пятигорске этот арест подтвердил. «По вопросу получения права собственности и снятия ареста мы обратились в Пятигорский городской суд, где судья очень предвзято отнесся к решению вопроса и мы получили отказ, — говорит Алла, — Хотя аналогичный вопрос рассматривался в Минераловодском суде и там судья вынесла положительное решение и в настоящий момент пайщик получил право собственности на свою квартиру. Предстоит рассмотрение в апелляционной инстанции — несмотря на то, что рассматривать нечего: ареста на недвижимость кооператива сейчас нет. А на квартиру Людмилы Дворцовой — почему-то есть».
«Кооператив — единственная возможность для людей со средним достатком получить жилье, — говорит Алла, — Нерешенный вопрос — налоги. Мы вынуждены платить налоги с юридических лиц, но мы же физические лица. К депутатам — с нас берут налог как с юрлица. Почему мы должны оплачивать — мы же физлица? Во многих регионах это сделано. Надеемся добиться того, чтобы это было сделано по всей стране».
«20 тыс. пайщиков кооператива не так просто остановить, — подчеркивает пайщица. — Это наши деньги, и мы за них боремся. Обязательно отстоим наш кооператив!»
«Активно участвую в защите кооператива»
У пайщицы кооператива, жительницы из Ленинградской области Татьяны Власенко старший сын находится на СВО с сентября 2022 года. «Три месяца проходил обучение, а потом оказался „за ленточкой“, — говорит Татьяна. — Мы должны были в конце лета 2022 года купить квартиру с помощью кооператива — квартиру планировали большую. Но работа кооператива оказалась заблокирована, и кооператив не может ничего купить уже более двух лет, так как и его счета, и его деятельность заблокированы. Деньги пайщиков, собранные для приобретения квартир, при этом обесцениваются».
«Я активно участвую в защите нашего кооператива, — говорит Татьяна. — Езжу на все суды в Санкт-Петербурге, хотя живу в деревне — ехать два с половиной, а то и три часа, мне 64 года, а путь неблизкий. И сын в этом поддерживает меня».
«Вот уже более двух лет работа кооператива заблокирована, деньги обесцениваются»
У пайщицы из Хабаровска Натальи Ромашко на СВО было два племянника — один, к сожалению, погиб. Планировалась покупка большой квартиры в Хабаровске, Наталья состояла в накопительной программе. «Вот уже более двух лет работа кооператива заблокирована, деньги обесцениваются, — говорит Наталья. — Я участвую в защите кооператива, пишу письма в прокуратуру, в высшие государственные инстанции, потому что нарушаются наши права, нас лишают возможности приобрести квартиру вне ипотечной системы. Пока приходят одни отписки, но борьбу мы продолжаем!»
«Ситуация — как на СВО»
Пайщица Светлана Иванова из Новосибирска, волонтер СВО, планировала приобрести с помощью кооператива две квартиры — для себя и для сына, который воюет на СВО добровольцем. «До этого я продала квартиру и сейчас вообще без собственного жилья, — говорит она. — Вступила в кооператив по накопительной программе, рассматривали для приобретения квартиры Крым и Новосибирск. В СВО участвую как волонтер — не остаюсь в стороне, когда Родине нужна помощь».
«От ситуации с кооперативом очень многие люди пострадали, — говорит Светлана. — Ситуация — как на СВО. И еще больше пострадают, если реализуется плохой сценарий. Хочется надеяться на то, что все образуется, правда восторжествует. Заявление о выходе из кооператива и возврате паевых средств я не подавала. Готова продолжить участие в нем, готова приобрести недвижимость с его помощью: это самый лучший вариант покупки квартиры».
«Сдаваться в планах нет»
Пайщица Марина Янковская родом из Хабаровска, она переехала в Новороссийск, ее супруг погиб на СВО. «У нас уже подходила очередь на покупку квартиры в Новороссийске для нашей семьи, мы должны были подбирать объект, — рассказывает она. — Но два года назад все остановилось. Хотели приобрести двухкомнатную квартиру в Новороссийске. Деньги лежат на арестованных счетах кооператива — а недвижимость за это время подорожала. Надеемся хоть что-то приобрести после того, как счета разблокируют».
«Мы всячески поддерживаем кооператив, — говорит Марина. — Записывали видео, что мы не обманутые, а вполне довольные пайщики, объясняли, что это никакая не пирамида, а организация, созданная в помощь людям. Ждем, когда снимут эти аресты. Сдаваться в планах нет».
Воины-пайщики оказались без защиты в тылу
Пайщики «Бест Вей» — участники СВО готовы защищать свой кооператив
[url=https://svpressa.ru/society/article/421904]Владимир Колокольцев[/url]
Многие пайщики кооператива «Бест Вей» и те, в чьих интересах приобретаются квартиры в кооперативе, — участники СВО. И они сами, и их родственники возмущены событиями, происходящими вокруг кооператива. Ведь защищая интересы страны на фронте, в тылу они не защищены от того, что не могут приобрести недвижимость из-за того, что счета кооператива с почти 4 млрд рублей уже более двух лет находятся под арестом — притом, что сумма ущерба по уголовному делу, рассматриваемому Приморским районным судом Санкт-Петербурга, согласно обвинительному заключению, составляет 282 млн рублей.
«Надеемся, новое руководство Министерства обороны поможет урегулировать ситуацию»
Наталья Пригаро, мать пайщика кооператива — участника СВО из Нефтеюганска Даниила Пригаро:
— Сын расплатился за однокомнатную квартиру, приобретенную с помощью кооператива, еще два года назад, подал пакет документов на оформление квартиры в собственность в Росреестр в начале сентября 2022 года — и все это время не может зарегистрировать собственность на нее из-за того, что в Росреестре она значится как арестованная. В конце сентября 2022 года он был мобилизован. Сын брал кредит, чтобы расплатиться за квартиру с кооперативом. Планировал продать эту квартиру и взять квартиру побольше в ипотеку. Однако неоднократно накладывался арест на недвижимость кооператива, несмотря на то что сейчас ареста нет — последний арест, накладывавшийся осенью 2023 года, 19 февраля истек, ходатайства о новом аресте дважды отклонены судом, по Росреестру арест с квартиры до сих пор не снят. В результате кредит приходится платить, квартира в собственность до сих пор не оформлена.
У сына нет никакой физической и финансовой возможности выяснять отношения с Росреестром — в административном или судебном порядке, он приезжает в отпуска на неделю-полторы, ему не до сутяжничества. Я нахожусь в Анапе, приобрела квартиру с помощью кооператива — сын не имеет возможности сделать на меня генеральную доверенность, а я не могу из Анапы приехать в Нефтеюганск и решать вопросы в Росреестре — кроме поезда, сейчас, сами понимаете, ничего не ходит (конец цитаты).
«На безобразие, которое происходит с кооперативом, мы пишем жалобы во все инстанции, — говорит она. — Надеемся, новое руководство Министерства обороны поможет урегулировать ситуацию».
«Надежда, что все разрешится, остается — не может же все быть коррумпировано»
Елена Бойко — пайщик кооператива из Новосибирска и мать солдата, воющего в СВО, для которого она собиралась приобрести одну из двух квартир в кооперативе. «Я стала пайщиком кооператива в 2021 году. В январе 2021 года подала заявление на покупку двух квартир с помощью кооператива по накопительной программе. У меня двое детей — сын и дочь, им нужно жилье. В 2020 году взяли в ипотеку загородный дом — причем оформили на сына как единственного, кому ее могли одобрить, а в 2021 году он ушел в армию как призывник, а потом подписал контракт. Его относительно недавно отправили „за ленточку“. Думали, что поживем в ипотечном доме, и тут как раз подойдет очередь приобретать квартиры с помощью кооператива. Мы останемся жить в частном доме, а дети въедут в кооперативные квартиры».
«Ипотека — это разорительно, — подчеркивает Елена, — мы взяли 2,5 млн, а за 30 лет отдадим 6 млн. Когда появился кооператив, для нас это была возможность предоставить каждому из детей жилье: за 10 лет или раньше они расплатятся с кооперативом, причем с минимальной по сравнению ипотекой переплатой, и у них будет собственное жилье. Из-за репрессий в отношении кооператива мы можем быть этой возможности лишены».
«Я всем сердцем переживаю за ситуацию с кооперативом, заявление о выходе из кооператива пока не пишу — надеюсь, что его деятельность „разморозится“, — говорит Елена. — Обращаемся во все инстанции, в прокуратуру — приходят отписки. Смысл происходящего я понимаю хорошо — когда-то занималась кредитным потребительским кооперативом: нас тогда закрыл Центральный банк. Нашел формальный повод. Но настоящая причина была в том, что мы мешали банкам обирать клиентов. Но надежда на то, что все разрешится, остается — не может же быть все коррумпировано».
«Больше всего в нашей ситуации возмущает то, что наши воины стоят на стороне государства на фронте, а здесь, в тылу, их интересы нарушаются государственными структурами. Такого быть не должно!»
«Для многих кооператив — единственная возможность приобрести квартиру»
Лариса Зискинд — пайщица «Бест Вей», переехавшая с помощью кооператива с семьей — с дочерью, зятем и маленьким внуком - с Дальнего Востока в Белгородскую область. Ее зять, живущий в кооперативной квартире, находится на фронте СВО. «С помощью кооператива я приобрела квартиру в двухэтажном доме в селе Пушкарное Белгородской области — мы с дочерью переехали сюда. Мы уже два года расплачиваемся за него с кооперативом. Не до конца сделали ремонт, кухни фактически нет: моем посуду в ванной. Финансовых средств не хватает, так как зять уже год на СВО, пока ни разу не был в отпуске, и перевести деньги с его карты в ВТБ сейчас непросто, потому что карта — в расположении, до ближайшего банка ехать далеко, а он сам — на линии фронта. Мы фактически живем с выплат на ребенка, на мне кредиты — у меня полпенсии уходит на их оплату. Квартира по Росреестру находится под арестом, хотя арест по квартирам кооператива арест истек — а я боюсь ехать в Белгород, чтобы решать по ней вопрос. Я с ребенком на улицу гулять не выхожу в нынешней военной обстановке — сирены воздушной тревоги постоянные. Побыстрее бы наши войска создали санитарную зону в Харьковской области!»
По поводу покупки квартиры Лариса сообщает, что «ипотека точно никак не светила. Кооператив — это 14 тыс. в месяц и всего лишь 10 лет. В кооперативе даже студентка может приобрести квартиру, почти не переплачивая. При покупке через банк придется заплатить еще как минимум за одну квартиру».
«Среди пайщиков кооператива очень много пенсионеров, социально незащищенных людей, это была их единственная возможность приобрести квартиру, — говорит Лариса. — Из-за репрессий против кооператива мы можем лишиться единственного жилья. Мы куда только не писали в защиту кооператива. Но, видимо, слишком большой куш. Кооперативу не дают даже платить налоги. Надеемся, что отношение к кооперативу в результате справедливого разбирательства изменится в лучшую сторону».
«Мы очень устали. Но боремся»
Людмила Гарина — пайщица кооператива из Саратова, зять которой, проживающий вместе с ней в кооперативной квартире, участвует в СВО. «Я проживаю в кооперативной квартире почти три года вместе с дочерью и внуком, вносим ежемесячные платежи через госуслуги».
«Возможности взять ипотеку не было, — говорит Людмила. — Фактически у нас один работающий человек — дочь, зять мобилизован, внук — студент, внучке — четыре года. Мы с мужем пенсионеры».
«У нас с кооперативом проблем не было никаких, — говорит она, — но сейчас уверенности в завтрашнем дне нет. Пишем во все возможные инстанции, в том числе в Администрацию президента, ходили на прием к Володину. Пока ничего не помогает. Вся надежда на то, что суд разберется».
«Хотелось, чтобы вопрос о возобновлении работы кооператива быстрее решился, — говорит она, — тогда стало бы проще. Когда кооператив заработает, будет совершенно иная ситуация. Мы очень устали. Но боремся».
«20 тыс. пайщиков кооператива не так просто остановить»
Алла Яровая — пайщик кооператива из Пятигорска и представитель пайщицы Людмилы Дворцовой, сын которой, Игорь Дворцов, находился на СВО, а теперь служит внутри страны. «Людмила приобрела с помощью кооператива в Пятигорске квартиру для своего сына. Вступила в кооператив она в 2017 году, сначала находилась в накопительной программе — накапливала на первоначальный паевый взнос, накопила, потом стояла в очереди на приобретение жилье, приобрела с помощью кооператива однокомнатную квартиру за 1450 тыс. рублей. Потом приобрела квартиру и постепенно полностью за нее расплатилась. Работала для этого на двух работах, в том числе санитаркой в инфекционном отделении».
