UNesT: Local Spatial Representation Learning with Hierarchical Transformer for Efficient Medical Segmentation
Xin Yu, Qi Yang, Yinchi Zhou, Leon Y. Cai , Riqiang Gao, Ho Hin Lee, Thomas Li, Shunxing Bao, Zhoubing Xu, Thomas A. Lasko, Richard G. Abramson, Zizhao Zhang, Yuankai Huo, Bennett A. Landman, Yucheng Tang
Paper: https://arxiv.org/abs/2209.14378
Code: https://github.com/Project-MONAI/model-zoo/tree/dev/models
Abstract
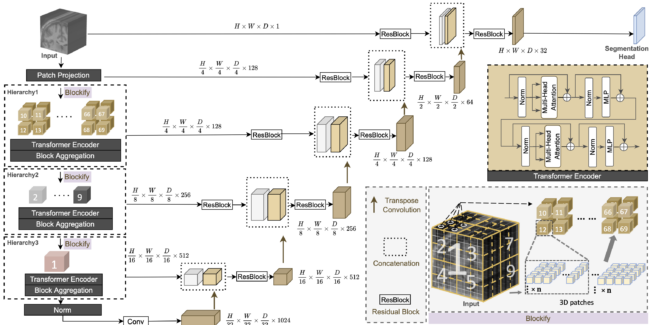
Transformer-based models, capable of learning better global dependencies, have recently demonstrated exceptional repre- sentation learning capabilities in computer vision and medical image analysis. Transformer reformats the image into separate patches and realizes global communication via the self-attention mechanism. However, positional information between patches is hard to preserve in such 1D sequences, and loss of it can lead to sub-optimal performance when dealing with large amounts of heterogeneous tissues of various sizes in 3D medical image segmentation. Additionally, current methods are not robust and efficient for heavy-duty medical segmentation tasks such as predicting a large number of tissue classes or modeling globally inter-connected tissues structures. To address such challenges and inspired by the nested hierarchical structures in vision transformer, we proposed a novel 3D medical image segmentation method (UNesT), employing a simplified and faster-converging transformer encoder design that achieves local communication among spatially adjacent patch sequences by aggregating them hierarchically. We extensively validate our method on multiple challenging datasets, consisting of multiple modalities, anatomies, and a wide range of tissue classes including 133 structures in the brain, 14 organs in the abdomen, 4 hierarchical components in the kidneys, and inter-connected kidney tumors. We show that UNesT consistently achieves state-of-the-art performance and evaluate its generalizability and data efficiency. Particularly, the model achieves whole brain segmentation task complete ROI with 133 tissue classes in a single network, outperforms prior state-of-the-art method SLANT27 ensembled with 27 networks, our model performance increases the mean DSC score of the publicly available Colin and CANDI dataset from 0.7264 to 0.7444 and from 0.6968 to 0.7025, respectively. Code, pre-trained models, and use case pipeline are integrated with the open-source framework MONAI available at: https://github.com/Project-MONAI/model-zoo/tree/dev/models.