Reducing Positional Variance in Cross-sectional Abdominal CT Slices with Deep Conditional Generative Models
Xin Yu*, Qi Yang*, Yucheng Tang, Riqiang Gao, Shunxing Bao, Leon Y. Cai, Ho Hin Lee, Ann Zenobia Moore, Luigi Ferrucci, Bennett A. Landman, “Reducing Positional Variance in Cross-sectional Abdominal CT Slices with Deep Conditional Generative Models”, MICCAI 2022
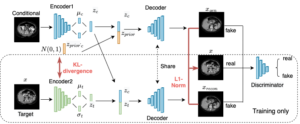
2D low-dose single-slice abdominal computed tomography (CT) slice enables direct measurements of body composition, which are critical to quantitatively characterizing health relationships on aging. However, longitudinal analysis of body composition changes using 2D abdominal slices is challenging due to positional variance between longitudinal slices acquired in different years. To reduce the positional variance, we extend the conditional generative models to our C-SliceGen that takes an arbitrary axial slice in the abdominal region as the condition and generates a defined vertebral level slice by estimating the structural changes in the latent space. Experiments on 1170 subjects from an in-house dataset and 50 subjects from BTCV MICCAI Challenge 2015 show that our model can generate high quality images in terms of realism and similarity. External experiments on 20 subjects from the Baltimore Longitudinal Study of Aging (BLSA) dataset that contains longitudinal single abdominal slices validate that our method can harmonize the slice positional variance in terms of muscle and visceral fat area. Our approach provides a promising direction of mapping slices from different vertebral levels to a target slice to reduce positional variance for single slice longitudinal analysis.