Но оформить квартиру в собственность не удалось, так как она оказалась под арестом, поясняет Алла. И хотя арест 19 февраля истек и больше не накладывался, суд первой инстанции в Пятигорске этот арест подтвердил. «По вопросу получения права собственности и снятия ареста мы обратились в Пятигорский городской суд, где судья очень предвзято отнесся к решению вопроса и мы получили отказ, — говорит Алла, — Хотя аналогичный вопрос рассматривался в Минераловодском суде и там судья вынесла положительное решение и в настоящий момент пайщик получил право собственности на свою квартиру. Предстоит рассмотрение в апелляционной инстанции — несмотря на то, что рассматривать нечего: ареста на недвижимость кооператива сейчас нет. А на квартиру Людмилы Дворцовой — почему-то есть».
«Кооператив — единственная возможность для людей со средним достатком получить жилье, — говорит Алла, — Нерешенный вопрос — налоги. Мы вынуждены платить налоги с юридических лиц, но мы же физические лица. К депутатам — с нас берут налог как с юрлица. Почему мы должны оплачивать — мы же физлица? Во многих регионах это сделано. Надеемся добиться того, чтобы это было сделано по всей стране».
«20 тыс. пайщиков кооператива не так просто остановить, — подчеркивает пайщица. — Это наши деньги, и мы за них боремся. Обязательно отстоим наш кооператив!»
«Активно участвую в защите кооператива»
У пайщицы кооператива, жительницы из Ленинградской области Татьяны Власенко старший сын находится на СВО с сентября 2022 года. «Три месяца проходил обучение, а потом оказался „за ленточкой“, — говорит Татьяна. — Мы должны были в конце лета 2022 года купить квартиру с помощью кооператива — квартиру планировали большую. Но работа кооператива оказалась заблокирована, и кооператив не может ничего купить уже более двух лет, так как и его счета, и его деятельность заблокированы. Деньги пайщиков, собранные для приобретения квартир, при этом обесцениваются».
«Я активно участвую в защите нашего кооператива, — говорит Татьяна. — Езжу на все суды в Санкт-Петербурге, хотя живу в деревне — ехать два с половиной, а то и три часа, мне 64 года, а путь неблизкий. И сын в этом поддерживает меня».
«Вот уже более двух лет работа кооператива заблокирована, деньги обесцениваются»
У пайщицы из Хабаровска Натальи Ромашко на СВО было два племянника — один, к сожалению, погиб. Планировалась покупка большой квартиры в Хабаровске, Наталья состояла в накопительной программе. «Вот уже более двух лет работа кооператива заблокирована, деньги обесцениваются, — говорит Наталья. — Я участвую в защите кооператива, пишу письма в прокуратуру, в высшие государственные инстанции, потому что нарушаются наши права, нас лишают возможности приобрести квартиру вне ипотечной системы. Пока приходят одни отписки, но борьбу мы продолжаем!»
«Ситуация — как на СВО»
Пайщица Светлана Иванова из Новосибирска, волонтер СВО, планировала приобрести с помощью кооператива две квартиры — для себя и для сына, который воюет на СВО добровольцем. «До этого я продала квартиру и сейчас вообще без собственного жилья, — говорит она. — Вступила в кооператив по накопительной программе, рассматривали для приобретения квартиры Крым и Новосибирск. В СВО участвую как волонтер — не остаюсь в стороне, когда Родине нужна помощь».
«От ситуации с кооперативом очень многие люди пострадали, — говорит Светлана. — Ситуация — как на СВО. И еще больше пострадают, если реализуется плохой сценарий. Хочется надеяться на то, что все образуется, правда восторжествует. Заявление о выходе из кооператива и возврате паевых средств я не подавала. Готова продолжить участие в нем, готова приобрести недвижимость с его помощью: это самый лучший вариант покупки квартиры».
«Сдаваться в планах нет»
Пайщица Марина Янковская родом из Хабаровска, она переехала в Новороссийск, ее супруг погиб на СВО. «У нас уже подходила очередь на покупку квартиры в Новороссийске для нашей семьи, мы должны были подбирать объект, — рассказывает она. — Но два года назад все остановилось. Хотели приобрести двухкомнатную квартиру в Новороссийске. Деньги лежат на арестованных счетах кооператива — а недвижимость за это время подорожала. Надеемся хоть что-то приобрести после того, как счета разблокируют».
«Мы всячески поддерживаем кооператив, — говорит Марина. — Записывали видео, что мы не обманутые, а вполне довольные пайщики, объясняли, что это никакая не пирамида, а организация, созданная в помощь людям. Ждем, когда снимут эти аресты. Сдаваться в планах нет».
Simulation games like Flight Simulator offer incredibly realistic experiences, allowing players to engage in activities they might not otherwise have the chance to. https://kpkesihatan.com/
Every weekend i used to visit this website, because i
wish for enjoyment, for the reason that this this site conations really pleasant funny information too.
Hi there! Do you use Twitter? I’d like to follow you if that would be ok.
I’m absolutely enjoying your blog and look forward to new posts.
Thank you a lot for sharing this with all people you actually understand what you are speaking approximately! Bookmarked. Please additionally talk over with my website =). We can have a hyperlink alternate contract among us
http://gazeta.sebastopol.ua/vazhlyvist-yakisnogo-skla-fary-chomu-tse-maye-znachennya
WELCOME TO PUGACHEV LUXURY CAR RENTALS EXCLUSIVE MOTORING LUXURY & EXOTIC RENTALS
Dive into the ultimate driving experience with Pugachev Luxury Car Rentals, your premier destination for luxury and exotic car rentals in Los Angeles. Whether you’re looking to impress at a corporate event, add a splash of glamour to your weekend, or simply enjoy the thrill of driving some of the world’s most sought-after vehicles, we’ve got you covered.
https://pugachev.la/
Our extensive fleet includes the pinnacle of automotive excellence from renowned brands such as Lamborghini, Rolls-Royce, Bentley, Ferrari, McLaren, and more. Indulge in the elegance of a Rolls-Royce Ghost, starting at $1,150 per day, or experience the exhilarating performance of a Lamborghini Huracan EVO, available from $875 per day. With options ranging from the powerful Ferrari GTB Red at $1,100 per day to the opulent Bentley Continental GT at $950 per day, our collection is meticulously curated to cater to the most discerning tastes and preferences.
Navigating the vibrant streets of Los Angeles or cruising along the scenic coastlines has never been more accessible or more stylish. At Pugachev Luxury Car Rentals, we make it easy and efficient for you to rent the luxury car of your dreams with straightforward daily and weekly rental options—like the versatile Mercedes-Benz AMG G63, which can be yours for $790 daily or $4,424 weekly.
Embrace the luxury, power, and finesse of our elite vehicles and elevate your next Los Angeles visit into a truly memorable journey. Whether you’re seeking an exotic sports car for that extra special thrill, a luxury sedan for sophisticated travel, or a sumptuous SUV for family comfort and safety, our fleet promises the perfect match for every occasion and desire.
OUR COLLECTION OF LUXURY CARS FOR DISCERNING DRIVERS
Elegant Sedans for Business Executives
At Pugachev Luxury Car Rentals, we understand the importance of a strong first impression. Our fleet of luxury sedans is designed to cater to business executives who demand sophistication and functionality. Each vehicle, such as the Rolls-Royce Ghost Anthracite, available at $1,100 per day, offers a sublime mix of comfort and prestige, ensuring you arrive at your business engagements in unparalleled style. For a slightly different but equally impressive experience, consider the Bentley Flying Spur at $1,000 daily, which combines classic elegance with cutting-edge technology.
luxury car rental los angeles
luxury car rental los angeles
Sumptuous SUVs for Family and Leisure
Our selection of high-end SUVs blends luxury with practicality, making them perfect for family trips or stylish outings with friends. The Bentley Bentayga, starting from $700 per day, offers spaciousness and state-of-the-art amenities that ensure comfort without compromise. For those seeking a bolder statement, the Rolls-Royce Cullinan Black Badge provides an imposing presence and unrivaled luxury for $1,300 daily, ensuring every journey is an event in itself.
Prestige Convertibles for Scenic Drives
Los Angeles is renowned for its beautiful landscapes and stunning coastlines, best enjoyed in one of our prestige convertibles. Feel the exhilarating freedom of the open road in a Ferrari Portofino Spyder, available for $850 daily. Its breathtaking performance and sleek design are perfect for scenic drives along the Pacific Coast Highway. Alternatively, the BMW M4 Competition Cabrio, at $390 per day, offers a more accessible but equally thrilling option for cruising under the Californian sun.
Our fleet at Pugachev Luxury Car Rentals is meticulously maintained and regularly updated to ensure that each car looks and performs like new. This commitment to quality and excellence reflects our dedication to providing only the best for our clients, who return time and again for the reliability and prestige our services offer. Each model in our fleet is more than just a vehicle; it’s a part of an exclusive lifestyle made accessible to our discerning clientele. Whether you are looking to impress at a corporate event or simply indulge in the pleasure of driving a luxury car, our collection is guaranteed to enhance your experience in Los Angeles.
exotic car rental los angeles
luxury and exotic car rental in los angeles
A lot of blog writers nowadays yet just a few have blog posts worth spending time on reviewing.
My website: порно римминг
I believe this is one of the so much vital information for me.
And i’m happy reading your article. But wanna commentary on some common issues, The site style is perfect, the
articles is in point of fact nice : D. Good job, cheers
It’s nearly impossible to find well-informed people about this
subject, but you sound like you know what you’re talking about!
Thanks
Hello there I am so thrilled I found your website, I really
found you by error, while I was researching on Yahoo
for something else, Nonetheless I am here now and would just like to say kudos for a marvelous post and a all
round interesting blog (I also love the theme/design),
I don’t have time to read through it all at the minute but I have bookmarked
it and also added in your RSS feeds, so when I have time I will be back to read a great deal more,
Please do keep up the awesome work.
Hi there! Quick question that’s entirely off topic.
Do you know how to make your site mobile friendly?
My site looks weird when viewing from my iphone 4.
I’m trying to find a template or plugin that might be able to fix this problem.
If you have any recommendations, please share. Thank you!
Pretty! This was an incredibly wonderful article. Thanks for supplying this info.
Hi there friends, its impressive article on the topic of educationand
completely defined, keep it up all the time.
I love reading an article that can make men and women think. Also, many thanks for allowing me to comment.
I always spent my half an hour to read this website’s posts daily along with a cup
of coffee.
A fairly new player in the Russian darknet arena, blacksprut Blacksprut has quickly gained attention for its interesting features and growing popularity. While some aspects raise questions, it has the potential to become a prominent figure in the darknet scene.
Features:
Blacksprut https://bs-gl-darknet.com offers an “Instant Transactions” feature, inspired by the success of similar systems on other platforms like Hydra. Couriers hide goods within the city and provide buyers with coordinates, adding an adventurous element to the purchasing process.
Its not my first time to pay a quick visit this website, i
am visiting this web page dailly and get good information from here all
the time.
I could not resist commenting. Very well written!
If you are going for best contents like myself, only pay a visit this site everyday for
the reason that it offers quality contents, thanks
Военный адвокат Запорожье. Бесплатная консультация
юрист Днепр
— это опытный специалист имеющий высшее юридическое образование, сдавший квалификационный государственный экзамен на право осуществления адвокатской деятельностью и специализирующийся в основном на военных делах
Вся правовая помощь военного адвоката осуществляется в индивидуальном порядке, грамотно, четко и в соответствии с действующими нормативно-правовыми актами.
Мы как военные юристы действуем не против органов Украины или министерства обороны, мы действуем во благо Украины — наших защитников и граждан Украины, которые попали в тяжелую жизненную ситуацию связанную с незнанием военного и действующего законодательства.
Поскольку, проявив патриотизм и чувство гражданской ответственности – став на защиту суверенитета страны, граждане участвующие и помогавшие в обороне после, становятся никому не нужными, особенно если военнослужащий стал инвалидом, потерял часть тела или конечность, и не может самостоятельно защитить свои права. Именно в таких ситуациях мы как военные адвокаты приходим на помощь, и добиваемся в установленном законом порядке справедливости, необходимых выплат, установление статуса, оформление пенсий, льгот и т.п.
Тоже касается, и получение отсрочки от мобилизации, когда например, безосновательно призывают сына у которого отец инвалид 2 группы, или мать прикованная из-за тяжелой болезни к постели, и требующая постороннего ухода. Это же относится и к военнослужащим, рапорта которых не регистрируются в канцелярии воинской части и полностью игнорируются, под прикрытием суеты боевых действий..
Именно в таких ситуациях, мы приходим на помощь и с помощью ЗАКОННЫХ методов правовой защиты, используя свой опыт полученный при ведении аналогичных военных дел добиваемся справедливости.
