Semi-Supervised Multi-Organ Segmentation through Quality Assurance Supervision
Ho Hin Lee, Yucheng Tang, Olivia Tang, Yuchen Xu, Yunqiang Chen, Dashan Gao, Shizhong Han, Riqiang Gao, Michael R. Savona, Richard G. Abramson, Yuankai Huo, Bennett A. Landman, “Semi-Supervised Multi-Organ Segmentation through Quality Assurance Supervision”, SPIE MI:IP 2020. Houston, TX.
Link: https://arxiv.org/abs/1911.05113
Abstract
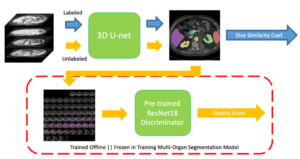
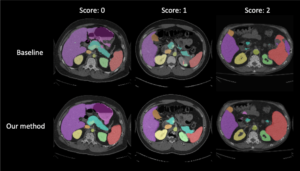
Human in-the-loop quality assurance (QA) is typically performed after medical image segmentation to ensure that the systems are performing as intended, as well as identifying and excluding outliers. By performing QA on large-scale, previously unlabeled testing data, categorical QA scores (e.g. “successful” versus “unsuccessful”) can be generated. Unfortunately, the precious use of resources for human in-the-loop QA scores are not typically reused in medical image machine learning, especially to train a deep neural network for image segmentation. Herein, we perform a pilot study to investigate if the QA labels can be used as supplementary supervision to augment the training process in a semi-supervised fashion. In this paper, we propose a semi-supervised multi-organ segmentation deep neural network consisting of a traditional segmentation model generator and a QA involved discriminator. An existing 3-D abdominal segmentation network is employed, while the pre-trained ResNet-18 network is used as discriminator. A large-scale dataset of 2027 volumes are used to train the generator, whose 2-D montage images and segmentation mask with QA scores are used to train the discriminator. To generate the QA scores, the 2-D montage images were reviewed manually and coded 0 (success), 1 (errors consistent with published performance), and 2 (gross failure). Then, the ResNet-18 network was trained with 1623 montage images in equal distribution of all three code labels and achieved an accuracy 94% for classification predictions with 404 montage images withheld for the test cohort. To assess the performance of using the QA supervision, the discriminator was used as a loss function in a multi-organ segmentation pipeline. The inclusion of QA-loss function boosted performance on the unlabeled test dataset from 714 patients to 951 patients over the baseline model. Additionally, the number of failures decreased from 606 (29.90%) to 402 (19.83%). The contributions of the proposed method are threefold: We show that (1) the QA scores can be used as a loss function to perform semi-supervised learning for unlabeled data, (2) the well trained discriminator is learnt by QA score rather than traditional “true/false”, and (3) the performance of multi-organ segmentation on unlabeled datasets can be fine-tuned with more robust and higher accuracy than the original baseline method. The use of QA-inspired loss functions represents a promising area of future research and may permit tighter integration of supervised and semi-supervised learning.