Deep Multi-path Network Integrating Incomplete Biomarker and Chest CT Data for Evaluating Lung Cancer Risk
Riqiang Gao, Yucheng Tang, Kaiwen Xu, Michael N. Kammer, Sanja L. Antic, Steve Deppen, Kim L. Sandler, Pierre P. Massion, Yuankai Huo, Bennett A. Landman,Deep Multi-path Network Integrating Incomplete Biomarker and Chest CT Data for Evaluating Lung Cancer Risk, SPIE, Medical Imaging, 2021.
Full text: https://arxiv.org/abs/2010.09524
Abstract
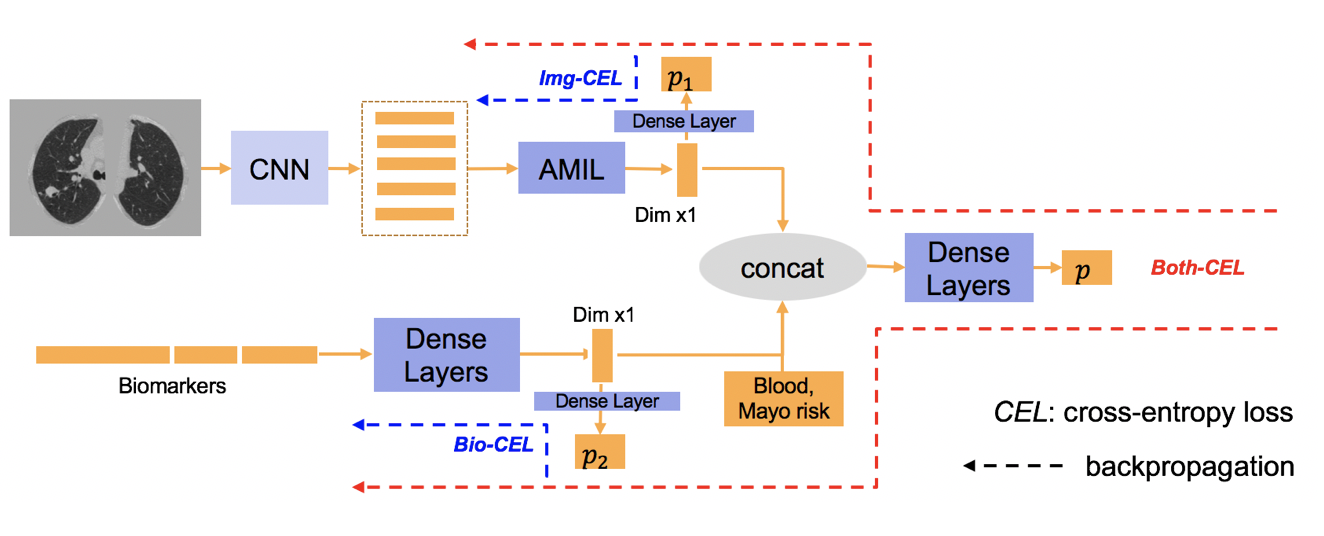
Clinical data elements (CDEs) (e.g., age, smoking history), blood markers and chest computed tomography (CT) structural features have been regarded as effective means for assessing lung cancer risk. These independent variables can provide complementary information and we hypothesize that combining them will improve the prediction accuracy. In practice, not all patients have all these variables available. In this paper, we propose a new network design, termed as multi-path multi-modal missing network (M3Net), to integrate the multi-modal data (i.e., CDEs, biomarker and CT image) considering missing modality with multiple paths neural network. Each path learns discriminative features of one modality, and different modalities are fused in a second stage for an integrated prediction. The network can be trained end-to-end with both medical image features and CDEs/biomarkers, or make a prediction with single modality. We evaluate M3Net with datasets including three sites from the Consortium for Molecular and Cellular Characterization of Screen-Detected Lesions (MCL) project. Our method is cross validated within a cohort of 1291 subjects (383 subjects with complete CDEs/biomarkers and CT images), and externally validated with a cohort of 99 subjects (99 with complete CDEs/biomarkers and CT images). Both cross-validation and external-validation results show that combining multiple modality significantly improves the predicting performance of single modality. The results suggest that integrating subjects with missing either CDEs/biomarker or CT imaging features can contribute to the discriminatory power of our model (p < 0.05, bootstrap two-tailed test). In summary, the proposed M3Net framework provides an effective way to integrate image and non-image data in the context of missing information.