Appreciation to my father who stated to me about
this webpage, this web site is actually awesome.
Hi there! I could have sworn I’ve been to your blog before but after looking at a few of the posts I realized it’s new to me. Anyways, I’m definitely delighted I discovered it and I’ll be book-marking it and checking back regularly!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.nfomedia.com/profile?uid=rOiZcbE
https://rentry.org/tdrzb4m5
http://www.nfomedia.com/profile?uid=rOiYekB
https://www.hentai-foundry.com/user/mormitor1963/profile
https://fgfhgfhgf1964.bandcamp.com/album/submission-by-pleasure
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/350992
http://www.babelcube.com/user/sarah-williams-3
https://tubeteencam.com/user/ereds1952/profile
http://www.babelcube.com/user/kim-richardson
https://cannabis.net/user/151282
Разбираю ситуацию вокруг “кооператива” Life is Good
гей порно
Я решил написать этот пост, чтоб пройтись по доводам тех, кто привлекает новых “пайщиков” в структуры г-на Василенко (на деле – оплачивающих существование пирамиды Life is Good). Понятно, что, в этом сообществе крутятся сотни тех, кто занимается привлечением и теперь ЦБ с прокуратурой их оставят не только без заработка, но, возможно, придется отвечать за свои действия в суде. А деньги уже, наверняка, потрачены. Также есть люди, которые уже живут в квартире от LIG или выплатили некую сумму, они верят до конца и отказываются принимать ситуацию, в которой надо признать, что они могут остаться и без денег, и без квартиры. В любом случае, даже если кто-то живет в квартире от LIG, она ему не принадлежит и, в отличие от банковского залога, урегулировать вопрос не получится (их просто выселят через суд при банкротстве LIG, хотя ситуация скандальная и возможно государство что-то предпримет, чтоб сгладить проблему).Теперь по сути, что такое LIG.
Компания LIG – «Лайф-из-гуд» заявляет два основных направления деятельности: инвестиционные счета «Виста» в компании Hermes Management Ltd. и жилищный кооператив «Бест-вей» — альтернатива ипотеке.
Если верить консультантам «Лайф-из-гуд», доходность счетов «Виста» в компании «Гермес-менеджмент» последние годы держится в районе 20% годовых в долларах. Это довольно много (Уоррен Баффет позавидовал бы такой доходности), при этом непонятно, за счет чего достигается такая доходность.
Деятельность «Гермес-менеджмента» непрозрачна, на сайте не видно полноценных отчетов, нет сведений о структуре компании и ее руководстве.
«Гермес-менеджмент» привлекает российских клиентов через компанию «Лайф-из-гуд», но у него нет лицензии Центрального банка России на брокерскую деятельность или доверительное управление.
«Гермес-менеджмент» зарегистрирован в Белизе, офшорной зоне в Центральной Америке. Там же расположен офис компании. Еще в документах упоминается компания Hermes M в Шотландии. В случае проблем придется разбираться с зарубежной компанией, что делает задачу бесперспективной. Более того, учитывая реалии мошеннических схем, бывшие пайщики столкнутся еще и с лже-юристами, обещающими вернуть им вложенные деньги и конечно, останутся в крайне бедственном положении, с кредитами, без жилья и перспектив (у кого-то могут опуститься руки со всеми вытекающими).
Компания платит за привлечение новых клиентов, открывающих инвестиционные счета «Виста» (по сути – ничем не обеспеченные цифры на сайте. Заведение средств происходит через перевод средств на счета частных лиц(!), выполняющих роль консультантов LIG (а это сразу нарушает законы РФ и привлекает внимание регулятора), вывод – через перепродажу воздуха других буратинам. После внесения LIG в черный список, чаты структуры полны объявлений о продажи токенов Виста, но покупателей почти нет)) и вступающих в жилищный кооператив «Бест-вей». Система комиссионных выглядит такой разветвленной и продуманной, что становится понятно, что именно в ней суть компании, а не в инвестициях или альтернативе ипотеке.
Сам по себе сетевой маркетинг — вполне законный способ привлечения новых клиентов. Но когда он выглядит как основной вид деятельности, а клиентам при этом рассказывают про инвестиции, пассивный доход и успех в жизни, это наводит на очевидные выводы.
Суть предложения «Гермеса» такая: инвесторы открывают счет и перечисляют компании доллары или евро, а взамен получают пассивный доход. Минимальная сумма инвестиционного контракта — 10 тысяч долларов или евро. При этом первый взнос может быть 1130 долларов или 1100 евро, а оставшуюся сумму можно довнести позже.
Чтобы открыть счет «Виста», надо заполнить регистрационную форму на сайте «Лайф-из-гуд». При регистрации сайт просит поставить галочку «Я ознакомился и принимаю условия указанные в публичной оферте» (пунктуация сохранена).
По ссылке открывается не оферта, а договор об оказании услуг. В договоре не указаны название и реквизиты компании, с которой заключается договор. Кроме того, речь в нем идет не об управлении активами, а только об «образовательных и консультационных услугах об инвестиционных механизмах и способах управления капиталом». Это не похоже на обещанный пассивный доход от слова совсем. Таким образом, зазывалы в пирамиду пытаются уйти от ответственности.
домашний анальный секс
Чтобы открыть счет и пользоваться им, придется заплатить различные комиссии. Все они прописаны в договоре:
Комиссия за открытие счета: 100 евро или 130 долларов.
Ажио, или входная комиссия: 7% от суммы лицевого счета. На сайте есть пример расчета: при вложении 100 000 $ ажио составит 7000 $.
Комиссия за управление счетом: 1% в год от размера счета, взимается каждый месяц по 1/12.
Комиссия с прибыли: 20% от ежемесячной прибыли, «полученной от инвестиционных продуктов сторонних партнерских компаний или физических лиц, в результате рекомендаций и деятельности Исполнителя».
Итак, LIG состоит из двух основных компаний: Гермес, которая предлагает участникам инвестиционный доход (с идеей полученную прибыль направить на квартиру) и Бест-Вэй, являющаяся ЖНК. Что это такое?
Это Жилищный накопительный кооператив (общепринятое сокращение — ЖНК) — вид потребительского кооператива, сочетающий в себе качества финансового и жилищно-строительного кооператива. ЖНК работают в соответствии с № 215-ФЗ «О жилищных накопительных кооперативах». Целью деятельности жилищных накопительных кооперативов является удовлетворения потребностей членов кооператива в жилых помещениях путём объединения членами кооператива паевых взносов. Жилищный накопительный кооператив имеет право:
– привлекать и использовать денежные средства граждан на приобретение жилых помещений;
– вкладывать имеющиеся у него денежные средства в строительство жилых помещений (в том числе в многоквартирных домах), а также участвовать в строительстве жилых помещений в качестве застройщика или участника долевого строительства;
– приобретать жилые помещения;
– привлекать заёмные денежные средства, общая величина не должна превышать сорок процентов стоимости имущества кооператива.
Членами жилищного накопительного кооператива могут быть только граждане РФ. Жилищный накопительный кооператив создаётся по инициативе не менее чем пятидесяти человек и не более чем пяти тысяч человек. Контроль за деятельностью кооператива по привлечению и использованию денежных средств граждан на приобретение жилых помещений, а также за соблюдением кооперативом требований действующего законодательства Российской Федерации осуществляет Центральный банк Российской Федерации.
Уже отсюда понятно, что заявления заинтересованных лиц из структур LIG о том, что ЦБ не в праве вмешиваться в деятельность собственно LIG лишены оснований. Может и вмешался.
Если говорить о цифрах, то на 13 декабря 2021 года в LIG было свыше 19 000 пайщиков изо всех уголков РФ. LIG выкупил и передал в безвозмездное пользование пайщикам примерно 2 500 объектов недвижимости, из них по 247 пайщики полностью выплатили пай, оформив недвижимость в собственность. Интересная статистика – 19 000 вступили, заплатив членский взнос в 160 000 рублей, но лишь 8 часть из них накопила необходимые 35% для получения недвижимости при том, что фактически недвижимость их собственностью не является. Заявления о том, что получившие недвижимость прописаны в квартирах – это пережитки советского прошлого и, вероятно, сознательное введение остальных в заблуждение. В современной РФ не существует института прописки, есть регистрация, постоянная и временная. Есть найм жилья и есть собственность. Допускаю, что какая-то часть людей все еще проживает в государственных квартирах и не оформила их в собственность, но это – давно не норма, как это было в СССР (где собственность как раз была редким явлением).
По сути кооператив — посредник между покупателем и продавцом жилья, а это увеличивает риски. Приходится полагаться на то, что кооператив соблюдает закон и что он не закроется до того, как вы оформите квартиру на себя.
Даже если все по закону, риск потерять деньги остается. Если кооператив по каким-то причинам перестанет работать, его собственность распродадут. Сначала он рассчитается с кредиторами и только затем вернет деньги пайщикам — пропорционально их паевым взносам. Хватит ли у кооператива денег, чтобы вернуть все паевые взносы, неизвестно.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/chcoste
https://tubeteencam.com/user/jerevinan1966/profile
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3194432
https://onedio.ru/profile/varchik-199-5
https://rentry.org/wez2z4x3
Hello! I could have sworn I’ve been to this site before but after browsing through some of the posts I realized it’s new to me. Regardless, I’m certainly delighted I came across it and I’ll be book-marking it and checking back often!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/irenegate1982/profile
https://okwave.jp/profile/u3115732.html
https://launchpad.net/~irelend19521
https://rentry.org/4yidbn22
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195275
You need to be a part of a contest for one of the most useful websites online. I most certainly will highly recommend this blog!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/shef681957
https://permacultureglobal.org/users/59007-amanda-greene
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195504
https://chyoa.com/user/fuvas1968
https://anotepad.com/notes/pn9h7ctc
Hey very interesting blog!
We are a bunch of volunteers and opening a brand new scheme in our community.
Your website provided us with useful information to work on. You
have done a formidable task and our whole neighborhood will probably be grateful to you.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://launchpad.net/~sdsw19501
https://anotepad.com/notes/p2x7aeyh
https://chyoa.com/user/ijl1992
https://www.haikudeck.com/presentations/G0ecfypDVh
https://launchpad.net/~testobur19741
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/59310-jenifer-winkler
https://imageevent.com/seell1995
https://cannabis.net/user/151932
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196266
https://www.quia.com/profiles/roberto198b
카 심바 슬롯
정말 큰 변화가 일어나면 나라의 근간이 흔들리고 큰 일이 일어날 것입니다.
Vanderbilt University (informally Vandy or VU) is a private research university in Nashville, Tennessee. Founded in 1873
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.haikudeck.com/presentations/bLzAoFx0xB
https://imageevent.com/leutnant1989
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196848
https://tubeteencam.com/user/jkkkkz1992/profile
https://imageevent.com/nicklaym1954
religious good night images
good morning images 5d
good night images best friend
3d good night images
magical good night images
https://picwali.in/beautiful-good-night-images/
good night images 2024 gif
good night images lord krishna
good night images tenor
independence day good night images
good night images 15 august
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cohia1980.bandcamp.com/album/twin-sister-delights-chapter-5-twins-wicked-evening
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196482
https://onedio.ru/profile/sveta-25198-2
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195615
https://anotepad.com/notes/iekfh3rq
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
http://www.babelcube.com/user/melissa-patterson
http://www.nfomedia.com/profile?uid=rOjRdkI
https://okwave.jp/profile/u3117399.html
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196911
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195285
Военный адвокат Запорожье. Бесплатная консультация
адвокат Запорожье
— это опытный специалист имеющий высшее юридическое образование, сдавший квалификационный государственный экзамен на право осуществления адвокатской деятельностью и специализирующийся в основном на военных делах
Вся правовая помощь военного адвоката осуществляется в индивидуальном порядке, грамотно, четко и в соответствии с действующими нормативно-правовыми актами.
Мы как военные юристы действуем не против органов Украины или министерства обороны, мы действуем во благо Украины — наших защитников и граждан Украины, которые попали в тяжелую жизненную ситуацию связанную с незнанием военного и действующего законодательства.
Поскольку, проявив патриотизм и чувство гражданской ответственности – став на защиту суверенитета страны, граждане участвующие и помогавшие в обороне после, становятся никому не нужными, особенно если военнослужащий стал инвалидом, потерял часть тела или конечность, и не может самостоятельно защитить свои права. Именно в таких ситуациях мы как военные адвокаты приходим на помощь, и добиваемся в установленном законом порядке справедливости, необходимых выплат, установление статуса, оформление пенсий, льгот и т.п.
