Diffusion MRI microstructural models in the cervical spinal cord – application, normative values, and correlations with histological analysis
Kurt G. Schilling, Samantha By, Haley Feiler, Bailey Box, Kristin P. O’Grady, Atlee Witt, Bennett A. Landman, Seth A. Smith. “Diffusion MRI microstructural models in the cervical spinal cord – application, normative values, and correlations with histological analysis”. NeuroImage. doi: 10.1016/j.neuroimage.2019.116026. 2019.
Full text: https://www.ncbi.nlm.nih.gov/pubmed/31326569
Abstract
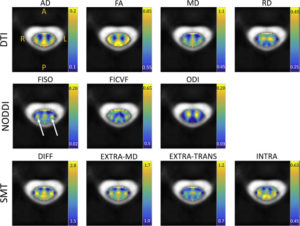
Multi-compartment tissue modeling using diffusion magnetic resonance imaging has proven valuable in the brain, offering novel indices sensitive to the tissue microstructural environment in vivo on clinical MRI scanners. However, application, characterization, and validation of these models in the spinal cord remain relatively under-studied. In this study, we apply a diffusion “signal” model (diffusiontensor imaging, DTI) and two commonly implemented “microstructural” models (neurite orientation dispersion and density imaging, NODDI; spherical mean technique, SMT) in the human cervical spinal cord of twenty-one healthy controls. We first provide normative values of DTI, SMT, and NODDI indices in a number of white matter ascending and descending pathways, as well as various gray matter regions. We then aim to validate the sensitivity and specificity of these diffusion-derived contrasts by relating these measures to indices of the tissue microenvironment provided by a histological template. We find that DTI indices are sensitive to a number of microstructural features, but lack specificity. The microstructural models also show sensitivity to a number of microstructure features; however, they do not capture the specific microstructural features explicitly modelled. Although often regarded as a simple extension of the brain in the central nervous system, it may be necessary to re-envision, or specifically adapt, diffusion microstructural models for application to the human spinal cord with clinically feasible acquisitions – specifically, adjusting, adapting, and re-validating the modeling as it relates to both theory (i.e. relevant biology, assumptions, and signal regimes) and parameter estimation (for example challenges of acquisition, artifacts, and processing).