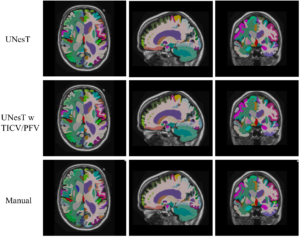
Enhancing Hierarchical Transformers for Whole Brain Segmentation with Intracranial Measurements Integration
Xin Yu, Yucheng Tang, Qi Yang, Ho Hin Lee, Shunxing Bao, Yuankai Huo, and Bennett A. Landman. “Enhancing Hierarchical Transformers for Whole Brain Segmentation with Intracranial Measurements Integration.” SPIE Medical Imaging 2024
Full text: NIHMSID
Abstract:
Whole brain segmentation with magnetic resonance imaging (MRI) enables the non-invasive measurement of brain regions, including total intracranial volume (TICV) and posterior fossa volume (PFV). Enhancing the existing whole brain segmentation methodology to incorporate intracranial measurements offers a heightened level of comprehensiveness in the analysis of brain structures. Despite its potential, the task of generalizing deep learning techniques for intracranial measurements faces data availability constraints due to limited manually annotated atlases encompassing whole brain and TICV/PFV labels. In this paper, we enhancing the hierarchical transformer UNesT for whole brain segmentation to achieve segmenting whole brain with 133 classes and TICV/PFV simultaneously. To address the problem of data scarcity, the model is first pretrained on 4859 T1-weighted (T1w) 3D volumes sourced from 8 different sites. These volumes are processed through a multi-atlas segmentation pipeline for label generation, while TICV/PFV labels are unavailable. Subsequently, the model is finetuned with 45 T1w 3D volumes from Open Access Series Imaging Studies (OASIS) where both 133 whole brain classes and TICV/PFV labels are available. We evaluate our method with Dice similarity coefficients(DSC). We show that our model is able to conduct precise TICV/PFV estimation while maintaining the 132 brain regions performance at a comparable level. Code and trained model are available at: https://github.com/MASILab/UNesT/wholebrainSeg.