Тоже касается, и получение отсрочки от мобилизации, когда например, безосновательно призывают сына у которого отец инвалид 2 группы, или мать прикованная из-за тяжелой болезни к постели, и требующая постороннего ухода. Это же относится и к военнослужащим, рапорта которых не регистрируются в канцелярии воинской части и полностью игнорируются, под прикрытием суеты боевых действий..
Именно в таких ситуациях, мы приходим на помощь и с помощью ЗАКОННЫХ методов правовой защиты, используя свой опыт полученный при ведении аналогичных военных дел добиваемся справедливости.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://permacultureglobal.org/users/58878-rhonda-naylor
https://okwave.jp/profile/u3117208.html
https://okwave.jp/profile/u3118337.html
https://www.metal-archives.com/users/juyhtgrf1995197819981964
https://wertrew1970.bandcamp.com/album/great-everyday-sex
I must thank you for the efforts you have put in writing this blog. I’m hoping to check out the same high-grade blog posts by you later on as well. In fact, your creative writing abilities has encouraged me to get my own blog now 😉
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://cannabis.net/user/151601
https://www.quia.com/profiles/chad370le
https://anotepad.com/notes/gfndy6yb
https://www.quia.com/profiles/chrisal224
https://rentry.org/vx756x4z
Right here is the right webpage for anybody who wishes to find out about this topic. You know so much its almost hard to argue with you (not that I really will need to…HaHa). You definitely put a fresh spin on a subject which has been discussed for years. Great stuff, just excellent.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/maksidrom1955
https://onedio.ru/profile/taaa-197-4
https://permacultureglobal.org/users/59507-maria-jensen
https://haveagood.holiday/users/351634
https://rentry.org/3rdhx4uf
슬롯 게임 추천
처음에는 Fang Jifan을 경멸했기 때문에 온갖 비꼬는 말을했습니다.처음에는 왜 생각을 못했을까, 보고를 했을까?
Great items from you, man. I’ve remember your stuff prior to and
you’re just extremely great. I really like what you have received here, really like
what you are stating and the best way in which you
say it. You’re making it entertaining and you still take
care of to stay it sensible. I can not wait to read far more from you.
That is really a tremendous web site.
Today, I went to the beachfront with my kids.
I found a sea shell and gave it to my 4 year old daughter and said “You can hear the ocean if you put this to your ear.” She placed the
shell to her ear and screamed. There was a hermit crab
inside and it pinched her ear. She never wants to go back!
LoL I know this is totally off topic but I had to tell someone!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://sdsadasd199619921968.diary.ru/
https://haveagood.holiday/users/350614
http://www.babelcube.com/user/samantha-grant
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3194508
https://rentry.org/ty245uxt
О команде Экипировка Эксперт
ТЕПЛОВИЗИОННЫЙ ПРИЦЕЛ GUIDE TR450
Боец, Экипировка Эксперт — это розничный интернет-магазин при оптовом складе. Это значит, что при должном количестве товара мы дадим очень хорошие цены.
Название взяли независимо от того, что наша страна сейчас проводит Специальную Военную Операцию, хорошая снаряга и экипировка нужна всегда. Готовишься в бой, мобилизован, привык активно проводить время или решил подготовить тревожный чемоданчик, мы поможем тебе. Наши клиенты: фонды, медики, такие же как ты бойцы СВО и обычные неравнодушные граждане.
ГАРПИЯ (ГРОМ) 6-КАНАЛЬНЫЙ ПОДАВИТЕЛЬ БПЛА, ОБЛЕГЧЕННЫЙ
https://ekipirovka.shop/katalog/taktika/sibz/plitonostsy/
Самое главное, что нужно о нас знать, мы детально объясняем, что и как работает, чтобы ты сделал правильный выбор не переплачивая.
Обращаясь к нам, не удивляйся, если ты получишь честный и жесткий ответ – часто случается так, что мы знаем лучше, что именно нужно нашему гостю. Особенно это касается мобилизованных без опыта боевых действий. Здесь ты можешь полагаться на нашу экспертность.
Одна из наших основных целей предоставить тебе возможность удобной и безопасной покупки: хоть за наличку, хоть по карте, хоть по счету. Повторимся, если нужна оптовая поставка, согласуем и отгрузим.
Именно от того, как ты производишь оплату, зависит цена заказа. Для нас важно предоставить тебе качественную экипировку и снаряжение соблюдая при этом законы нашей страны.
Боец, помни, мы помогаем фондам, нуждающимся людям, подразделениям в зоне СВО. Отчеты об этом вскоре будут опубликованы как на сайте, так и на наших каналах в социальных сетях. На эту деятельность уходит значительная часть выручки. Делая покупки в нашем магазине, ты помогаешь людям и фронту. Уверен, что это найдет отзыв в твоем сердце.
У нашей команды есть набор ценностей: честность, справедливость, сопереживание, взаимопомощь, мужество, патриотичность. Уверены, ты их разделяешь, и мы легко найдем общий язык.
Ну а если что-то пойдет не так, не руби с плеча, объясни, где мы ошиблись и поверь, мы разберемся и исправим.
Наш девиз “In hostem omnia licita” – по отношению к врагу дозволено все. Возьми этот девиз, он поможет тебе принять правильное решение в трудной ситуации, с честью выполнить боевую задачу и вернуться домой живым и здоровым!
bookmarked!!, I love your website.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://koreo1969.bandcamp.com/album/orgy-2
https://tubeteencam.com/user/alienka1985/profile
https://imageevent.com/plut951975
https://rentry.org/ebi69hfd
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3196105
Hi, I do believe your web site could possibly be having web browser compatibility issues. When I take a look at your website in Safari, it looks fine however, if opening in I.E., it’s got some overlapping issues. I merely wanted to give you a quick heads up! Other than that, great site.
I need to to thank you for this great read!! I certainly loved every little bit of it. I have got you book marked to check out new things you post…
Hi, I do believe this is an excellent blog. I stumbledupon it 😉 I am going to revisit yet again since i have bookmarked it. Money and freedom is the greatest way to change, may you be rich and continue to guide others.
Thankfulness to my father who shared with me about this blog, this website is in fact awesome.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/z77awmva
https://imageevent.com/kiska20091986
http://www.babelcube.com/user/brian-cuevas
http://www.nfomedia.com/profile?uid=rOiYYeC
https://www.hentai-foundry.com/user/esfwer1958/profile
This website was… how do I say it? Relevant!! Finally I’ve found something that helped me. Kudos!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.quia.com/profiles/dan397g
https://coka21950.diary.ru/
https://okwave.jp/profile/u3117947.html
https://chyoa.com/user/deal1967
https://www.hentai-foundry.com/user/bodyanvk1995/profile
Я работаю с компанией «лайф из гуд» уже несколько лет и могу с уверенностью сказать, что это одна из лучших финансовых компаний на рынке. Их продукты и услуги всегда были на высшем уровне, они помогли многим людям улучшить их финансовое положение. Я никогда не сталкивался с проблемами или задержками в выплатах, все было четко и прозрачно. Когда начались судебные процессы, я был в шоке. Как можно обвинять компанию, которая столько лет помогала людям, в мошенничестве? Это просто абсурд. Все клиенты и партнеры знают, что «лайф из гуд» всегда действовала честно и законно. Я уверен, что это дело сфабриковано, чтобы уничтожить успешную компанию и захватить ее активы. Надеюсь, что суд разберется и оправдает всех обвиняемых. Мы будем поддерживать «лайф из гуд» и бороться за их справедливое имя.
[url=https://www.infpol.ru/219600-roman-viktorovich-vasilenko-dostizheniya-i-uspekhi/]лайф из гуд[/url]
There is certainly a lot to find out about this issue. I like all the points you made.
Title: Mejores estrategias para jugar en Spin Casino Brasil: Consejos de los profesionales
Description: Descubre estrategias expertas y consejos profesionales para optimizar tus juegos en Spin Casino Brasil. ?Aprende de los mejores!
spin casino online
H1: Mejores estrategias para jugar en Spin Casino Brasil: Consejos de los profesionales
Bienvenidos al emocionante mundo de Spin Casino Brasil, donde la oportunidad de ganar se combina con el entretenimiento de primera clase. En este articulo, exploraremos estrategias efectivas y consejos profesionales que te ayudaran a maximizar tus experiencias de juego. Desde comprender la estructura del casino hasta aplicar tacticas especificas en tus juegos favoritos, te guiaremos paso a paso para mejorar tus habilidades y aumentar tus posibilidades de exito.
H2: Conociendo Spin Casino Brasil
Introduccion al Casino
Spin Casino Brasil se destaca en el mercado por ofrecer una experiencia de juego segura y dinamica. Con una licencia valida y un compromiso con la seguridad del jugador, este casino se presenta como una opcion confiable para los entusiastas de los juegos de azar en linea.
Variedad de Juegos
Explora la amplia gama de juegos que Spin Casino Brasil tiene para ofrecer. Desde tragamonedas clasicas hasta juegos de mesa sofisticados como el blackjack y la ruleta. Cada juego viene con una descripcion detallada de las reglas y los porcentajes de retorno al jugador (RTP), asegurando que los jugadores puedan elegir sabiamente segun sus preferencias.
Software y Tecnologia
Spin Casino Brasil utiliza software de los lideres de la industria, garantizando graficos de alta calidad y una interfaz de usuario fluida. Este apartado discutira como la tecnologia de Spin Casino Brasil mejora la experiencia del usuario y mantiene la integridad del juego.
Atencion al Cliente
El soporte al cliente es esencial en cualquier plataforma de juego en linea. Detalla como los jugadores pueden acceder al soporte 24/7 a traves de chat en vivo, correo electronico y telefono, asegurando que todas las consultas y problemas se resuelvan de manera eficiente.
H2: Estrategias generales de juego
Gestion del Bankroll
Un buen jugador sabe que la gestion del bankroll es crucial. Explica tecnicas para determinar cuanto dinero destinar al juego sin comprometer la estabilidad financiera personal. Se incluiran metodos para establecer limites de apuestas y la importancia de adherirse a estos limites.
Comprender las Probabilidades
El juego de casino no solo se trata de suerte; las matematicas juegan un papel crucial. Ofrece una explicacion sencilla de como entender las probabilidades y como pueden influir en las decisiones de juego. Este apartado demuestra como un conocimiento basico de estadisticas y probabilidad puede ayudar a tomar decisiones mas informadas.
Importancia de la Seleccion del Juego
No todos los juegos de casino ofrecen las mismas probabilidades de ganar. Discute como la eleccion del juego puede afectar las posibilidades de exito, destacando aquellos juegos que ofrecen mejores probabilidades y explicando por que algunos juegos son mas amigables con el jugador que otros.
Estrategias de Apuestas
Finalmente, se abordaran diversas estrategias de apuestas que los jugadores pueden emplear. Desde la estrategia conservadora hasta la mas arriesgada, cada estilo de apuesta sera discutido con ejemplos practicos, ayudando a los jugadores a entender cuando y como ajustar sus apuestas segun el flujo del juego.
H2: Estrategias especificas para juegos populares en Spin Casino Brasil
Este apartado se centra en las tacticas especializadas que los jugadores pueden emplear en algunos de los juegos mas populares disponibles en Spin Casino Brasil, asegurando que los jugadores de todos los niveles puedan optimizar sus oportunidades de ganar.
H3: Tragamonedas
Comprender la Varianza y el RTP
Inicia con una explicacion sobre la importancia del retorno al jugador (RTP) y la varianza de las tragamonedas. Describe como seleccionar maquinas con un RTP alto puede aumentar las posibilidades de ganancias a largo plazo y como la varianza afecta la frecuencia y cantidad de los pagos.
Estrategias de Apuestas
Detalla estrategias de apuestas eficaces para las tragamonedas, como ajustar el tamano de las apuestas basado en el ciclo de ganancias/perdidas y la importancia de maximizar las lineas de pago para aumentar las posibilidades de ganar en cada giro.
Aprovechar los Bonos
Explica como los jugadores pueden utilizar bonos y giros gratis ofrecidos por Spin Casino para maximizar sus jugadas sin aumentar el riesgo financiero, destacando la importancia de leer los terminos y condiciones asociados a estos bonos.
H3: Blackjack
Estrategia Basica
Proporciona una guia detallada sobre la estrategia basica de blackjack, incluyendo cuando pedir carta, plantarse, dividir o doblar la apuesta, basado en la mano del jugador y la carta visible del crupier.
Gestion del Riesgo
Discute la importancia de la gestion del riesgo y del bankroll especificamente para el blackjack, enfatizando como evitar las apuestas impulsivas y como adaptar el tamano de las apuestas en funcion de la situacion del juego.
Contar Cartas
Introduce conceptos basicos del conteo de cartas (donde legalmente sea aplicable), explicando como esta tecnica puede dar a los jugadores una ventaja estadistica sobre el casino y como practicar de manera efectiva sin infringir las reglas.
H3: Ruleta
Apuestas Exteriores vs. Interiores
Explica la diferencia entre apuestas exteriores e interiores, sus riesgos y beneficios, y por que las apuestas exteriores suelen ser una mejor opcion para aquellos que buscan minimizar riesgos.
Sistemas de Apuestas
Analiza varios sistemas de apuestas como el Martingala, Fibonacci y D’Alembert, evaluando su efectividad y como pueden ser aplicados en la ruleta para gestionar mejor las perdidas y potenciar las ganancias.
H3: Poker
Conocimientos Fundamentales de Poker
Detalla los conocimientos basicos del poker, como las reglas del Texas Hold’em, las clasificaciones de manos y la importancia de la posicion en la mesa.
Estrategias de Apuesta y Bluff
Describe estrategias avanzadas como el arte del bluff, saber leer a los oponentes, y la importancia de ajustar las apuestas basado en el estilo de juego de los contrincantes y la dinamica de la mesa.
Importancia del Control Emocional
Destaca la importancia de mantener el control emocional, como gestionar las ganancias y las perdidas, y por que un enfoque disciplinado y una mente clara son esenciales para el exito en el poker a largo plazo.
H2: Consejos de los profesionales
Aprender de la Experiencia
Introduccion a las lecciones aprendidas por jugadores experimentados, enfatizando la importancia de aprender tanto de los exitos como de los fracasos. Resalta como la experiencia es un recurso invaluable que puede ayudar a prever situaciones de juego y tomar decisiones mas informadas.
Adaptabilidad y Flexibilidad
Describe como los profesionales se adaptan a diferentes situaciones de juego, ajustando sus estrategias segun la dinamica del juego y las acciones de otros jugadores. Discute la importancia de ser flexible y no apegarse rigidamente a una sola estrategia.
Mantener la Disciplina
Enfatiza la necesidad de mantener la disciplina, especialmente en momentos de ganancias o perdidas significativas. Explica como la gestion efectiva del bankroll y el control emocional son cruciales para el exito a largo plazo en los juegos de casino.
H2: Maximizando bonificaciones y promociones
Entender los Tipos de Bonificaciones
Explica los diferentes tipos de bonificaciones que ofrece Spin Casino Brasil, como bonos sin deposito, bonos por deposito, y giros gratis. Detalla como cada tipo puede beneficiar a los jugadores dependiendo de su estilo de juego y objetivos.
Estrategias para Aprovechar Promociones
Ofrece estrategias especificas para aprovechar al maximo las promociones, como planificar depositos durante eventos de bonificacion y elegir promociones que complementen el estilo de juego del usuario. Discute la importancia de leer los terminos y condiciones para evitar sorpresas.
Cumpliendo con los Requisitos de Apuesta
Proporciona consejos practicos sobre como cumplir con los requisitos de apuesta sin comprometer el bankroll. Explora tecnicas para maximizar las oportunidades de ganancia mientras se juega a traves de los requisitos de apuesta de manera eficiente.
H2: Herramientas y recursos utiles
Software y Aplicaciones de Ayuda
Presenta software y aplicaciones que pueden ayudar a los jugadores a mejorar su juego, como calculadoras de probabilidades, diarios de juego, y herramientas de analisis de manos. Evalua como estas herramientas pueden proporcionar una ventaja competitiva.
Recursos Educativos
Recomienda libros, tutoriales en video y cursos que pueden proporcionar a los jugadores nuevos y experimentados conocimientos avanzados sobre estrategias de juego y psicologia del juego. Destaca recursos especificos que han sido bien recibidos por la comunidad de juego.
Foros y Comunidades
Alude a la importancia de participar en foros y comunidades en linea donde los jugadores pueden intercambiar estrategias, experiencias y consejos. Subraya como el aprendizaje colaborativo y el apoyo mutuo pueden mejorar la experiencia de juego y el desarrollo de habilidades.
Al aplicar las estrategias y consejos discutidos en este articulo, estaras bien equipado para afrontar los desafios que Spin Casino Brasil ofrece. Recuerda que la clave del exito en los juegos de casino reside no solo en conocer las reglas, sino en aplicar estrategias informadas y mantener una disciplina ferrea. Juega de manera responsable y dentro de tus limites para disfrutar de una experiencia tanto segura como emocionante. ?Buena suerte y que disfrutes de cada juego!
FAQ:
?Es seguro jugar en Spin Casino Brasil?
?Absolutamente! Spin Casino Brasil esta completamente regulado y ofrece un entorno seguro para todos sus jugadores. Disfruta de tu juego sin preocupaciones.
?Que juegos ofrecen las mejores probabilidades en Spin Casino Brasil?
Los juegos como el blackjack y el video poker generalmente ofrecen las mejores probabilidades, especialmente cuando aplicas las estrategias adecuadas.
?Puedo jugar en Spin Casino Brasil desde mi movil?
Si, Spin Casino Brasil ofrece una plataforma movil excelente que te permite jugar tus juegos favoritos desde cualquier lugar y en cualquier momento.
?Como puedo maximizar las bonificaciones ofrecidas por Spin Casino Brasil? Asegurate de leer los terminos y condiciones de las bonificaciones. Aprovecha los bonos de bienvenida, recargas y programas de fidelidad para obtener el maximo beneficio.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/varchik-199-5
https://www.hentai-foundry.com/user/gabriel921961/profile
https://feho1974.diary.ru/
https://rentry.org/wn5u3f9z
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3197205
I really like it whenever people get together and share ideas. Great website, continue the good work.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/351230
https://ivansmith1954.bandcamp.com/album/leprechaun
https://www.dnnsoftware.com/activity-feed/my-profile/userid/3195636
https://haveagood.holiday/users/351170
https://okwave.jp/profile/u3117114.html
Woah! I’m really loving the template/theme of this site. It’s simple, yet effective.
A lot of times it’s very hard to get that “perfect balance” between superb usability and visual appearance.
I must say you’ve done a amazing job with this.
Additionally, the blog loads very quick
for me on Chrome. Superb Blog!
Колокольцев – предатель Родины
Армия восстанет против ОПГ Колокольцева и Плугина
Целенаправленно разрушая кооператив для военных «Бест Вей», захватывая его активы в интересах своей камарильи, глава МВД и его команда ударили по тылу армии, наказав тысячи пайщиков кооператива – военных и членов их семей.
Приморский районный суд Санкт-Петербурга (судья Екатерина Богданова) сейчас рассматривает так называемое уголовное дело «Лайф-из-Гуд» – «Гермес» – «Бест Вей». В рамках этого дела два года почти непрерывно арестованы счета кооператива «Бест Вей», пайщиками которого являются тысячи военнослужащих, в том числе участники СВО, на которых около 4 млрд рублей. В уголовном деле речь идет прежде всего об обязательствах иностранной инвесткомпании «Гермес» перед более чем 200 клиентами этой компании.
Следствие ГУ МВД по Санкт-Петербургу и прокуратура города на Неве объявили кооператив, «Гермес» и «Лайф-из-Гуд» частью некоего единого холдинга, которого в природе никогда не существовало. Кооператив объявлен гражданским ответчиком по уголовному делу.
Общая сумма претензий потерпевших, согласно обвинительному заключению, 282 млн рублей – даже их (незаконное) изъятие для погашения долгов перед клиентами «Гермеса» никак бы не сказалось ни на операционной деятельности, ни на ликвидности кооператива. Но из раза в раз арестовываются абсолютно все средства на счетах, что полностью блокирует как покупку квартир кооперативом, так и возврат денег пайщикам, которые изъявили желание забрать свои паевые взносы.
Недвижимость, на которую претендовали пайщики, обесценивается, деньги обесцениваются – и этот беспредел продолжается уже более двух лет по ходатайствам ГСУ питерского главка МВД и Прокуратуры Санкт-Петербурга.
Кто-то влиятельный пытается наложить лапу именно на кооперативные 4 млрд – хотя это деньги пайщиков: физических лиц, рядовых граждан России, и как минимум соучастниками грабежа являются сотрудники правоохранительных органов.
Армейский кооператив
Кооператив изначально создавался военнослужащими – одной из его задач в 2014 году было решение жилищной проблемы военнослужащих, увольняемых в запас, эта задача сохранилась и в последующие годы. Из почти 20 тыс. его пайщиков – тысячи военнослужащих со всей России. Значительная часть из них – участники СВО.
В результате действий ОПГ Колокольцева сотни семей военнослужащих, участников СВО стали потерпевшими от действий правоохранительных органов. Налицо преступные действия против участников СВО.
Камарилья, которая устроила репрессии против кооператива – руководители следственной группы Сапетова и Винокуров, начальник ГСУ питерского главка МВД Негрозов и начальник этого главка Плугин, замминистра внутренних дел по следствию Лебедев, пресс-секретарь министра Волк, транслировавшая на всю страну ложь про десятки тысяч потерпевших от работы кооператива, сам министр внутренних дел Колокольцев, – ведет вредительскую деятельность, все эти люди – предатели Родины.
Разоряет экономику и подрывает тыл
Колокольцев и Ко разрушают экономику страны, фабрикуют дела. При этом его камарилья наживается на честном бизнесе.
Колокольцев – миллиардер! Его боевая подруга генеральша Волк – миллиардерша! Зам. начальника питерского ГСУ МВД полковник Винокуров, который управляет Колокольцевым, заставляя его подписываться под липовым уголовным делом «Лайф-из-Гуд» – «Гермес» – «Бест Вей», – миллиардер! Колокольцев превратил МВД в коммерческое антинародное образование и зарабатывает деньги на простых людях.
Колокольцев и Ко прицепили звезды на погоны и возомнили себя офицерами – на деле они тыловые вороватые крысы. Настоящие офицеры против бутафорских чинуш со звездами на погонах.
Министр внутренних дел дестабилизирует ситуацию в народе и армии, разоряет армию в тылу. Колокольцев со своими тыловыми крысами подрывают доверие народа к власти и отбивают желание людей идти на военную службу, потому что благодаря таким, как он, государство не может обеспечить безопасность военных в тылу.
Армия против Колокольцева
Армия восстанет против ОПГ Колокольцева. Военные требуют за совершенные преступления против народа, за воровство и коррупцию арестовать Колокольцева, Негрозова, Винокурова, Сапетову, оправдать и отпустить невинных людей – рядовых сотрудников «Лайф-из-Гуд», которые более двух лет томятся в тюрьме.
Новый министр обороны Белоусов наверняка поддержит своих бойцов, поможет им защитить собственные и своих семей имущественные интересы в тылу: по-другому и быть не может.
Одного любителя запускать руку в казну и любителя генеральш – Шойгу – уже отправили на пенсионную должность, Колокольцев должен последовать за ним: это – в интересах страны, в интересах народа!
Колокольцев – предатель Родины
Армия восстанет против ОПГ Колокольцева и Плугина
Целенаправленно разрушая кооператив для военных «Бест Вей», захватывая его активы в интересах своей камарильи, глава МВД и его команда ударили по тылу армии, наказав тысячи пайщиков кооператива – военных и членов их семей.
Приморский районный суд Санкт-Петербурга (судья Екатерина Богданова) сейчас рассматривает так называемое уголовное дело «Лайф-из-Гуд» – «Гермес» – «Бест Вей». В рамках этого дела два года почти непрерывно арестованы счета кооператива «Бест Вей», пайщиками которого являются тысячи военнослужащих, в том числе участники СВО, на которых около 4 млрд рублей. В уголовном деле речь идет прежде всего об обязательствах иностранной инвесткомпании «Гермес» перед более чем 200 клиентами этой компании.
Следствие ГУ МВД по Санкт-Петербургу и прокуратура города на Неве объявили кооператив, «Гермес» и «Лайф-из-Гуд» частью некоего единого холдинга, которого в природе никогда не существовало. Кооператив объявлен гражданским ответчиком по уголовному делу.
Общая сумма претензий потерпевших, согласно обвинительному заключению, 282 млн рублей – даже их (незаконное) изъятие для погашения долгов перед клиентами «Гермеса» никак бы не сказалось ни на операционной деятельности, ни на ликвидности кооператива. Но из раза в раз арестовываются абсолютно все средства на счетах, что полностью блокирует как покупку квартир кооперативом, так и возврат денег пайщикам, которые изъявили желание забрать свои паевые взносы.
Недвижимость, на которую претендовали пайщики, обесценивается, деньги обесцениваются – и этот беспредел продолжается уже более двух лет по ходатайствам ГСУ питерского главка МВД и Прокуратуры Санкт-Петербурга.
Кто-то влиятельный пытается наложить лапу именно на кооперативные 4 млрд – хотя это деньги пайщиков: физических лиц, рядовых граждан России, и как минимум соучастниками грабежа являются сотрудники правоохранительных органов.
Армейский кооператив
Кооператив изначально создавался военнослужащими – одной из его задач в 2014 году было решение жилищной проблемы военнослужащих, увольняемых в запас, эта задача сохранилась и в последующие годы. Из почти 20 тыс. его пайщиков – тысячи военнослужащих со всей России. Значительная часть из них – участники СВО.
В результате действий ОПГ Колокольцева сотни семей военнослужащих, участников СВО стали потерпевшими от действий правоохранительных органов. Налицо преступные действия против участников СВО.
Камарилья, которая устроила репрессии против кооператива – руководители следственной группы Сапетова и Винокуров, начальник ГСУ питерского главка МВД Негрозов и начальник этого главка Плугин, замминистра внутренних дел по следствию Лебедев, пресс-секретарь министра Волк, транслировавшая на всю страну ложь про десятки тысяч потерпевших от работы кооператива, сам министр внутренних дел Колокольцев, – ведет вредительскую деятельность, все эти люди – предатели Родины.
Разоряет экономику и подрывает тыл
Колокольцев и Ко разрушают экономику страны, фабрикуют дела. При этом его камарилья наживается на честном бизнесе.
Колокольцев – миллиардер! Его боевая подруга генеральша Волк – миллиардерша! Зам. начальника питерского ГСУ МВД полковник Винокуров, который управляет Колокольцевым, заставляя его подписываться под липовым уголовным делом «Лайф-из-Гуд» – «Гермес» – «Бест Вей», – миллиардер! Колокольцев превратил МВД в коммерческое антинародное образование и зарабатывает деньги на простых людях.
Колокольцев и Ко прицепили звезды на погоны и возомнили себя офицерами – на деле они тыловые вороватые крысы. Настоящие офицеры против бутафорских чинуш со звездами на погонах.
Министр внутренних дел дестабилизирует ситуацию в народе и армии, разоряет армию в тылу. Колокольцев со своими тыловыми крысами подрывают доверие народа к власти и отбивают желание людей идти на военную службу, потому что благодаря таким, как он, государство не может обеспечить безопасность военных в тылу.
Армия против Колокольцева
Армия восстанет против ОПГ Колокольцева. Военные требуют за совершенные преступления против народа, за воровство и коррупцию арестовать Колокольцева, Негрозова, Винокурова, Сапетову, оправдать и отпустить невинных людей – рядовых сотрудников «Лайф-из-Гуд», которые более двух лет томятся в тюрьме.
Новый министр обороны Белоусов наверняка поддержит своих бойцов, поможет им защитить собственные и своих семей имущественные интересы в тылу: по-другому и быть не может.
Одного любителя запускать руку в казну и любителя генеральш – Шойгу – уже отправили на пенсионную должность, Колокольцев должен последовать за ним: это – в интересах страны, в интересах народа!
Truly all kinds of excellent knowledge.
Feel free to surf to my blog post – http://links.musicnotch.com/mllermelinda
It’s nearly impossible to find experienced people on this topic, but you seem like you know what you’re talking about! Thanks
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://www.hentai-foundry.com/user/umisho1968/profile
https://tubeteencam.com/user/punkmen1963/profile
https://launchpad.net/~sdsw19501
https://wwoww1973.bandcamp.com/album/the-agreement-1
https://www.obesityhelp.com/members/bullkamc1997/about_me/
https://xrumer.ru/ – роблокс r34
https://xrumer.xyz/ – роблокс r34
i-guru – Ремонт Iphone и другой техники Apple в Минске
перепайка процессора iphone 13
Профессиональный ремонт любых устройств из линейки Apple, без посредников
80% ремонтов iPhone занимает около 20 минут
Несложные ремонты делаются быстро, модульный ремонт в большинстве случаев занимает не более 20 минут.
Почему выбирают i-guru
Ремонт в тот же день
Даже большинство сложных ремонтов выполняем день в день.
Гарантировано низкие цены
Мы беремся за сложные задачи, не передавая их сторонним исполнителям, и зарабатываем не как посредники.
Честная гарантия
Предлагаем честную гарантию на выполненные работы до 12 месцев в зависимости от вида работ.
Beneficial advice, Appreciate it.
Feel free to visit my blog :: https://Guyanaexpatforum.com/question/justin-bieber-can-%ec%b6%9c%ec%9e%a5%eb%a7%88%ec%82%ac%ec%a7%80-can-you-77/
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/reoe2odw
https://haveagood.holiday/users/350669
https://www.hentai-foundry.com/user/bodyanvk1995/profile
https://www.quia.com/profiles/angie436wa
https://tubeteencam.com/user/rokketa1996/profile
That is a good tip especially to those new to the blogosphere.
Simple but very precise info… Many thanks for sharing this one.
A must read post!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://haveagood.holiday/users/351118
http://www.nfomedia.com/profile?uid=rOjQYcB
https://cannabis.net/user/150928
https://imageevent.com/jeller1968
https://rentry.org/t7kw6p32
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://onedio.ru/profile/aterryan-199-0
https://anotepad.com/notes/s7ijrgyj
https://www.haikudeck.com/presentations/FqJB8kvl3z
https://cannabis.net/user/151073
https://dahaka19891950.diary.ru/
Премиум users get off on a inappropriate range of benefits not close by on a free package. · You will download documents at tied speeds without unrequired limitations. https://tezfiless.com/
Компания Септик-Нара-купить септик для частного дома
занимается продажей, установкой и обслуживанием септиков в Наро-Фоминске Наро-Фоминском районе. Основное направление деятельности нашей компании именно установка под ключ септиков любых видов и размеров. С момента основания нашей компании, мы произвели монтаж более 1000 септиков по Московской и Калужской области. Благодаря этому у нас огромный опыт работы с любыми станциями, представленными в нашем регионе!
Если Вы решили купить септик для дома или дачи, мы с радостью поможем Вам с выбором модели, доставим и установим септик на вашем участке в кратчайшие сроки.
Мы занимаемся продажей септиков таких марок: Топас Юнилос Астра Евролос Тверь Аквалос Дочиста Фекалов Волгарь Удача. Мы работаем напрямую с производителями септиков, поэтому Вы можете быть уверены, что не переплачиваете ни копейки. Вся продукция в нашей компании имеет соответствующие сертификаты и лицензии. Время выезда на осмотр, установки или привоза оборудования согласовывается с клиентами и выполняется в срок. Мы заботимся о своей репутации, поэтому выполняем работу надежно и быстро.
Если Вы не знаете, какой септик больше всего Вам подходит, мы предоставим консультацию и выезд специалиста на Ваш участок абсолютно бесплатно!
Благодаря приобретению качественных септиков в нашей компании, каждый клиент получает большое количество преимуществ:
» компактные размеры и небольшой вес устройства;
» доступная стоимость очистной установки;
» быстрый и простой монтаж;
» невысокая стоимость эксплуатации;
» отсутствие неприятных запахов;
» высокая производительность;
» длительный срок эксплуатации;
» высокая степень очистки;
» практически полностью автономный режим работы.
A fairly new player in the Russian darknet arena, blacksprut Blacksprut has quickly gained attention for its interesting features and growing popularity. While some aspects raise questions, it has the potential to become a prominent figure in the darknet scene.
Features:
Blacksprut https://bs-gl-darknet.com offers an “Instant Transactions” feature, inspired by the success of similar systems on other platforms like Hydra. Couriers hide goods within the city and provide buyers with coordinates, adding an adventurous element to the purchasing process.
Attractive element of content. I simply stumbled upon your website and in accession capital to assert that I get in fact loved account your weblog posts. Anyway I’ll be subscribing in your augment or even I success you access persistently fast.
https://thetvdb.plex.tv/movies/real-gangsters
Hi there everyone, it’s my first go to see at this web site, and post is truly fruitful for me, keep up posting these types of articles or reviews.
https://thetvdb.plex.tv/movies/real-gangsters
You actually suggested this well.
my web site :: https://forum.atwimamponuarb.com/viewtopic.php?id=870150
Ирина Волк: расследование уголовного дела пирамиды Life is Good завершено
[url=https://telltrue.net/eto-lohotron/chernyj-spisok-lohotronov-utl-coinexfom-uno-club-bitlans-profitcrypto/?ysclid=ly1sujvcbx993980390]первый анальный секс[/url]
Официальный представитель МВД России Ирина Волк сообщила об окончании предварительного расследования уголовного дела Life is Good. Материалы с утвержденным обвинительным заключением в отношении десяти человек направлены в Приморский районный суд Санкт-Петербурга для рассмотрения по существу. Они обвиняются в организации деятельности по привлечению денежных средств и мошенничестве, совершенных в особо крупном размере в составе преступного сообщества. Всего, по версии следствия, в состав криминальной организации входили 35 соучастников. Потерпевшими по уголовному делу признаны 221 человек – люди заключили договоры с целью получения пассивного дохода. Общая сумма материального ущерба составила свыше 280 млн рублей.
«Кроме того, в деятельность финансовой пирамиды были вовлечены более 18 тысяч россиян. Они вложили в жилищный кооператив более 15 миллиардов рублей, два миллиарда из которых организаторы криминальной схемы присвоили. Десять предполагаемых участников преступного сообщества были задержаны в феврале 2022 года оперативниками ГУЭБиПК МВД России совместно с коллегами из Санкт-Петербурга и сотрудниками ФСБ России. Остальные обвиняемые в противоправной деятельности объявлены в розыск. Возбуждено уголовное дело по признакам преступления, предусмотренного частью 3 статьи 210, частью 4 статьи 159, частью 2 статьи 172.2 УК РФ. Расследование было взято на контроль руководством Следственного департамента МВД России», – сообщила Ирина Волк.
По информации МВД, в общей сложности полицейскими и сотрудниками ФСБ проведено более 100 обысков по местам проживания фигурантов и в офисах подконтрольных им организаций. Изъята компьютерная техника, средства связи, документы и другие предметы, имеющие доказательственное значение.
«В качестве обеспечительной меры по возмещению причиненного материального ущерба наложен арест на более чем 2300 находящихся в собственности кооператива квартир, а также на денежные средства на счетах компании в сумме свыше 3,7 млрд рублей. Также арестованы активы фигурантов на общую сумму порядка 1 млрд рублей», – рассказала генерал-майор полиции.
По версии следствия, противоправная деятельность велась с 2014 года на территории девяти регионов России. Злоумышленниками были созданы подконтрольные организации с сетью региональных филиалов, которые работали под единым брендом «Лайф из Гуд». Гражданам предлагали стать пайщиками кооператива, обещая либо новое жилье, либо получение дохода в размере до 30% годовых. При этом реальным инвестированием вкладов фигуранты не занимались. Приобретение недвижимости и обещанные выплаты осуществлялись за счет привлечения средств новых клиентов. Кроме того, граждан вводили в заблуждение об уровне доходности вложений, указывая в договорах параметры гораздо ниже обещанных. Полученные от вкладчиков деньги похищались.
Ранее стало известно, что Приморский районный суд Санкт-Петербурга заочно арестовал топ-лидера инвестиционного проекта Life is Good в Набережных Челнах Сергея Санникова и его жену Юлию. Также суд избрал меру пресечения в виде заключения под стражу и в отношении директора холдинга Игоря Маланчука. В июле этого года они были объявлены в федеральный розыск – соответствующая информация появилась в базе данных министерства.
Кооператив для военных
Как пайщики — участники СВО относятся к событиям вокруг «Бест Вей»
Уголовное дело №12101400038003637
Потребительский кооператив «Бест Вей» оказался затронут уголовным делом, касающимся в основном иностранной инвесткомпании «Гермес», которое сейчас рассматривается Приморским районным судом Санкт-Петербурга. Более двух лет более 3,5 млрд рублей на счетах кооператива почти непрерывно арестованы по ходатайству сначала ГСУ ГУ МВД по Санкт-Петербургу, а затем Прокуратуры Санкт-Петербурга: пайщики не имеют возможности ни приобрести недвижимость, ни вернуть средства.
По данным совета потребительского кооператива «Бест Вей», в числе его пайщиков, страдающих от блокировки средств, тысячи военнослужащих, в том числе сотни участников СВО, часть из которых успела приобрести квартиру, часть собирает или собрала первоначальный взнос, а часть — планировала вступить в кооператив. «СП» пообщалась с некоторыми из них и их родственниками, чтобы узнать отношение к «Бест Вей» и событиям вокруг кооператива.
«К кооперативу отношение очень хорошее»
Александр Голдман на СВО с мая 2022 года как доброволец, три ранения. Пришел на СВО рядовым, сейчас — начальник штаба батальона. Был пайщиком кооператива, сейчас пайщик — его мама.
«С помощью кооператива в 2019 году приобретена двухкомнатная квартира во Владивостоке, в которой проживает мама, — рассказывает он. — Расплачиваемся за нее, в ближайшие месяцы намерены погасить задолженность перед кооперативом и оформить квартиру в собственность. К кооперативу отношение очень хорошее, полностью его поддерживаю — он дает возможность без больших переплат приобрести недвижимость. К действиям в отношении кооператива отношусь отрицательно, так как „Бест Вей“ — единственная возможность приобрести жилье в рассрочку».
«Происходящее вокруг кооператива вызывает шок»
Гвардии рядовой Глушков Иван Васильевич — пайщик кооператива из Челябинской области. Танкист, мобилизованный, проходил службу в 15-й отдельной гвардейской мотострелковой Александрийской бригаде 2-й гвардейской общевойсковой армии ЦВО. Погиб 11 октября 2023 года.
Рассказывает его вдова Татьяна Неручева:
«Мы внесли первоначальный паевый взнос и во второй половине 2021 года встали в очередь на приобретение квартиры в Челябинске, когда начались события вокруг кооператива — была заблокирована его возможность приобретать недвижимость и заблокированы его счета. Наша очередь должна была подойти примерно через год. Мы заявили для приобретения небольшую квартиру, но планировали при покупке увеличить ее стоимость до 3 млн и купить двухкомнатную квартиру — с увеличением первоначального паевого взноса: уставом кооператива это позволяется, а затем переехать из Коркино в Челябинск. Сейчас 3 млн хватит только на небольшую квартиру-студию метров 25, то есть мы понесли материальный ущерб из-за блокирования деятельности кооператива, так как лишены были возможности приобрести квартиру, на которую собрали первоначальный паевый взнос. Работа кооператива заблокирована, счета арестованы уже более двух лет. В наследование пая я только вступаю, так как муж очень долго считался пропавшим без вести — долго шла экспертиза, и свидетельство о смерти мы получили только 18 июня этого года».
Татьяна — юрист: «Как у юриста у меня происходящее вокруг кооператива вызывает шок, и любой непредвзятый юрист вам скажет то же самое». «Кооператив, — подчеркивает она, — абсолютно прозрачен, он полностью соответствует законодательству о кооперации — что и подтверждалось многократно государственными органами и судами. Если бы мы с мужем не были уверены, что все прозрачно и законно, мы бы не вкладывали в него деньги. Мы не подавали заявление о выходе из кооператива. Я, несмотря ни на что, жду счастливого завершения рукотворного кризиса вокруг кооператива. Для меня это еще и память о муже — он продал долю в квартире, которая ему принадлежала, чтобы вложиться в кооператив. Кроме того, мне хочется понять — до какого маразма может дойти ситуация у нас в государстве в плане незаконных действий в отношении организации, которая по тем или иным причинам не понравилась каким-то чиновникам?».
«Рассчитываю, что ситуация закончится благополучно»
Сергей Логинов, рядовой, мобилизованный, представлен к награде «Честь и доблесть».
«Пять лет назад я стал пайщиком кооператива, участвовал в накопительной программе, планировал прибрести однокомнатную квартиру в Самарской области. Уже нужно было подбирать объект недвижимости и вставать в очередь на покупку, как работа кооператива была заблокирована по инициативе правоохранительных органов, и это продолжается уже более двух лет. По-прежнему надеюсь получить квартиру и рассчитываю, что ситуация закончится благополучно. Поддерживаю кооператив».
«В кооперативе минимум переплат — несопоставимо с ипотекой»
Егор Ивков, офицер флота, дирижер военного оркестра, пайщик кооператива с 2019 года.
«Я нахожусь в накопительной программе — планировал покупку квартиры в Санкт-Петербурге. До постановки в очередь на покупку дело не дошло, но сумма внесена серьезная — и на два года все зависло. Есть друзья-военнослужащие, которые также являются пайщиками и тоже накапливали деньги на первоначальный паевый взнос — они, как и я, не могут ни продолжать накапливать, ни получить деньги обратно, потому что счета арестованы. Отношение наше к ситуации, создавшейся вокруг кооператива, крайне негативное».
«Кооператив поддерживаю, — говорит пайщик. — Моя сестра живет в квартире, приобретенной с помощью „Бест Вей“ — у нее многодетная семья, сейчас кооперативная квартира переходит в ее собственность. Минимум переплат — несопоставимо с ипотекой. Надеюсь, что ситуация с арестом счетов разрешится в ближайшее время».
I’m not sure why but this website is loading extremely slow
for me. Is anyone else having this problem or is it a problem on my end?
I’ll check back later on and see if the problem still
exists.
It’s the best time to make some plans for the future and it’s time to be happy.
I’ve learn this publish and if I may I want to counsel you few interesting issues or tips.
Maybe you could write subsequent articles relating to this article.
I want to learn more issues about it!
Hello! I know this is kind of off topic but I was wondering which blog platform are you using for this website?
I’m getting sick and tired of WordPress because I’ve had problems with hackers
and I’m looking at options for another platform. I would be awesome if you
could point me in the direction of a good platform.
Звон Колокольцева. Питерская полицейская мафия виляет Министерством внутренних дел
Пайщики ПК Бествей
Поздравляем вас, гражданин министр, соврамши!
Выступая прошлой осенью в Совете Федерации, министр внутренних дел Владимир Колокольцев рассказывал, о так называемом уголовном деле «Лайф-из-Гуд» – «Гермес» – «Бест Вей», обещал миллиарды рублей ущерба и десятки тысяч потерпевших. Пресс-служба МВД под руководством его боевой подруги Ирины Волк заявила о том, что вскрыта деятельность крупнейшей в истории России финансовой пирамиды.
Однако в уголовном деле, расследованном или, вернее сказать, изготовленном ГСУ питерского главка МВД, которое в феврале начал рассматривать Приморский районный суд Санкт-Петербурга, 282 млн рублей ущерба и 221 лицо, признанное следствием потерпевшим: никаких миллиардов и десятков тысяч потерпевших.
Министр и его Волк публично солгали – на основе информации, переданной замначальника ГСУ руководителем следственной группы полковником юстиции А.Н. Винокуровым, фактически даже руководившей СГ его заместительницей майором, а затем подполковником юстиции Е.А. Сапетовой по материалам, состряпанным опером УЭБиПК питерского главка МВД майором полиции А.Ю. Машевским, при попустительстве (или соучастии?) начальника ГСУ Негрозова и замначальника Следственного департамента федерального министерства Вохмянина. Полковник и подполковник с примерно пятого-шестого уровня иерархии МВД виляет генералом полиции Российской Федерации, постоянным членом Совета безопасности России – это позор для государства.
Так называемые потерпевшие и реально пострадавшие
Большинство «потерпевших» на суде заявляют суммы около 1–2 млн рублей, при этом в ходе судебного следствия выясняется, что они, как правило, получали немалый доход, причем, они еще и налоговые/валютные преступники, так как этот доход не декларировали. Среди «потерпевших» есть граждане, заявляющие смехотворные суммы в 50–70 тыс., то есть количество потерпевших специально накручивалось следствием. И даже с этим накручиванием удалось набрать так мало – учитывая, что у «Гермеса», по данным самого же следствия, более 200 тыс. клиентов в России, а в кооперативе «Бест Вей» – около 20 тыс. пайщиков.
То есть большинство и клиентов «Гермеса», и пайщиков кооператива не считают себя потерпевшими от деятельности этих организаций.
Судя по тысячам обращений во все инстанции, нескольким волнам митингов, прокатившихся по России, они считают себя потерпевшими от деятельности органов внутренних дел.
Ведь именно завербованный питерским УЭБиПК сисадмин российской платежной системы «Гермеса» Набойченко заблокирован и разгромил в феврале 2022 года эту платежную систему, повесив на сайте дисклеймер: «Обращайтесь в правоохранительные органы», что на месяцы прекратило вывод средств. Именно действия правоохранительных органов в отношении компании до и после затруднили вывод средств. Дело в том, что для вывода средств многими использовался механизм p2p, позволяющий не платить комиссию, то есть для вывода средств нужно, чтобы кто-то вносил средства (что, понятно, резко сократилось из-за уголовного дела) и происходил обмен. Однако этот способ не единственный, вывод средств так или иначе осуществляется.
Тысячи пайщиков кооператива тем более не считают себя потерпевшими от его деятельности – потому что именно правоохранительные органы воспрепятствовали приобретению недвижимости с помощью кооператива, а она из-за более чем двухлетнего ареста его счетов, на которых около 4 млрд рублей, не может быть приобретена по прежней цене.
Именно правоохранительные органы прямо запрещают выплаты пайщикам кооператива, решившим забрать свой пай, – даже по исполнительным листам судов. И клиентам «Гермеса», и пайщикам кооператива правоохранительными органами нанесен колоссальный ущерб – и материальный, и моральный, который они намерены взыскать с государства.
«Следователи»-преступники должны сидеть в тюрьме
Весьма скромный результат следствия МВД был достигнут откровенно преступным путем.
ГСУ Санкт-Петербурга беспредел
1. Некоторые из преступлений следствия были фактически признаны судами. 1 декабря прошлого года Приморский районный суд города Санкт-Петербурга признал незаконным, нарушающим УПК фактический отказ кооперативу в ознакомлении с материалами уголовного дела.При рассмотрении дела в суде выяснилось, что следственная группа ГСУ питерского главка МВД, формально руководимая замначальника ГСУ полковником юстиции А.Н. Винокуровым, а фактически – подполковником юстиции Е.А. Сапетовой, подделала документы. Автор подделки – Сапетова – еще в феврале была уволена из ГСУ «по собственному желанию».
Уличенная адвокатами кооператива в нарушении УПК, следственная группа составила письмо об удовлетворении ходатайства задним числом и попыталась представить дело так, что кооператив не получил письмо по своей вине. Ложь была выявлена в том числе и с помощью системы электронного документооборота питерского главка МВД.
2. Подделка документов была вынужденным преступлением для сокрытия более серьезного: незаконного содержания под стражей. Следственная группа грубо нарушила права гражданских истцов и ответчиков, потому что без этого нарушения она не успела за 30 суток до истечения предельного срока содержания четверых обвиняемых под стражей начать ознакомление обвиняемых с материалами дела –а это было единственное основание продления им срока содержания под стражей свыше предельного.
Следственная группа из-за спешки даже толком не смогла завершить следственные действия, незаконно вела параллельное расследование по «резервному» делу, но позднее, отбросив стыд, из-за отсутствия материала для составления «нужного» обвинительного заключения, незаконно продолжила расследование «основного» дела, в том числе проводила следственные действия, которые, согласно УПК, невозможны после начала ознакомления обвиняемых с материалами дела. Все эти ухищрения были необходимы для того, чтобы ни в коем случае не выпускать обвиняемых и продолжать держать их в заложниках.
3. Де-факто происходит уголовное наказание неосужденных людей – четверо подсудимых уже второй год сидят в тюрьме. При этом наказываются явно ни в чем неповинные люди – технические сотрудники «Лайф-из-Гуд»: даже если предположить, что действительно работала пирамида – что опровергается показаниями свидетелей самого обвинения, которые сообщают суду, что компания «Гермес» хорошо работала, они были довольны получаемым доходом, и проблемы начались после того, как российская платежная система компании была обрушена завербованным полицией петербургским сисадмином компании Набойченко. Все подсудимые – технические сотрудники компании «Лайф-из-Гуд» и ни к каким управленческим решениям отношения никогда не имели. Их взяли в заложники для того, чтобы они дали показания на руководство компании.
4. Еще одно преступление – заведомо подложное постановление руководителя следственной группы А.Н. Винокурова о привлечении кооператива «Бест Вей» в качестве гражданского ответчика на 16 млрд рублей, тогда как сумма ущерба в уголовном деле –282 млн, и в деле нет ни одного искового заявления – даже на 100 рублей.
5. Следствие стимулировало двух особенно активных так называемых потерпевших написать заявления о моральном ущербе на миллиард (!) рублей каждое – исключительно для ложного обоснования ареста активов кооператива, но понятно, что это ничтожные документы, так как моральный ущерб во всех случаях, не связанных с причинением смерти, присуждается российскими судами в размере не более десятков тысяч рублей.
6. Ни один из «потерпевших» не доказал обоснованность своих претензий в гражданском суде. При этом арестованы активы кооператива почти на 4 млрд рублей и активы частных лиц на такую же сумму. Это не что иное, как попытка захвата активов при участии органов внутренних дел некой заинтересованной группой клиентов «Гермеса» – необязательно из числа «потерпевших», то есть коррупционное преступление, которое упорно игнорирует ГУСБ МВД.
Механизм для такого захвата есть – это передача средств под управление Федерального общественно-государственного фонда по защите прав вкладчиков и акционеров. Понятно, почему не ограничиваются активами подсудимых и обвиняемых: этого недостаточно для удовлетворения аппетитов тех, кто стоит за заказным уголовным делом. И понятно, почему одно юридическое лицо – кооператив «Бест Вей» – незаконно пытаются привлечь к ответственности за другое – компанию «Гермес»: активы «Гермеса» – за рубежом.
7. Следствие, а теперь и прокуратура совершают еще одно преступление: незаконно удерживает средства пайщиков кооператива, отказываясь их вернуть, то есть совершают хищение.
Звон Колокольцева. Питерская полицейская мафия виляет Министерством внутренних дел
Министр Колокольцев
Поздравляем вас, гражданин министр, соврамши!
Выступая прошлой осенью в Совете Федерации, министр внутренних дел Владимир Колокольцев рассказывал, о так называемом уголовном деле «Лайф-из-Гуд» – «Гермес» – «Бест Вей», обещал миллиарды рублей ущерба и десятки тысяч потерпевших. Пресс-служба МВД под руководством его боевой подруги Ирины Волк заявила о том, что вскрыта деятельность крупнейшей в истории России финансовой пирамиды.
Однако в уголовном деле, расследованном или, вернее сказать, изготовленном ГСУ питерского главка МВД, которое в феврале начал рассматривать Приморский районный суд Санкт-Петербурга, 282 млн рублей ущерба и 221 лицо, признанное следствием потерпевшим: никаких миллиардов и десятков тысяч потерпевших.
Министр и его Волк публично солгали – на основе информации, переданной замначальника ГСУ руководителем следственной группы полковником юстиции А.Н. Винокуровым, фактически даже руководившей СГ его заместительницей майором, а затем подполковником юстиции Е.А. Сапетовой по материалам, состряпанным опером УЭБиПК питерского главка МВД майором полиции А.Ю. Машевским, при попустительстве (или соучастии?) начальника ГСУ Негрозова и замначальника Следственного департамента федерального министерства Вохмянина. Полковник и подполковник с примерно пятого-шестого уровня иерархии МВД виляет генералом полиции Российской Федерации, постоянным членом Совета безопасности России – это позор для государства.
Так называемые потерпевшие и реально пострадавшие
Большинство «потерпевших» на суде заявляют суммы около 1–2 млн рублей, при этом в ходе судебного следствия выясняется, что они, как правило, получали немалый доход, причем, они еще и налоговые/валютные преступники, так как этот доход не декларировали. Среди «потерпевших» есть граждане, заявляющие смехотворные суммы в 50–70 тыс., то есть количество потерпевших специально накручивалось следствием. И даже с этим накручиванием удалось набрать так мало – учитывая, что у «Гермеса», по данным самого же следствия, более 200 тыс. клиентов в России, а в кооперативе «Бест Вей» – около 20 тыс. пайщиков.
То есть большинство и клиентов «Гермеса», и пайщиков кооператива не считают себя потерпевшими от деятельности этих организаций.
Судя по тысячам обращений во все инстанции, нескольким волнам митингов, прокатившихся по России, они считают себя потерпевшими от деятельности органов внутренних дел.
Ведь именно завербованный питерским УЭБиПК сисадмин российской платежной системы «Гермеса» Набойченко заблокирован и разгромил в феврале 2022 года эту платежную систему, повесив на сайте дисклеймер: «Обращайтесь в правоохранительные органы», что на месяцы прекратило вывод средств. Именно действия правоохранительных органов в отношении компании до и после затруднили вывод средств. Дело в том, что для вывода средств многими использовался механизм p2p, позволяющий не платить комиссию, то есть для вывода средств нужно, чтобы кто-то вносил средства (что, понятно, резко сократилось из-за уголовного дела) и происходил обмен. Однако этот способ не единственный, вывод средств так или иначе осуществляется.
Тысячи пайщиков кооператива тем более не считают себя потерпевшими от его деятельности – потому что именно правоохранительные органы воспрепятствовали приобретению недвижимости с помощью кооператива, а она из-за более чем двухлетнего ареста его счетов, на которых около 4 млрд рублей, не может быть приобретена по прежней цене.
Именно правоохранительные органы прямо запрещают выплаты пайщикам кооператива, решившим забрать свой пай, – даже по исполнительным листам судов. И клиентам «Гермеса», и пайщикам кооператива правоохранительными органами нанесен колоссальный ущерб – и материальный, и моральный, который они намерены взыскать с государства.
«Следователи»-преступники должны сидеть в тюрьме
Весьма скромный результат следствия МВД был достигнут откровенно преступным путем.
Потерпевшие Лайф из Гуд
1. Некоторые из преступлений следствия были фактически признаны судами. 1 декабря прошлого года Приморский районный суд города Санкт-Петербурга признал незаконным, нарушающим УПК фактический отказ кооперативу в ознакомлении с материалами уголовного дела.При рассмотрении дела в суде выяснилось, что следственная группа ГСУ питерского главка МВД, формально руководимая замначальника ГСУ полковником юстиции А.Н. Винокуровым, а фактически – подполковником юстиции Е.А. Сапетовой, подделала документы. Автор подделки – Сапетова – еще в феврале была уволена из ГСУ «по собственному желанию».
Уличенная адвокатами кооператива в нарушении УПК, следственная группа составила письмо об удовлетворении ходатайства задним числом и попыталась представить дело так, что кооператив не получил письмо по своей вине. Ложь была выявлена в том числе и с помощью системы электронного документооборота питерского главка МВД.
2. Подделка документов была вынужденным преступлением для сокрытия более серьезного: незаконного содержания под стражей. Следственная группа грубо нарушила права гражданских истцов и ответчиков, потому что без этого нарушения она не успела за 30 суток до истечения предельного срока содержания четверых обвиняемых под стражей начать ознакомление обвиняемых с материалами дела –а это было единственное основание продления им срока содержания под стражей свыше предельного.
Следственная группа из-за спешки даже толком не смогла завершить следственные действия, незаконно вела параллельное расследование по «резервному» делу, но позднее, отбросив стыд, из-за отсутствия материала для составления «нужного» обвинительного заключения, незаконно продолжила расследование «основного» дела, в том числе проводила следственные действия, которые, согласно УПК, невозможны после начала ознакомления обвиняемых с материалами дела. Все эти ухищрения были необходимы для того, чтобы ни в коем случае не выпускать обвиняемых и продолжать держать их в заложниках.
3. Де-факто происходит уголовное наказание неосужденных людей – четверо подсудимых уже второй год сидят в тюрьме. При этом наказываются явно ни в чем неповинные люди – технические сотрудники «Лайф-из-Гуд»: даже если предположить, что действительно работала пирамида – что опровергается показаниями свидетелей самого обвинения, которые сообщают суду, что компания «Гермес» хорошо работала, они были довольны получаемым доходом, и проблемы начались после того, как российская платежная система компании была обрушена завербованным полицией петербургским сисадмином компании Набойченко. Все подсудимые – технические сотрудники компании «Лайф-из-Гуд» и ни к каким управленческим решениям отношения никогда не имели. Их взяли в заложники для того, чтобы они дали показания на руководство компании.
4. Еще одно преступление – заведомо подложное постановление руководителя следственной группы А.Н. Винокурова о привлечении кооператива «Бест Вей» в качестве гражданского ответчика на 16 млрд рублей, тогда как сумма ущерба в уголовном деле –282 млн, и в деле нет ни одного искового заявления – даже на 100 рублей.
5. Следствие стимулировало двух особенно активных так называемых потерпевших написать заявления о моральном ущербе на миллиард (!) рублей каждое – исключительно для ложного обоснования ареста активов кооператива, но понятно, что это ничтожные документы, так как моральный ущерб во всех случаях, не связанных с причинением смерти, присуждается российскими судами в размере не более десятков тысяч рублей.
6. Ни один из «потерпевших» не доказал обоснованность своих претензий в гражданском суде. При этом арестованы активы кооператива почти на 4 млрд рублей и активы частных лиц на такую же сумму. Это не что иное, как попытка захвата активов при участии органов внутренних дел некой заинтересованной группой клиентов «Гермеса» – необязательно из числа «потерпевших», то есть коррупционное преступление, которое упорно игнорирует ГУСБ МВД.
Механизм для такого захвата есть – это передача средств под управление Федерального общественно-государственного фонда по защите прав вкладчиков и акционеров. Понятно, почему не ограничиваются активами подсудимых и обвиняемых: этого недостаточно для удовлетворения аппетитов тех, кто стоит за заказным уголовным делом. И понятно, почему одно юридическое лицо – кооператив «Бест Вей» – незаконно пытаются привлечь к ответственности за другое – компанию «Гермес»: активы «Гермеса» – за рубежом.
7. Следствие, а теперь и прокуратура совершают еще одно преступление: незаконно удерживает средства пайщиков кооператива, отказываясь их вернуть, то есть совершают хищение.
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://smalik1954.bandcamp.com/album/twincest-is-best-part-i
http://www.babelcube.com/user/ali-landan
https://www.obesityhelp.com/members/adrial19781954/about_me/
https://rentry.org/q2nik3xc
https://poxah41972.bandcamp.com/album/summer-at-pond-cove-chapter-05
Somebody essentially help to make critically articles I’d state.
That is the first time I frequented your web page and to this point?
I surprised with the research you made to make this particular put up amazing.
Wonderful process!
МОСКВА, 7 мая — РИА Новости. Правительство России, возглавляемое Михаилом Мишустиным, уйдет в отставку во вторник.
Сразу после инаугурации президента Владимира Путина нынешний кабинет министров сложит полномочия перед вступившим в должность главой государства.
Прогон по комментария
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://rentry.org/p67gckf8
https://rentry.org/vqbpa5qs
https://www.metal-archives.com/users/ololjkeee1965
https://www.haikudeck.com/presentations/MuWoluf6GU
https://onedio.ru/profile/iibonni-196-3
obviously like your web-site however you have to take a look at
the spelling on quite a few of your posts. Several of them are rife with spelling issues
and I in finding it very bothersome to inform the truth then again I’ll surely come back again.
Hurrah, that’s what I was seeking for, what a material!
existing here at this webpage, thanks admin of this website.
Also visit my homepage agen slot online
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://tubeteencam.com/user/kiruli4ka1970/profile
https://www.metal-archives.com/users/shenlu1983
https://kos1111971.diary.ru/
https://dra671957.diary.ru/
https://www.obesityhelp.com/members/andreyjs1955/about_me/
You actually make it seem so easy with your presentation but I find this matter to be actually
something that I think I would never understand.
It seems too complex and very broad for me. I’m looking forward for your
next post, I’ll try to get the hang of it!
There are many popular live sex cam sites that cater to all different preferences and desires. Some of the most popular ones include Chaturbate, MyFreeCams, LiveJasmin, and Flirt4Free.
Chaturbate is known for its diverse selection of cam models, ranging from amateur performers to professional porn stars. It offers a unique “tip-based” system, where viewers can tip the performers for special requests or to show their appreciation.
MyFreeCams is a popular choice for those looking for a more personalized experience, as many of the models offer private shows for a fee. It also has a large community aspect, with forums and chat rooms for viewers to interact with each other and the models.
LiveJasmin is known for its high-quality video and audio, making it a top choice for viewers who value a visually stimulating experience. It also has a wide range of categories, allowing viewers to easily find the type of performer they are looking for.
Flirt4Free is a popular site for those looking for a more intimate and interactive experience. It offers a variety of features such as cam-to-cam shows and interactive sex toys, making it a favorite among viewers who enjoy a more immersive experience.
Overall, these live sex cam sites offer a diverse range of performers and features to cater to all types of desires and preferences. Their popularity shows that the demand for live sex cams continues to grow as people seek out new and exciting forms of sexual entertainment.
https://imageevent.com/maksdark1977
http://www.babelcube.com/user/candice-lowe
https://www.metal-archives.com/users/vitaliya1963
https://www.hentai-foundry.com/user/lflflflf1974/profile
https://www.hentai-foundry.com/user/mucaltin1990/profile
Люди, вы видите, что творится с «Бест Вей»? Эти сволочи из МВД, под руководством Колокольцева, просто издеваются над нами! Они выдумали обвинения, солгали про миллиарды ущерба, а на деле – только 282 миллиона и 221 человек. Эти уроды в погонах, Винокуров и Сапетова, подделывают документы, незаконно держат людей в тюрьме. Четверо невинных уже второй год сидят за решеткой! А что с нашими деньгами? Заблокированы! Мы не можем их вывести, не можем купить недвижимость! Эти мрази думают, что могут делать всё, что угодно. Мы не оставим это просто так! Мы будем бороться до конца, чтобы эти твари понесли заслуженное наказание!
I got what you intend,bookmarked, very decent website.
My website: анал порно